Abstract
The ability to regenerate missing body parts exists throughout the animal kingdom. Positional information is crucial for regeneration, but how it is harboured and used by differentiated tissues is poorly understood. In planarians, positional information has been identified from study of phenotypes caused by RNA interference in which the wrong tissues are regenerated. For example, inhibition of the Wnt signalling pathway leads to regeneration of heads in place of tails1,2,3. Characterization of these phenotypes has led to the identification of position control genes (PCGs)—genes that are expressed in a constitutive and regional manner and are associated with patterning. Most PCGs are expressed within planarian muscle4; however, how muscle is specified and how different muscle subsets affect regeneration is unknown. Here we show that different muscle fibres have distinct regulatory roles during regeneration in the planarian Schmidtea mediterranea. myoD is required for formation of a specific muscle cell subset: the longitudinal fibres, oriented along the anterior–posterior axis. Loss of longitudinal fibres led to complete regeneration failure because of defects in regeneration initiation. A different transcription factor-encoding gene, nkx1-1, is required for the formation of circular fibres, oriented along the medial–lateral axis. Loss of circular fibres led to a bifurcated anterior–posterior axis with fused heads forming in single anterior blastemas. Whereas muscle is often viewed as a strictly contractile tissue, these findings reveal that different muscle types have distinct and specific regulatory roles in wound signalling and patterning to enable regeneration.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Petersen, C. P. & Reddien, P. W. Smed-βcatenin-1 is required for anteroposterior blastema polarity in planarian regeneration. Science 319, 327–330 (2008)
Gurley, K. A., Rink, J. C. & Sánchez Alvarado, A. β-catenin defines head versus tail identity during planarian regeneration and homeostasis. Science 319, 323–327 (2008)
Iglesias, M., Gomez-Skarmeta, J. L., Saló, E. & Adell, T. Silencing of Smed-βcatenin-1 generates radial-like hypercephalized planarians. Development 135, 1215–1221 (2008)
Witchley, J. N., Mayer, M., Wagner, D. E., Owen, J. H. & Reddien, P. W. Muscle cells provide instructions for planarian regeneration. Cell Reports 4, 633–641 (2013)
Cebrià, F., Vispo, M., Newmark, P., Bueno, D. & Romero, R. Myocyte differentiation and body wall muscle regeneration in the planarian Girardia tigrina. Dev. Genes Evol. 207, 306–316 (1997)
Cebrià, F. Planarian body-wall muscle: Regeneration and function beyond a simple skeletal support. Front. Cell Dev. Biol. 4, 8 (2016)
Buckingham, M. & Rigby, P. W. Gene regulatory networks and transcriptional mechanisms that control myogenesis. Dev. Cell 28, 225–238 (2014)
Kassar-Duchossoy, L. et al. Mrf4 determines skeletal muscle identity in Myf5:Myod double-mutant mice. Nature 431, 466–471 (2004)
Baugh, L. R. & Hunter, C. P. MyoD, modularity, and myogenesis: conservation of regulators and redundancy in C. elegans. Genes Dev. 20, 3342–3346 (2006)
Balagopalan, L., Keller, C. A. & Abmayr, S. M. Loss-of-function mutations reveal that the Drosophila nautilus gene is not essential for embryonic myogenesis or viability. Dev. Biol. 231, 374–382 (2001)
Reddien, P. W., Bermange, A. L., Murfitt, K. J., Jennings, J. R. & Sánchez Alvarado, A. Identification of genes needed for regeneration, stem cell function, and tissue homeostasis by systematic gene perturbation in planaria. Dev. Cell 8, 635–649 (2005)
Cowles, M. W. et al. Genome-wide analysis of the bHLH gene family in planarians identifies factors required for adult neurogenesis and neuronal regeneration. Development 140, 4691–4702 (2013)
