Abstract
Astrocytes are complex glial cells with numerous fine cellular processes that infiltrate the neuropil and interact with synapses. The mechanisms that control the establishment of astrocyte morphology are unknown, and it is unclear whether impairing astrocytic infiltration of the neuropil alters synaptic connectivity. Here we show that astrocyte morphogenesis in the mouse cortex depends on direct contact with neuronal processes and occurs in parallel with the growth and activity of synaptic circuits. The neuroligin family cell adhesion proteins NL1, NL2, and NL3, which are expressed by cortical astrocytes, control astrocyte morphogenesis through interactions with neuronal neurexins. Furthermore, in the absence of astrocytic NL2, the formation and function of cortical excitatory synapses are diminished, whereas inhibitory synaptic function is enhanced. Our findings highlight a previously undescribed mechanism of action for neuroligins and link astrocyte morphogenesis to synaptogenesis. Because neuroligin mutations have been implicated in various neurological disorders, these findings also point towards an astrocyte-based mechanism of neural pathology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Clarke, L. E. & Barres, B. A. Emerging roles of astrocytes in neural circuit development. Nat. Rev. Neurosci. 14, 311–321 (2013)
Khakh, B. S. & Sofroniew, M. V. Diversity of astrocyte functions and phenotypes in neural circuits. Nat. Neurosci. 18, 942–952 (2015)
Ma, Z., Stork, T., Bergles, D. E. & Freeman, M. R. Neuromodulators signal through astrocytes to alter neural circuit activity and behaviour. Nature 539, 428–432 (2016)
Oberheim, N. A. et al. Uniquely hominid features of adult human astrocytes. J. Neurosci. 29, 3276–3287 (2009)
Freeman, M. R. Specification and morphogenesis of astrocytes. Science 330, 774–778 (2010)
Burda, J. E. & Sofroniew, M. V. Reactive gliosis and the multicellular response to CNS damage and disease. Neuron 81, 229–248 (2014)
Stichel, C. C., Müller, C. M. & Zilles, K. Distribution of glial fibrillary acidic protein and vimentin immunoreactivity during rat visual cortex development. J. Neurocytol. 20, 97–108 (1991)
Morel, L., Higashimori, H., Tolman, M. & Yang, Y. VGluT1+ neuronal glutamatergic signaling regulates postnatal developmental maturation of cortical protoplasmic astroglia. J. Neurosci. 34, 10950–10962 (2014)
Cahoy, J. D. et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J. Neurosci. 28, 264–278 (2008)
Li, M. et al. Synaptogenesis in the developing mouse visual cortex. Brain Res. Bull. 81, 107–113 (2010)
Akerman, C. J., Smyth, D. & Thompson, I. D. Visual experience before eye-opening and the development of the retinogeniculate pathway. Neuron 36, 869–879 (2002)
Srinivasan, R. et al. New transgenic mouse lines for selectively targeting astrocytes and studying calcium signals in astrocyte processes in situ and in vivo. Neuron 92, 1181–1195 (2016)
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014)
Zhang, Y. et al. Purification and characterization of progenitor and mature human astrocytes reveals transcriptional and functional differences with mouse. Neuron 89, 37–53 (2016)
Bemben, M. A., Shipman, S. L., Nicoll, R. A. & Roche, K. W. The cellular and molecular landscape of neuroligins. Trends Neurosci. 38, 496–505 (2015)
Baudouin, S. & Scheiffele, P. SnapShot: Neuroligin-neurexin complexes. Cell 141, 908 (2010)
Craig, A. M. & Kang, Y. Neurexin-neuroligin signaling in synapse development. Curr. Opin. Neurobiol. 17, 43–52 (2007)
Venkatesh, H. S. et al. Neuronal activity promotes glioma growth through neuroligin-3 secretion. Cell 161, 803–816 (2015)
Samarelli, A. V. et al. Neuroligin 1 induces blood vessel maturation by cooperating with the α6 integrin. J. Biol. Chem. 289, 19466–19476 (2014)
Proctor, D. T. et al. Axo-glial communication through neurexin-neuroligin signaling regulates myelination and oligodendrocyte differentiation. Glia (2015). 10.1002/glia.22875
Graf, E. R., Zhang, X., Jin, S. X., Linhoff, M. W. & Craig, A. M. Neurexins induce differentiation of GABA and glutamate postsynaptic specializations via neuroligins. Cell 119, 1013–1026 (2004)
Murai, K. K., Nguyen, L. N., Irie, F., Yamaguchi, Y. & Pasquale, E. B. Control of hippocampal dendritic spine morphology through ephrin-A3/EphA4 signaling. Nat. Neurosci. 6, 153–160 (2003)
Dean, C. et al. Neurexin mediates the assembly of presynaptic terminals. Nat. Neurosci. 6, 708–716 (2003)
Gokce, O. & Südhof, T. C. Membrane-tethered monomeric neurexin LNS-domain triggers synapse formation. J. Neurosci. 33, 14617–14628 (2013)
Liang, J. et al. Conditional neuroligin-2 knockout in adult medial prefrontal cortex links chronic changes in synaptic inhibition to cognitive impairments. Mol. Psychiatry 20, 850–859 (2015)
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010)
Baldwin, K. T. & Eroglu, C. Molecular mechanisms of astrocyte-induced synaptogenesis. Curr. Opin. Neurobiol. 45, 113–120 (2017)
Poulopoulos, A. et al. Neuroligin 2 drives postsynaptic assembly at perisomatic inhibitory synapses through gephyrin and collybistin. Neuron 63, 628–642 (2009)
Varoqueaux, F. et al. Neuroligins determine synapse maturation and function. Neuron 51, 741–754 (2006)
Wang, Y. et al. Norrin/Frizzled4 signaling in retinal vascular development and blood brain barrier plasticity. Cell 151, 1332–1344 (2012)
Liddelow, S. A. et al. Neurotoxic reactive astrocytes are induced by activated microglia. Nature 541, 481–487 (2017)
Harris, K. D. & Shepherd, G. M. The neocortical circuit: themes and variations. Nat. Neurosci. 18, 170–181 (2015)
Allen, N. J. et al. Astrocyte glypicans 4 and 6 promote formation of excitatory synapses via GluA1 AMPA receptors. Nature 486, 410–414 (2012)