Scimone, M. L., Lapan, S. W. & Reddien, P. W. A forkhead transcription factor is wound-induced at the planarian midline and required for anterior pole regeneration. PLoS Genet. 10, e1003999 (2014)
Vogg, M. C. et al. Stem cell-dependent formation of a functional anterior regeneration pole in planarians requires Zic and Forkhead transcription factors. Dev. Biol. 390, 136–148 (2014)
Vásquez-Doorman, C. & Petersen, C. P. zic-1 expression in planarian neoblasts after injury controls anterior pole regeneration. PLoS Genet. 10, e1004452 (2014)
LoCascio, S. A., Lapan, S. W. & Reddien, P. W. Eye absence does not regulate planarian stem cells during eye regeneration. Dev. Cell 40, 381–391.e3 (2017)
Wurtzel, O. et al. A generic and cell-type-specific wound response precedes regeneration in planarians. Dev. Cell 35, 632–645 (2015)
Wenemoser, D., Lapan, S. W., Wilkinson, A. W., Bell, G. W. & Reddien, P. W. A molecular wound response program associated with regeneration initiation in planarians. Genes Dev. 26, 988–1002 (2012)
Wenemoser, D. & Reddien, P. W. Planarian regeneration involves distinct stem cell responses to wounds and tissue absence. Dev. Biol. 344, 979–991 (2010)
Pellettieri, J. et al. Cell death and tissue remodeling in planarian regeneration. Dev. Biol. 338, 76–85 (2010)
Kakugawa, S. et al. Notum deacylates Wnt proteins to suppress signalling activity. Nature 519, 187–192 (2015)
Zhang, X. et al. Notum is required for neural and head induction via Wnt deacylation, oxidation, and inactivation. Dev. Cell 32, 719–730 (2015)
Petersen, C. P. & Reddien, P. W. Polarized notum activation at wounds inhibits Wnt function to promote planarian head regeneration. Science 332, 852–855 (2011)
Gaviño, M. A., Wenemoser, D., Wang, I. E. & Reddien, P. W. Tissue absence initiates regeneration through Follistatin-mediated inhibition of Activin signaling. eLife 2, e00247 (2013)
Roberts-Galbraith, R. H. & Newmark, P. A. Follistatin antagonizes Activin signaling and acts with Notum to direct planarian head regeneration. Proc. Natl Acad. Sci. USA 110, 1363–1368 (2013)
Adler, C. E. & Sánchez Alvarado, A. PHRED-1 is a divergent neurexin-1 homolog that organizes muscle fibers and patterns organs during regeneration. Dev. Biol. 427, 165–175 (2017)
Scimone, M. L., Cote, L. E., Rogers, T. & Reddien, P. W. Two FGFRL-Wnt circuits organize the planarian anteroposterior axis. eLife 5, e12845 (2016)
Knirr, S., Azpiazu, N. & Frasch, M. The role of the NK-homeobox gene slouch (S59) in somatic muscle patterning. Development 126, 4525–4535 (1999)
Liu, S. Y. et al. Reactivating head regrowth in a regeneration-deficient planarian species. Nature 500, 81–84 (2013)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010)
Rouhana, L. et al. RNA interference by feeding in vitro-synthesized double-stranded RNA to planarians: methodology and dynamics. Dev. Dyn. 242, 718–730 (2013)
Ross, K. G. et al. Novel monoclonal antibodies to study tissue regeneration in planarians. BMC Dev. Biol. 15, 2 (2015)
Schierwater, B. et al. The early ANTP gene repertoire: insights from the placozoan genome. PLoS One 3, e2457 (2008)
Fahey, B. & Degnan, B. M. Origin and evolution of laminin gene family diversity. Mol. Biol. Evol. 29, 1823–1836 (2012)
Scimone, M. L., Kravarik, K. M., Lapan, S. W. & Reddien, P. W. Neoblast specialization in regeneration of the planarian Schmidtea mediterranea. Stem Cell Reports 3, 339–352 (2014)
Acknowledgements
We thank N. Watson and A. Mahowald for transmission electron microscopy; S. LoCascio for eye resections; C.-C. Chen for V5277; and M. Fedorovsky for illustrations. We acknowledge support from NIH R01GM080639 and the Eleanor Schwartz Charitable Foundation. P.W.R. is an Investigator of the HHMI and an associate member of the Broad Institute of Harvard and MIT.