Christopherson, K. S. et al. Thrombospondins are astrocyte-secreted proteins that promote CNS synaptogenesis. Cell 120, 421–433 (2005)
Kucukdereli, H. et al. Control of excitatory CNS synaptogenesis by astrocyte-secreted proteins Hevin and SPARC. Proc. Natl Acad. Sci. USA 108, E440–E449 (2011)
Singh, S. K. & Eroglu, C. Neuroligins provide molecular links between syndromic and nonsyndromic autism. Sci. Signal. 6, re4 (2013)
Sun, C. et al. Identification and functional characterization of rare mutations of the neuroligin-2 gene (NLGN2) associated with schizophrenia. Hum. Mol. Genet. 20, 3042–3051 (2011)
Gao, R. & Penzes, P. Common mechanisms of excitatory and inhibitory imbalance in schizophrenia and autism spectrum disorders. Curr. Mol. Med. 15, 146–167 (2015)
Windrem, M. S. et al. Human iPSC glial mouse chimeras reveal glial contributions to schizophrenia. Cell Stem Cell 21, 195–208 (2017)
Chih, B., Gollan, L. & Scheiffele, P. Alternative splicing controls selective trans-synaptic interactions of the neuroligin–neurexin complex. Neuron 51, 171–178 (2006)
Singh, S. K. et al. Astrocytes assemble thalamocortical synapses by bridging NRX1α and NL1 via Hevin. Cell 164, 183–196 (2016)
Acknowledgements
This work was supported by grants from the National Institutes of Health (RO1 DA031833 to C.E., RO1 DE022743 to R.-R.J., F31 NS092419 to J.A.S.) and a Holland Trice Brain Research Award to C.E. K.T.B. was supported by Foerster-Bernstein Family and the Hartwell Foundation. We thank the NHLBI light microscopy core for STED imaging. We thank N. Allen, M. Bagnat, D. Silver and S. Soderling for critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
J.A.S. and C.E. designed the experiments and wrote the paper. All authors reviewed and edited the manuscript. J.A.S. performed experiments and analysed data. J.A.S. and J.R. performed immunohistochemistry and cell/synapse count analysis. R.-R.J. designed and D.L. and Y.-H.K. performed and analysed the electrophysiology experiments. J.A.S. and K.T.B. performed western blot analysis. J.A.S., E.E., and T.E. performed in vitro experiments and analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Astrocyte morphology is developmentally regulated in V1 cortex and occurs in parallel with sensory activity.
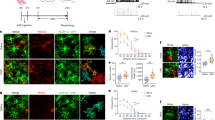
a–d, Dark rearing reduces astrocyte morphogenesis in V1 visual cortex. a, High-magnification images of V1 L1 astrocytes from Aldh1L1–EGFP mice labelled with EGFP and GFAP at P7 and P21. Finer EGFP-labelled processes, which emerge by P21, do not stain with GFAP (arrow). b, Representative single optical section confocal images of V1 visual cortex (left) and A1 auditory cortex (right) L4 astrocytes from normal reared (NR, top) and dark reared (DR, bottom) Aldh1L1–EGFP mice at P21. c, Fold change in astrocyte coverage of the neuropil in normal and dark-reared mice at each layer of V1 cortex normalized to normal L1. n = 10 ROIs per layer, 3 images per mouse, 4 mice per condition. d, Fold change in astrocyte coverage of the neuropil in normal and dark-reared mice at each layer of A1 cortex normalized to normal L1. n = 10 ROIs per layer, 3 images per mouse, 4 mice per condition. e–g, Postnatal astrocyte labelling by electroporation (PALE) effectively transfects L4–5 astrocytes with plasmid DNA. e, Schematic of PALE in newborn mice. Late P0 or early P1 mouse pups are sedated by hypothermia and injected with 1 μg plasmid DNA into the lateral ventricle of one hemisphere with a pulled glass pipette. The plasmid solutions are mixed with a small volume of fast green dye for visualizing injections. Plasmids used in this study for PALE encoded EGFP, membrane-targeted mCherry (mCherry–CAAX), shRNAs also encoding EGFP, HA-tagged NL-related constructs, or Cre recombinase. Five pulses of 100 V are applied to the mouse head, using tweezer-trodes with the positive terminal situated above one hemisphere of V1 cortex. Please see Supplementary Methods for details. f, Diagram of PALE injection site and cartoon of V1 cortical development following PALE. Electroporation of plasmids at P0–P1 transfects the radial glial stem cells (light green), which give rise to astrocytes and other glia. At P7, labelled radial glial remnants persist and sparsely labelled developing astrocytes (dark green) are present predominantly in the lower cortical layers, L4–5. Two weeks later, radial glial remnants disappear and labelled mature astrocytes (dark green) are present. g, Representative tile scan image of V1 cortex at P7 following PALE with shCtrl plasmid. Radial glial cell bodies are visible near the ventricular surface. The labelled radial glia extend basal processes dorsally where they branch to form multiple endfeet associated with the pia (dotted line). Labelled astrocytes are dispersed throughout L4-5 of V1 cortex. These labelled astrocytes are imaged by high magnification confocal microscopy for more detailed volumetric analysis. h–q, Dark rearing reduces astrocyte morphogenesis at the single-cell level in V1. h, Schematic of PALE to test soluble and membrane-targeted fluorophores in astrocyte neuropil infiltration measurements. pZac2.1_gfaABC1D plasmid vectors expressing EGFP or membrane-targeted mCherry (mCherry–CAAX) reporters under the control of the minimal human GFAP promoter are co-electroporated into P0 wild-type D1 mice. Mice are housed in normal (NR) or dark rearing (DR) conditions, then killed at P7 or P21. i, Representative high-magnification confocal images and neuropil infiltration volume (NIV) reconstructions of PALE V1 cortex L4–5 astrocytes at P7 (left) and P21 (right). EGFP+ (green) and mCherry–CAAX+ (red) astrocytes (co-localized equals yellow) were uploaded to Imaris and the NIVs for each fluorophore were 3D reconstructed (middle and bottom). j, Average NIVs of V1 L4–5 PALE astrocytes at P7 and P21 for both EGFP and mCherry–CAAX fluorophores. Three NIVs per cell, 18–20 cells per condition, 4 mice per condition. k, Fold change in NIV growth curves from P7 to P21 astrocytes of EGFP and mCherry–CAAX fluorophores, normalized to P7 for each fluorophore. Three NIVs per cell, 18–20 cells per condition, 4 mice per condition. As expected, the membrane-bound fluorescent protein mCherry–CAAX slightly but significantly increased the visualization of astrocyte infiltration into the neuropil (mCherry–CAAX:EGFP; P7 = 1.28 ± 0.07-fold, P = 0.001; P21 = 1.17 ± 0.04-fold, P = 0.005). l, Average NIV of P7 astrocytes from normal and dark-reared mice using EGFP and mCherry–CAAX fluorophores. Three NIVs per cell, 18–20 cells per condition, 4 mice per condition. m, Average NIV of P21 astrocytes from normal and dark-reared mice using EGFP and mCherry–CAAX fluorophores. Three NIVs per cell, 18–20 cells per condition, 4 mice per condition. n, Representative high-magnification confocal images and territory reconstructions of V1 P7 and P21 PALE astrocytes from normal and dark-reared mice. The EGFP and mCherry–CAAX fluorophores (pseudo-coloured yellow) were used to reconstruct the astrocyte territory (red, see Supplementary Methods for further details). o, Average astrocyte territory volumes of V1 L4–5 PALE astrocytes at P7 and P21. Between 18 and 20 cells per condition, 4 mice per condition. p, Average territory volumes of V1 P7 PALE astrocytes from normal and dark-reared mice. Between 18 and 20 cells per condition, 4 mice per condition. q, Average territory volumes of V1 P21 PALE astrocytes from normal and dark-reared mice. Between 18 and 20 cells per condition, 4 mice per condition. One-sided t-test (c, d, o, p, q), one-way ANOVA (j, l, m), ANCOVA (k). Data are means ± s.e.m. Scale bars, 10 μm (a, i, n), 50 μm (b), 100 μm (g).
Extended Data Figure 2 Direct neuronal contact controls astrocyte morphogenesis in vitro.
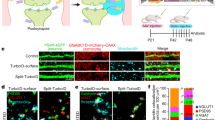
a–d, Neurons induce astrocyte morphogenesis. a, Top, diagram of rat co-culture strategy used to test the cellular and molecular mechanisms of astrocyte–neuron interactions. Astrocytes and neurons are independently isolated from rat cortices (see Supplementary Methods for further details). Astrocytes are transfected after nine days in vitro (DIV9) with plasmid constructs (encoding EGFP, for example) and then trypsinized and plated onto cortical neurons at DIV11. Astrocytes and neurons are co-cultured for 48 h, unless otherwise indicated. Bottom, representative images of astrocytes alone or cultured with neurons (not shown). b, Sholl quantification of EGFP-transfected astrocyte morphology in astrocyte-only cultures at the indicated time points. The length of culture includes only the time of culture on PDL and laminin-coated coverslips (DIV11+), and not the isolation, purification, or transfection stages (DIV0–11, see Supplementary Methods). c, Sholl quantification of EGFP-transfected astrocyte morphology in co-culture with cortical neurons at the indicated time points. The length of culture includes only the time of co-culture with DIV11 rat cortical neurons and not the astrocyte isolation, purification, or transfection stages (DIV0–11). d, Quantification of total intersections obtained from Sholl analysis (b, c) of EGFP-transfected astrocytes grown alone (black) or on cortical neurons (red). b–d, Data represent one experiment with three biological replicates. Similar results were obtained in two independent experiments. n > 15 cells per condition per experiment. e–n, Direct contact of astrocytes with neurons or synapses regulates astrocyte morphogenesis in vitro. e, Representative images of EGFP-labelled astrocytes co-cultured with Cos7 cells, cortical neurons, or alone with neuron-conditioned medium (NCM). f, Representative images of EGFP-labelled astrocytes co-cultured with neurons in the presence of 50 μM or 200 μM MPEP or 1 μM TTX. g, Representative images of EGFP-labelled astrocytes plated on DIV11 methanol (MeOH)-fixed neurons or after neuronal processes were extracted with 8 M urea. h–j, Sholl quantification of astrocyte complexity under different culture conditions from e–g. Data represent one experiment with three biological replicates. Similar results were obtained in at least two independent experiments. n > 25 cells per condition per experiment. k, Representative images of EGFP-transfected astrocytes grown on live or MeOH-fixed Cos7 cells. l, Sholl quantification of EGFP-labelled astrocyte complexity from k. Data represent one experiment with three biological replicates. Similar results were obtained in two independent experiments. n > 20 cells per condition per experiment. m, Representative images of DIV11 cortical (Cx) neurons, processed normally or after neuronal processes were extracted with 8 M urea, stained for the following: DAPI (DNA, blue), phalloidin-conjugated 647 (filamentous actin/neuronal cell bodies and processes, red), gelatin-conjugated Oregon green 488 (ECM proteins, green). n, Representative stimulated emission depletion (STED) super-resolution micrographs of 12-h and 48-h astrocyte–neuron co-cultures immunostained for the pre- and postsynaptic markers Bassoon (red) and Homer-1 (green), respectively. Membrane-targeted EGFP-transfected astrocytes (pseudo-coloured white) extend lamellipodia-like structures towards synapses at the 12-h co-culture time point. By 48-h co-culture, astrocyte processes come in very close proximity to synaptic puncta demarcated by the co-localization of Bassoon and Homer-1. ANCOVA (b, c, h–j, l), one-tailed t-test (d). Data are means ± s.e.m. Scale bars, 10 μm (a, e, f, g, k, m), 5 μm and 500 nm (n).