Author information
Authors and Affiliations
Contributions
M.L.S. and L.E.C. carried out RNAi characterization, RNA-seq and TEM; L.E.C. carried out phylogenetic analysis; M.L.S., L.E.C. and P.W.R. discussed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks C. Petersen and A. Sánchez Alvarado for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 myoD is a myogenic gene in planarians that is specific for longitudinal muscle fibres.
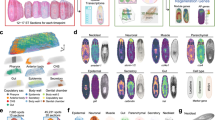
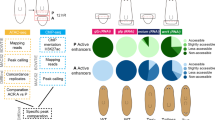
a, Co-expression of myoD (myoD and snail probes pooled) and the neoblast marker smedwi-1 (five animals, two experiments; white, double-positive cells). b, Live image shows a longer and thinner myoD(RNAi) animal (quantification in Fig. 1c). c, Reduction in BWM (collagen+) but not intestinal muscle (dd_6811+)17 in an uninjured myoD(RNAi) animal after ten dsRNA feedings. d, myoD or control RNAi animals have comparable diagonal fibre numbers. Scale bars, 10 μm. e, Heat map shows genes downregulated (log2[fold change] < 0, adjusted P value < 0.001) in uninjured myoD(RNAi) animals. Green, muscle-enriched genes from single-cell RNA-seq data17 (AUC > 0.8; 43 out of 123 genes). Each column is a replicate. f, Heat map showing that other tissue-enriched gene expression is not affected in myoD(RNAi) animals. Mean of tissue-enriched genes17 (AUC > 0.8) is used. Each column is a replicate. g, Phylogenetic analysis of homeodomain transcription factors. Accession numbers are in Supplementary Table 2. Tree shows 105 proteins from diverse organisms. Maximum likelihood analyses were run using PhyML with 100 bootstrap replicates. All maximum likelihood bootstrap values >20 are shown. Bf: Branchiostoma floridae; Ce: Caenorhabditis elegans; Ct: Capitella teleta; Dm: Drosophila melanogaster; Dr: Danio rerio; Dt: Discocelis tigrina; Gg: Gallus gallus; Hs: Homo sapiens; Lg: Lottia gigantea; Nv: Nematostella vectensis; Od: Oikopleura dioica; Sm: Schmidtea mediterranea; Sp: Strongylocentrotus purpuratus; Xl: Xenopus laevis. The lhx2/9 tree was previously reported36. All FISH panels are representative images of two independent experiments. Bottom left number: animals with phenotype out of total tested. Anterior, up. Scale bars, 100 μm unless indicated.
Extended Data Figure 2 A subset of transcription factors is expressed in longitudinal muscle fibres.
a, Seurat maps show expression of transcription factors downregulated in myoD(RNAi) animals within a reported17 single-cell RNA-seq experiment. Each dot represents a cell. b, Co-expression of those transcription factors and the BWM marker collagen in uninjured animals. Scale bars, 100 μm; insets, 10 μm. Number of cells expressing both genes (white) within the total number of cells expressing the transcription factor (magenta) is shown (five animals, two experiments). c, Co-expression of myoD and transcription factors (white) within the total myoD+ cells (green) in uninjured animals (five animals, two experiments). Scale bars, 10 μm. d, Reduced expression of transcription factors in uninjured myoD(RNAi) animals (five animals, two experiments). Scale bars, 100 μm. e, Heat map shows two clusters of muscle cells (myoD+ and nkx1-1+) and co-expression with other muscle transcription factors, PCGs and muscle regional expressed genes. Most PCG expression is widely distributed across all muscle cells. Each column is a single cell. Analyses using previously reported single-cell muscle data27. Asterisks denote best human BLAST hit. Cartoon shows in different colours the regions from which single muscle cell data were collected27. f, No major differences in BWM structure and expression of longitudinal fibre-enriched PCGs in the different RNAi conditions tested (10 dsRNA feedings, uninjured animals, five animals per group, two experiments). Scale bars, 50 μm. Anterior, up.