Extended Data Figure 3 Neuroligins are expressed in astrocytes.
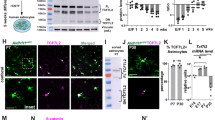
a, Average fragments per kilobase of transcript per million reads (FPKM) of neuroligin mRNAs from RNA sequencing of P7 mouse neurons (black) and astrocytes (red; all data from ref. 13). b, Average FPKM values of cell-type specific transcripts from P7 neurons and astrocytes (Aldh1L1, astrocyte; Eno2, neuron; MOG, oligodendrocyte; all data from ref. 13). c, Average FPKM values of neuroligin mRNAs from 25-year-old human neurons (black), 18–18.5gw (gestational week) fetal astrocytes (red), and 8–63-year-old human mature astrocytes (yellow; all data from ref. 14). d, Average FPKM values of cell-type specific transcripts from specimens listed in c (all data from ref. 14). e, Average FPKM values of neuroligin mRNAs engaged with translating ribosomes from P80 mouse astrocytes (red) and total input (black) using Aldh1L1–creERT2/Ribo-tag mice (all data from ref. 12). f, Average FPKM values of cell-type specific transcripts from specimens listed in e (all data from ref. 12). g, Single optical section confocal images (top) and 3D surface renderings (bottom) of FISH experiments with V1 L1 astrocytes (green) from P21 Aldh1L1–EGFP mice. NL1 (left, red), NL2 (middle, red), and NL3 (right, red) mRNAs are detected inside astrocyte somas and large branches (green, outlined). h, Representative images of RNA FISH experiments on NL1+/+ (WT) and NL1−/− (KO) mice also containing the Aldh1L1–EGFP transgene labelling all astrocytes (green). NL1-specific RNA-FISH probes (red) were used to detect NL1 in both genetic backgrounds. Note: there is substantial fluorescence detection inside (dotted line, arrows) and outside NL1 wild-type astrocytes (top), but no detection of NL1 RNA-FISH signal in NL1 KO tissue sections (bottom). i, RT–PCR amplicons generated using transcript-specific exon-skipping primers. Lanes: P7 rat cortex (Cx), DIV11 rat cortical neuron cultures (Neu.), DIV11 rat cortical astrocyte cultures (Astro.), and DIV11 rat cortical astrocyte cultures without the reverse transcriptase enzyme to control for gDNA contamination (−RT). Specific primer sets were used to detect the following transcripts: NL1, NL2, NL3, neuron-specific enolase (NSE, neuron control), CX3C chemokine receptor 1 R1 (CX3CR1, microglia control), myelin oligodendrocyte glycoprotein (MOG, oligodendrocyte control), glial fibrillary acid protein (GFAP, astrocyte control), aldehyde dehydrogenase 1 family member 1 (Aldh1L1, astrocyte control), and β-actin (loading control). Astrocyte cultures do not contain detectible neuron, microglial or oligodendrocyte contamination. Cortical neuron cultures contain around 10% glial (astrocyte and oligodendrocyte) contamination. j, Western blot images. Lanes: P7 rat cortex (Cx), DIV11 rat cortical neuron cultures (Neu.), DIV11 rat cortical astrocyte cultures (Astro.), and rat cortical astrocyte-conditioned medium (ACM). Specific and validated antibodies were used to detect the following proteins: NL1 intracellular domain (NL1 ICD), NL1 extracellular domain (NL1 ECD), NL2 ICD, NL3 ECD, Hevin (astrocyte-specific secreted protein control), GFAP (intracellular astrocyte specific control), NSE (neuron control), Tuj1 (β3-tubulin, neuron control), tubulin (intracellular protein loading control). Note the presence of two bands for NL3 in the astrocyte sample. Single asterisks denote the expected full length protein. Astrocytes express a lower molecular weight NL3 (double asterisks) to a greater extent, but this lower molecular weight product is not present in the ACM, indicating that it is not a secreted NL3 species. For gel source data, see Supplementary Fig. 1. Data are means ± s.e.m. Scale bars, 10 μm.
Extended Data Figure 4 Neuroligin CAMs control neuron-induced astrocyte morphogenesis.