Extended Data Figure 3 A subset of PCGs is enriched in longitudinal muscle fibres.
a, Phylogenetic analysis of all planarian netrins. Accession numbers used for the tree are in Supplementary Table 2. Tree shows 20 Netrin proteins from diverse organisms, which were aligned using MUSCLE with default settings and trimmed with Gblocks. Maximum likelihood analyses were run using PhyML with 1,000 bootstrap replicates. All maximum likelihood bootstrap values are shown above or below the respective branch. Dm, Drosophila melanogaster; Sp, Strongylocentrotus purpuratus; Bf, Branchiostoma floridae; Hs, Homo sapiens; Dr, Danio rerio; Ce, Caenorhabditis elegans; Pd, Platynereis dumerilii; Cg, Crassostrea gigas; Sm, Schmidtea mediterranea; Nv, Nematostella vectensis. Cartoon shows protein domain structure. b, Reduced expression of PCGs following myoD RNAi and co-expression of pooled myoD and snail with those PCGs in uninjured animals. Scale bars, 100 μm (top panels), 10 μm (bottom panels). Red box in left cartoon depicts location of image shown. FISH images are representative of two independent experiments. Bottom left number: animals with phenotype out of total tested. Anterior, up.
Extended Data Figure 4 myoD is required for regeneration.
a, Normal wound contraction in myoD(RNAi) animals. Trunk fragments are shown 30 min after amputation (15 animals, 3 experiments). b, Lack of anterior (notum+) and posterior (wnt1+) pole cells (top) and BWM structure (bottom) at 72 hpa during regeneration in myoD(RNAi) animals. c, Anterior pole (notum+) and BWM structure in uninjured animals (b, c; ten animals per group, three experiments). Scale bars, 50 μm. d, Neoblasts did not specify into eye progenitors (ovo+) in myoD(RNAi) animals at 72 hpa (one experiment). e, Homeostatic eye replacement 11 days after eye resection in myoD(RNAi) animals (ten animals per group, one experiment). Scale bars, 500 μm. f, Normal epidermis, neoblast and muscle expression of wound-induced genes in myoD(RNAi) animals 6 hpa. g, Graphs show reduced numbers of notum+ and fst+ cells and longitudinal fibres in myoD(RNAi) animals at 18 hpa after different numbers of dsRNA feedings. Cartoon shows the region counted. Linear correlation between fst+ cells and longitudinal fibres. Regression coefficient, R2 = 0.6928. Two-tailed Student’s t-test was performed. P values are shown in graphs. Mean ± s.d. shown in all graphs. Bottom left number: animals with phenotype out of total tested. Anterior, up. Scale bars, 100 μm unless indicated.
Extended Data Figure 5 myoD is required for the regenerative response.
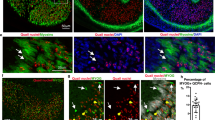
a, Partial co-expression of myoD (myoD and snail probes pooled) and muscle-wound induced genes inhibin-1 and nlg-1 at 6 hpa (four animals, one experiment). Number in white indicates co-expression within total number of counted cells expressing the wound-induced gene (green). Scale bars, 10 μm. b, Heat map shows expression of all 128 wound-induced genes17 from anterior-facing wounds of regenerating tail fragments of control, myoD and fst RNAi animals at different time points after amputation. Each column is a replicate. c, Heat map shows expression of neoblast genes17 (AUC > 0.8) in anterior-facing wounds of regenerating tail fragments of control, myoD and fst RNAi animals at different time points after amputation. Each column is a replicate. Right, no neoblast (smedwi-1+) accumulation at wounds (yellow arrow) 48 hpa in myoD(RNAi) tail fragments after eight dsRNA feedings (six animals, one experiment).
Extended Data Figure 6 The lack of a regenerative response in myoD(RNAi) animals is suppressed by act-1 inhibition.
a, Heat map shows failure to re-scale posterior or initiate expression of anterior regionally expressed muscle genes defined in a previous study27 in tail fragments of myoD and fst RNAi animals. Each column represents a replicate. Asterisks denote best human BLAST hit. Right, failure to re-scale wntP-2 in tail fragments of myoD(RNAi) animals at 48 hpa (eight animals for control, ten animals for myoD RNAi, three experiments). White arrows point to wound site, yellow arrows point to wntP-2 expression. Dotted line indicates wound site. b, Re-scaling of wntP-2 and expression of anterior pole cells (notum+) in tail fragments of double myoD and act-1 RNAi animals at 48 hpa (two experiments). White arrows point to wound site, yellow arrows point to wntP-2 expression. c, Comparable loss of longitudinal fibres (yellow arrows) in both groups (myoD and control RNAi animals, and myoD and act-1 RNAi animals; quantification in Fig. 2i). Scale bars, 10 μm. Loss of snail expression in both groups (two experiments). d, Long-term double myoD; control and myoD; act-1 RNAi animals failed to regenerate (two experiments). e, β-catenin-1 inhibition did not suppress the regeneration defect of myoD(RNAi) animals (one experiment). However, homeostatic ectopic stretching head-like outgrowths (red arrows) formed around the periphery of β-catenin-1(RNAi) animals. Asterisk denotes absence of anterior blastema, yellow arrows point to ectopic eyes. Bottom left number, animals with phenotype out of total tested. Anterior, up. Scale bars, 100 μm unless indicated.
Extended Data Figure 7 Transverse injuries trigger more fst expression than longitudinal injuries.
a, Higher numbers of fst+ cells relative to nlg-1+ cells in transverse versus sagittal amputations at 6 hpa. b, Higher numbers of wound-induced fst+ cells relative to nlg-1+ cells in transverse versus longitudinal incisions at 6 hpa. Yellow dotted lines show site of incision. Right graph shows total numbers of fst+, inhibin-1+ and nlg-1+ cells per length of wound at 6 hpa. Two-tailed Student’s t-test was performed. Mean ± s.d. are shown. ns, not significant. P values are shown. Red box in cartoon depicts location of image shown; dotted line, plane of injury performed. c, Live images show that myoD(RNAi) animals regenerated small blastemas following sagittal amputations (three experiments). Scale bars, 500 μm. All FISH and live images shown are anterior, up. Bottom left number, animals with phenotype out of total tested. Scale bars, 100 μm unless indicated.
Extended Data Figure 8 nkx1-1 specifies circular muscle fibres.
a, A minor fraction of nkx1-1+ cells co-expresses the neoblast marker smedwi-1. b, Expression of nkx1-1 within the collagen+ BWM in an uninjured animal (five animals, two experiments (a, b)). c, Animals become wider after nkx1-1 RNAi (quantification in Fig. 3b, 13 control and 17 nkx1-1 RNAi animals, three experiments). d, Loss of circular fibres in nkx1-1(RNAi) animals (quantification in Fig. 3c). Scale bars in expanded views, 10 μm. e, Comparable numbers of diagonal fibres in nkx1-1 and control RNAi animals (three experiments). Scale bars, 10 μm. f, Heat map shows 43 muscle-enriched genes downregulated (log2[fold change] < 0, adjusted P value < 0.001) in both uninjured myoD and nkx1-1 RNAi animals. Each column represents a replicate. g, Co-expression of nkx1-1 and PCGs (top) and reduced PCG expression in uninjured nkx1-1(RNAi) animals after 12 dsRNA feedings. Numbers in white indicate double-positive cells within the total number of counted cells expressing the PCG (green) (three experiments). Red box on cartoons depicts location of image shown. Bottom left number, animals with phenotype out of total tested. Anterior, up. Scale bars, 100 μm unless indicated.