a–c, Specificity of shRNAs against neuroligin sequences. a, Schematic representation of EGFP-expressing shRNA plasmids used to silence rat NLs in vitro. Sequences of shRNAs used on rat astrocytes against NL1, NL2, and NL3, as well as a scrambled shNL1 sequence used as a control (shCtrl). Rat-specific shRNAs against NL1, NL2, and NL3 were previously verified for efficiency and specificity40,41. The shNL2 sequence matches both rat and mouse NL2 sequences and effectively silences NL2 expression in cells from both species (Extended Data Fig. 6c, d). b, Western blot analyses of cell lysates from HEK293 cells transfected with shCtrl or shrtNL1 with various HA-tagged NL1 constructs. Full length NL1 (HA–NL1) is effectively silenced by shrtNL1, whereas the NL1 shRNA-resistant mutant (HA–NL1-RM) and NL1-SWAP are not silenced by shrtNL1. Note that NL1-SWAP runs smaller than full-length NL1 (previously published in ref. 23). c, Western blot analyses of cell lysates from HEK293 cells transfected with shCtrl or shNL2 with HA-tagged NL2 or HA-tagged NL2-RM. Full length NL2 (HA–NL2) is effectively silenced by shNL2, whereas NL2-RM is not. b, c, Antibodies against HA were used to detect the expression of neuroligins and the lysates were blotted for using an EGFP-specific antibody to verify transfection with shRNA plasmids. d-k, Neuroligins play significant roles in controlling neuron-induced astrocyte morphogenesis. d, Sholl quantification of astrocyte complexity of NL1, NL2, NL3, or NL1/2/3 silenced astrocytes cultured on cortical neurons (not visible, compare with Fig. 2a–e). Data represent one experiment with three biological replicates. Similar results were obtained in three independent experiments. n > 25 cells per condition per experiment. e, Representative images of shNL-transfected astrocytes (green) cultured with MeOH-fixed neurons (not visible). f, Sholl quantification of astrocyte complexity of neuroligin-silenced astrocytes cultured on MeOH-fixed neurons. Data represent one experiment with three biological replicates. Similar results were obtained in two independent experiments. n > 20 cells per condition per experiment. g, Representative images of transfected astrocytes (green) with shCtrl or an shRNA against EphrinA3 (EFNA3) in co-culture with neurons (not visible). EFNA3 is a CAM expressed in astrocytes that regulates astrocyte–synapse interactions in the hippocampus22. h, Sholl quantification of astrocyte complexity in shCtrl and shEFNA3-transfected astrocytes in co-culture with neurons. Data represent one experiment with three biological replicates. Similar results were obtained in two independent experiments. n > 20 cells per condition per experiment. i, Cartoon representation of NL1 domain structure. Neuroligins are type I transmembrane proteins with a large N-terminal extracellular domain (ECD). The NL1-ECD contains a cholinesterase (ChoE)-like domain that interacts with Nrxβs transcellularly. In NL1-SWAP, the ChoE-like domain is swapped for the ChoE sequence. This chimaera is efficiently trafficked to the cell surface, but fails to interact transcellularly with Nrxβs23. j, Representative images of astrocytes overexpressing EGFP (green) and HA-tagged neuroligins (red) in co-culture with neurons (not visible). Neuroligin-overexpressing astrocytes do not show any reductions in neuron-induced astrocyte morphogenesis. k, Sholl quantification of astrocyte complexity when overexpressing HA-tagged neuroligins. Data represent one experiment with three biological replicates. Similar results were obtained in two independent experiments. n > 20 cells per condition per experiment. ANCOVA (d, f, h, k). For gel source data, see Supplementary Fig. 1. Data are means ± s.e.m. Scale bars, 10 μm.
Extended Data Figure 5 Astrocytic NL1 interacts with neuronal neurexins to promote astrocyte morphogenesis in vitro.
a–d, Lentiviral knockdown of rat Nrx1 and Nrx2. a, Domain structures of Nrxα and Nrxβs. Nrxα and Nrxβs are expressed by alternative promoters and share the same LNS6 (laminin-α, neurexin and sex hormone-binding globulin 6), transmembrane and intracellular domains. Nrxαs contain five additional LNS domains and three EGF domains in their extracellular region. Gokce et al. generated a lentiviral construct with tandem shRNAs with recognition sequences localized to the LNS6 and TM domains of mouse Nrxs, such that both Nrxα and Nrxβs are silenced24. b, shRNA sequences that silence mouse Nrxα and Nrxβs24 compared to the corresponding rat sequences. Based on sequence homology, we predicted that this lentiviral construct would silence rat Nrx1α/β and Nrx2α/β, but not Nrx3α/β owing to base-pair mismatches (red). c, RT–PCR amplicons from cortical neurons transduced with a scrambled control lentivirus (shScr) or pan-shNrx lentivirus. Specific primers against Nrx1α, Nrx1β, Nrx2α, Nrx2β, and Nrx3 were used to detect the presence and levels of Nrx transcripts. β-Actin was used as a control. As predicted, pan-shNrx24 effectively diminished expression of rat Nrx1 and Nrx2 but not Nrx3. d, Quantification of Nrx cDNA levels from shScr or pan-shNrx-transduced cortical neurons. c, d, Results are from two independent experiments. e–o, Neuronal neurexins are required to promote astrocyte morphogenesis. e, Diagram of astrocyte–neuron co-culture strategy used to test the requirement for neuronal neurexins in regulating astrocyte complexity. Neurons are transduced on DIV2 with a lentivirus expressing EGFP and tandem shRNAs that silence rat Nrx1 and Nrx2 (green). Astrocytes are transfected with shRNA plasmids, which also encode mCherry, on DIV9 and then plated with neurons on DIV11 for 48 h. f, Representative images of shrtNL1 mCherry-transfected astrocytes (red) in co-culture with shNrx1/2 or shScr lentivirus-transduced neurons (green). g, Sholl quantification of astrocyte complexity in shrtNL1-transfected astrocytes in co-culture with shScr or shNrx1/2-transduced neurons. Data represent one experiment with three biological replicates. Similar results were obtained in two independent experiments. n > 25 cells per