Extended Data Figure 9 nkx1-1 is required for normal medial–lateral patterning during regeneration.
a, Normal wound contraction in nkx1-1(RNAi) animals. Trunk fragments are shown 30 min after amputation (15 animals, three experiments). b, Live images of regenerating (9–14 dpa) nkx1-1(RNAi) animals after 12 dsRNA feedings following a transverse amputation (five experiments). Scale bars, 500 μm. Graph shows quantification of eye distance relative to total animal length (ten animals per group, two experiments). Two-tailed Student’s t-test was performed. Mean ± s.d. are shown. Lines in cartoon on top show where the measurements were taken. c, Increased width of midline (slit) and brain lobes (netrin-2) in nkx1-1(RNAi) animals (three experiments). d, Anterior pole (notum+) and BWM fibres in intact (top) and regenerating (bottom, 72 hpa and 9 dpa) nkx1-1(RNAi) animals (12 animals per group, two experiments). e, Live images of regenerating nkx1-1(RNAi) animals following a sagittal amputation (three experiments). Yellow arrow points to ectopic eye. Scale bars, 200 μm. Dotted lines in cartoons show amputation sites. Red box in cartoons depicts location of image shown. All FISH and live images shown are anterior, up. Bottom left number, animals with phenotype out of total tested. Scale bars, 100 μm, unless indicated.
Extended Data Figure 10 Longitudinal and circular muscle fibres are required for normal patterning during homeostatic tissue turnover.
a, Reduced numbers of longitudinal and circular muscle fibres but comparable numbers of pharynx muscle fibres in control and double myoD and nkx1-1 RNAi animals. Dotted line in cartoon indicates that pharynx muscle is more internal than the BWM fibres (ten animals per group, three experiments). Scale bars, 10 μm. b, Comparable numbers of diagonal fibres in control and double myoD and nkx1-1 RNAi animals (ten animals per group, three experiments). Scale bars, 10 μm. c, Cross sections showing comparable dorsal–ventral fibres (yellow arrows) in control and double myoD and nkx1-1 RNAi animals (eight animals per group, two experiments). d, Intestinal muscle (dd_6811) is not affected in double myoD and nkx1-1 RNAi animals (six animals per group, two experiments). e, Ectopic posterior eyes (opsin+, white arrows) and reduced expression of PCGs (anterior: sFRP-1, ndk, fz5/8-4; midbody: ndl-3 and sFRP-2; posterior: wntP-2) in double myoD and nkx1-1 RNAi animals compared to controls (four animals per FISH, two experiments). Elongated brain lobes were also observed in double myoD and nkx1-1 RNAi animals compared to controls (chat+ and gd+ cells). NB.22.1e marks epidermal cells at the boundary of the animals (five animals per group, two experiments). f, Graph shows similar length-to-width ratios in double myoD and nkx1-1 RNAi animals and control RNAi animals (eight for control, twelve for double RNAi, two experiments). Two-tailed Student’s t-test was performed. Mean ± s.d. are shown. Red lines in cartoon indicate where measurements were taken. g, Live image of a dying double myoD and nkx1-1 RNAi animal. Lysis occurs 12–20 weeks after first dsRNA feeding. Scale bars, 500 μm. Red box in cartoons depicts location of image shown. All FISH and live images shown are anterior, up. Bottom left number, animals with phenotype out of total tested. Scale bars, 100 μm unless indicated.
Supplementary information
Supplementary Table 1 - Summary of RNA-sequencing analyses.
This file contains RNA-sequencing analyses of myoD and nkx1-1 RNAi animals during normal tissue turnover and regeneration. RNA-sequencing analyses of follistatin RNAi animals during regeneration. (XLSX 10533 kb)
Supplementary Table 2
This file contains accession numbers for all proteins used for phylogenetic analyses. (XLSX 43 kb)
Body wall muscle fibres in a control RNAi animal
Immunostaining using 6G10 and V5277 antibodies show different layers of muscle fibres in the planarian BWM. The video shows the layers of muscle fibres on the ventral side of the animal, and goes from the outer layer (closer to the epidermis) to the inner layer (closer to the parenchyma). First in the video, the circular muscle fibres can be observed, next the diagonal fibres and last the longitudinal fibres are observed. Muscle fibres from a 5-7 mm animal have been previously estimated to measure between 150-200 μm of length (Baguñà, J. & Romero, R. Quantitative analysis of cell types during growth, degrowth and regeneration in the planarians Dugesia mediterranea and Dugesia tigrina. Hydrobiologia 84, 181-194, (1981)) (MP4 2509 kb)
Loss of longitudinal muscle fibres in a myoD(RNAi) animal
Immunostaining using 6G10 and V5277 antibodies shows loss of the longitudinal fiber layer in a myoD(RNAi) animal. Similar to SI video 1, the confocal z-stack was taken on the ventral side of the animal. From the epidermis into the animal parenchyma the circular and diagonal muscle fibre layers can be observed but very few longitudinal muscle fibres are left intact after inhibition of myoD. (MP4 1099 kb)
Loss of circular muscle fibres in a nkx1-1(RNAi) animal
Immunostaining using 6G10 and V5277 antibodies shows loss of the circular fibre layer (first layer observed in the video) in a nkx1-1(RNAi) animal. Similar to SI video 1, the confocal z-stack was taken on the ventral side of the animal. From the epidermis into the animal parenchyma few circular muscle fibres are left intact in the outer layer, but diagonal and longitudinal muscle layers are maintained after inhibition of nkx1-1. (MP4 954 kb)
Cephalic ganglia structure in a control RNAi animal
FISH shows glutamate decarboxylase (gd) RNA probe in green, DAPI in gray. The confocal z-stack was taken from the ventral side of the animal. In green, the arc of gd+ cells can be observed. (MP4 489 kb)
Duplication of a brain lobe in a nkx1-1(RNAi) regenerating animal
FISH shows ectopic glutamate decarboxylase expression in green, DAPI in gray. A wider brain with a small forming brain lobe can be observed. (MP4 477 kb)
Loss of body wall muscle fibres in a double myoD; nkx1-1(RNAi) animal
Immunostaining using 6G10 and V5277 antibodies shows loss of both circular fibre (first layer shown in the video) and longitudinal fibre (last layer shown in the video) layers and disorganization of BWM in a double myoD, nkx1-1(RNAi) animal. (MP4 501 kb)
Rights and permissions
About this article
Cite this article
Scimone, M., Cote, L. & Reddien, P. Orthogonal muscle fibres have different instructive roles in planarian regeneration. Nature 551, 623–628 (2017). https://doi.org/10.1038/nature24660
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24660
This article is cited by
-
The BAF chromatin remodeling complex licenses planarian stem cells access to ectodermal and mesodermal cell fates
BMC Biology (2023)
-
Acoel single-cell atlas reveals expression dynamics and heterogeneity of adult pluripotent stem cells
Nature Communications (2023)
-
Spatiotemporal transcriptomic atlas reveals the dynamic characteristics and key regulators of planarian regeneration
Nature Communications (2023)
-
Wnt/β-catenin signalling is required for pole-specific chromatin remodeling during planarian regeneration
Nature Communications (2023)
-
Finding the potency in planarians
Communications Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.