condition per experiment. h, Schematic of soluble Fc-tagged Nrxβ ectodomains. Recombinant Fc-only and Nrx1β–Fc proteins were previously described40. For this study, we generated recombinant constructs to express and purify Nrx2β–Fc and Nrx3β–Fc proteins (see Supplementary Methods). i, Coomassie staining shows the molecular mass and purity of the recombinant Fc-tagged proteins. Note that Nrx2β–Fc stains weakly with Coomassie dye, probably because it has fewer positively charged amino acids compared to Nrx1β–Fc and Nrx3β–Fc. j, Schematic of in vitro experiments testing the requirement for neurexin–neuroligin interactions in the control of astrocyte complexity. shRNA EGFP-transfected astrocytes are plated onto MeOH-fixed neurons in the presence or absence of soluble Fc-tagged Nrxβ-ectodomain proteins or control Fc protein. k, Quantification of complexity of astrocytes grown on MeOH-fixed neurons with or without Fc-tagged proteins added at various concentrations. Total Sholl intersections are plotted as a function of increasing Fc-tagged protein concentration (normalized to 0 nM). In this assay, Fc protein concentrations above 25 nM reduced astrocyte morphogenesis nonspecifically; thus, all experiments were conducted at 25 nM concentration. Data represent one experiment with three biological replicates. Similar results were obtained in two independent experiments. n > 25 cells per condition per experiment. l, Representative images of shCtrl or shrtNL1-transfected astrocytes (green) cultured with MeOH-fixed neurons (not visible) in the presence of 25 nM Fc (control) or 25 nM Nrx1β–Fc, Nrx2β–Fc and Nrx3β–Fc (8.3 nM concentration each, total of 25 nM). m, Sholl quantification of astrocytes cultured on MeOH-fixed neurons with or without recombinant Fc proteins (25 nM total concentration). Data represent one experiment with three biological replicates. Similar results were obtained in two independent experiments. n > 25 cells per condition per experiment. n, Sholl quantification of shCtrl-transfected astrocytes cultured on MeOH-fixed neurons supplemented with recombinant Fc proteins (25 nM total concentration). Data represent one experiment with three biological replicates. Similar results were obtained in two independent experiments. n > 25 cells per condition per experiment. o, Sholl quantification of shrtNL1-transfected astrocytes cultured on MeOH-fixed neurons supplemented with recombinant Fc proteins (25 nM total concentration). Data represent one experiment with three biological replicates. Similar results were obtained in two independent experiments. n > 25 cells per condition per experiment. One-tailed t-test (d), ANCOVA (g, k, m–o). For gel source data, see Supplementary Fig. 1. Data are means ± s.e.m. Scale bars, 10 μm.
Extended Data Figure 6 Astrocytic neuroligins control astrocyte morphogenesis in vivo.
a–d, Verification of shRNAs used in vivo. a, Schematic of pLKO.1_hU6 plasmid vector housing the shRNA sequence and EGFP reporter. shRNA constructs targeting both mouse and rat NL1 and NL3 transcripts were obtained from Dharmacon. shNL2 used40,41 effectively silences both mouse and rat NL2. In this plasmid backbone, the shRNA expression is driven from the human U6 minimal promoter. The commercially available plasmids do not encode a fluorescent protein reporter; therefore, we cloned EGFP under the control of a CAG promoter (see Supplementary Methods). b, shRNA sequences used to silence mouse neuroligins for PALE. Because they are a perfect match with the rat sequences, the same shRNAs also target rat neuroligins. shNL1 and shNL3 targeting vectors were verified here. shNL2 was verified previously40,41 (and see Extended Data Fig. 4c). c, d, Western blot analysis of lysates from cultured rat (c) or mouse (d) astrocytes transduced with lentiviruses expressing shCtrl or shNLs. The shNLs effectively silenced the expression of endogenous NL1 (left), NL2 (middle) and NL3 (right) in both rat and mouse astrocytes. β-tubulin levels are shown as a loading control. Blots represent one experiment. Similar results were obtained from three separate experiments. e–h, Astrocytic neuroligins control astrocyte morphogenesis in vivo. e, Data from Fig. 3e, f normalized to P7 shCtrl astrocyte NIV values and replotted to determine how neuroligin silencing affects the growth trajectory of astrocyte NIV. shNL2 and shNL3-transfected astrocytes failed to expand their neuropil infiltration from P7 to P21, whereas shCtrl and shNL1-transfected astrocytes displayed robust (~2.5-fold) growth. Three NIV per cell, 10–20 cells per condition, at least three mice per condition. f, Top, representative images of P21 neuroligin-overexpressing PALE astrocytes from L4–5 V1 cortex. The territories of the neuroligin-overexpressing PALE astrocytes were determined in Imaris Bitplane software with a Matlab Xtension. This method identifies the terminal fluorescent points of each astrocyte and connects these points to generate the territory of each cell (red outline). Bottom, representative NIV (magenta) for neuroligin-overexpressing PALE astrocytes. g, Fold change in average territory volume of NL1- or NL2-overexpressing PALE astrocytes normalized to HA–NL1-SWAP. In the brains from three cohorts of seven PALE NL3-overexpressing mice, we were unable to find NL3-overexpressing astrocytes at P21, indicating that NL3-overexpression starting at P1 is not compatible with astrocyte survival and/or maturation. h, Fold change in average NIV of neuroligin-overexpressing PALE astrocytes normalized to HA–NL1-SWAP. Astrocytes might already occupy the available neuropil space; thus, neuroligin overexpression primarily forces astrocytes to expand. Alternatively, each neuroligin might direct astrocyte processes to certain neuronal elements; thus, neuroligin overexpression drives the astrocyte towards such structures, expanding their domains. g, h, Three NIVs per cell (h only), 14–20 cells per condition, 4 mice per condition. ANCOVA (e), one-way ANOVA (g, h). For gel source data, see Supplementary Fig. 1. Data are means ± s.e.m. Scale bars, 10 μm.
Extended Data Figure 7 NL2 protein is expressed in astrocytes and is required for neuropil infiltration in vivo.
a–c, NL2 protein staining in NL2 PALE HET and NL2 PALE KO astrocytes. a, Representative high-magnification Airyscan confocal images of P21 td-Tomato/Cre+ PALE astrocytes from Nlgn2f/+ and Nlgn2f/f (bottom) mice immunostained using an antibody against the intracellular domain of NL2. Left, single optical section of td-Tomato/Cre+ astrocytes depicting NL2 staining (green) and td-Tomato/Cre+ astrocyte (red). Middle, zoomed-in view of boxed region from left. Right, representative Imaris 3D reconstructed surface renderings of the co-localized NL2 (green) and td-Tomato/Cre+ (red) signals in PALE NL2 HET and NL2 KO astrocytes (NL2 signal outside the td-Tomato/Cre+ cell is not surface rendered). b, Quantification of average NL2 puncta volume inside td-Tomato/Cre+ NIV of PALE NL2 HET and NL2 KO astrocytes. c, Fold change in NL2 puncta volume per td-Tomato/Cre+ volume, normalized to NL2 HET. b, c, Two NIVs per image, three cells per mouse, three mice per genotype. d–f, shNL2 has no off-target effects in astrocytes in vivo. d, Approach to test the specificity of shNL2 in vivo. pCAG–Cre plasmid was injected with or without the shNL2 plasmid into P1 Nlgn2f/+ or Nlgn2f/f pups that also contained two copies of the RTM reporter. Animals were killed at P7 for astrocyte morphological analysis. e, Representative images of P7 td-Tomato/Cre+ astrocytes (red) or co-expressing shNL2 (green and red, appears as yellow) from PALE NL2 HET and NL2 KO mice. Representative NIVs are shown below each astrocyte (magenta). f, Fold change in average NIV of P7 td-Tomato/Cre+ PALE NL2 HET and NL2 KO astrocytes with or without shNL2, normalized to td-Tomato/Cre+ PALE NL2 HET. Three NIVs per cell, 16 cells per condition, at least 2 mice per genotype. One-tailed t-test (b, c), one-way ANOVA (f). Data are means ± s.e.m. Scale bars, 10 μm.
Extended Data Figure 8 Controls determining the specificity of synaptic immunohistochemistry.
a–f, To test whether synaptic immunostaining reflects true protein localization or random signal, a rotation analysis was performed40. In this test, the presynaptic channel was rotated 90° with respect to the other two channels and re-merged to create a rotated image. Synapse density quantifications were re-run using the rotated image compared to the standard image. a, c, e, Representative images of standard and rotated images stained for excitatory VGluT1 (red)/PSD95 (green) (a), excitatory VGluT2 (red)/PSD95 (green) (c), and inhibitory VGAT (red)/Gephyrin (green) (e) synaptic staining. Images depict the territory boundary of td-Tomato/Cre+ (blue) astrocytes and td-Tomato/Cre− wild-type (unlabelled) astrocytes. b, d, f, Quantification of co-localized synaptic puncta density of standard and rotated images from excitatory VGluT1 (red)/PSD95 (green) (a), excitatory VGluT2 (red)/PSD95 (green) (c), and inhibitory VGAT (red)/Gephyrin (green) (e) synaptic staining. Standard images (left four bars) compared to the rotated images (right two bars). One-way ANOVA (b, d, f). Data are means ± s.e.m. Scale bar, 2 μm.
Extended Data Figure 9 Generation and verification of NL2 conditional knockout mice.
a, b, Generating astrocyte-specific conditional NL2 knockout mice using the tamoxifen-inducible GLAST-CreERT2 transgenic mouse line. a, Top, NL2 cHET and NL2 cKO mice each contain a single copy of the GLAST-CreERT2 transgene (tg/0) and the RTM reporter (f/+). (Bottom) Experimental timeline of tamoxifen-induced recombination of Cre-dependent loci (floxed Nlgn2 and RTM) in combination with the GLAST-CreERT2 transgene. Cre recombination was activated by administering 0.6 mg tamoxifen twice at P10 and P11 via intraperitoneal injection to achieve widespread deletion of NL2 in astrocytes. b, Representative sagittal tile-scan confocal images of the forebrains of P21 NL2 cHET and NL2 cKO mice. td-Tomato/Cre+ cells (red) are visible in all regions of the forebrain. A higher magnification image of the V1 cortex is shown in the dotted box. More than 50% of cortical astrocytes in V1 are td-Tomato/Cre+ (see Extended Data Fig. 10 for quantification and details). A, P, D, V, are anterior, posterior, dorsal, ventral, respectively. c, d, Work-flow for isolating td-Tomato/Cre+ cells by FACS. c, Experimental flow chart demonstrating the isolation and analysis of td-Tomato/Cre+ cells from the cortices of NL2 cHET and NL2 cKO animals via FACS. Cortices were microdissected and papain digested, and a single-cell suspension was generated. The cell suspension was passed through a BD FACS sorter to sequester and capture td-Tomato/Cre+ cells. RNA was isolated from the td-Tomato+ cells, reverse transcribed to cDNA, then quantified by quantitative PCR. In separate experiments, genomic DNA (gDNA) was isolated from td-Tomato/Cre+ FACS samples to verify Cre-mediated recombination by PCR. d, Representative contour plots of NL2 cHET and cKO FACS analysis. Total single-cell suspension input was sorted and gated for td-Tomato fluorescence intensity (x-axis). The high td-Tomato+ cluster (red, boxed) was sorted into fresh tubes for RNA or gDNA extraction. e–i, Verification of the loss of NL2 in NL2 cKO astrocytes. e, qPCR quantification of gene transcript levels of GFAP, NL1, NL2 and NL3 in td-Tomato/Cre+ cells isolated by FACS from NL2 cHET (black) and cKO (red) mice. Expression is shown as fold change normalized to NL2 cHET values. Three or four mice per genotype. f, Schematic for genotyping the wild-type non-recombined alleles of NL2. The NL2 wild-type and floxed alleles have seven exons (labelled 1–7, yellow boxes). The NL2 floxed allele contains two LoxP sites (green triangles) that flank exons 3–5. To verify the non-Cre-mediated recombination of the NL2 locus by PCR, a reverse primer that base pairs with a region inside exon 3 is combined with a common forward primer that pairs with the DNA inside intron 2. The expected band sizes produced by the NL2 wild-type and floxed allele are 413 bp and about 500 bp, respectively. g, Representative non-recombined PCR gel with the following samples: water (control), tail samples from Nlgn2+/+, Nlgn2f/+, and Nlgn2f/f mice, and td-Tomato/Cre+ FACS samples from NL2 cHET and NL2 cKO brains. The wild-type and floxed alleles are depicted (arrows). h, Schematic for genotyping the knockout recombined alleles of NL2. To verify the Cre-mediated recombination of the NL2 locus by PCR, a reverse primer that base pairs with a region downstream of the second LoxP site inside intron 5 is combined with the common forward primer. The Cre-mediated recombined Nlgn2 locus will produce a roughly 250-bp band. The non-recombined alleles (wild-type and non-recombined floxed alleles) will not produce a band small enough for detection. i, Representative Cre-mediated recombined PCR gel with the following samples: water (control), tail samples from Nlgn2+/+, Nlgn2f/+, and Nlgn2f/f mice, and td-Tomato+ FACS samples from NL2 cHET and NL2 cKO brains. For gel source data, see Supplementary Fig. 1. One-tailed t-test (e). Data are means ± s.e.m. Scale bar, 1 mm.
Extended Data Figure 10 Conditional deletion of NL2 in astrocytes does not alter cell number or distribution.
a–c, The number of astrocytes and td-Tomato/Cre+ cells are similar between NL2 cHET and NL2 cKO mice. a, Representative tile scan images of V1 cortex from NL2 cHET and NL2 cKO mice stained for GFAP (astrocyte marker, green), td-Tomato/Cre+ cells (red), and DAPI (blue). High magnification of selected regions is shown below each image (dotted box). More than 99% of td-Tomato/Cre+ cells stain positive for GFAP. b, Quantification of the number and distribution of V1 cortex GFAP+ astrocytes in NL2 cHET and NL2 cKO mice. The number of GFAP+ cells is plotted as a function of cortical layer (L1–6) in which they reside. The percentage of GFAP/td-Tomato double-positive cells in NL2 cHET and NL2 cKO mice does not differ significantly (NL2 cHET = 58.7 ± 4.0%; NL2 cKO = 53.9 ± 4.4%, P = 0.31483). Three or four images per mouse, three mice per genotype. c, Quantification of the number and distribution of td-Tomato/Cre+ cells from NL2 cHET and NL2 cKO mice in V1 cortex. The number of td-Tomato/Cre+ cells is plotted as a function of cortical layer. Three or four images per mouse, three mice per genotype. d, Representative tile scan images of V1 cortex from NL2 cHET and NL2 cKO mice stained for NeuN (neuronal marker, green) and td-Tomato/Cre+ cells (red). High magnification of selected regions is shown in dotted boxes. The percentage of NeuN/td-Tomato double-positive cells was extremely low, confirming that Cre recombination is specific to astrocytes in V1 cortex (number of NeuN/td-Tomato double-positive cells by genotype was: NL2 cHET, 36 double-positive cells out of 13,903 NeuN+ cells counted (0.26%); NL2 cKO, 36 double-positive cells out of 13,504 NeuN+ cells counted (0.25%). e, Quantification of the number and distribution of NeuN+ neurons from NL2 cHET and NL2 cKO mice. The number of NeuN+ cells is plotted as function of the cortical layer in which they reside. Three or four images per mouse, three mice per genotype. f, Top left, NL2 cHET and NL2 cKO mice used to test excitatory and inhibitory synaptic function. Bottom left, experimental timeline of tamoxifen-induced recombination of Cre-dependent loci (floxed Nlgn2 and RTM) in combination with the GLAST-CreERT2 transgene. Cre was activated by administering 0.6 mg tamoxifen at P10 and P11 via intraperitoneal injection to achieve widespread deletion of NL2 in astrocytes. Right, schematic representation of electrophysiological recording experiments performed on L5 V1 pyramidal neurons from NL2 cHET and NL2 cKO mice. L5 pyramidal neurons receive excitatory and inhibitory synaptic connections from all cortical layers that contain td-Tomato/Cre+ (red) and td-Tomato/Cre− (grey) astrocytes. mEPSCs and mIPSCs were recorded from L5 pyramidal neurons using acute slices from P21 NL2 cHET and NL2 cKO mice containing the V1 cortex. ANCOVA (b, c, e). Data are means ± s.e.m. Scale bars, 100 μm.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary References and Supplementary Figure 1, the uncropped scans. (PDF 10967 kb)
Rights and permissions
About this article
Cite this article
Stogsdill, J., Ramirez, J., Liu, D. et al. Astrocytic neuroligins control astrocyte morphogenesis and synaptogenesis. Nature 551, 192–197 (2017). https://doi.org/10.1038/nature24638
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24638
This article is cited by
-
Human neuronal maturation comes of age: cellular mechanisms and species differences
Nature Reviews Neuroscience (2024)
-
Astrocytic β-catenin signaling via TCF7L2 regulates synapse development and social behavior
Molecular Psychiatry (2024)
-
Synaptic input and Ca2+ activity in zebrafish oligodendrocyte precursor cells contribute to myelin sheath formation
Nature Neuroscience (2024)
-
Morphological diversification and functional maturation of human astrocytes in glia-enriched cortical organoid transplanted in mouse brain
Nature Biotechnology (2024)
-
The Roles of Non-coding RNA Targeting Astrocytes in Cerebral Ischemia
Molecular Neurobiology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.