Abstract
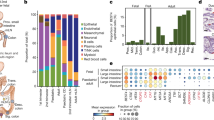
Intestinal epithelial cells absorb nutrients, respond to microbes, function as a barrier and help to coordinate immune responses. Here we report profiling of 53,193 individual epithelial cells from the small intestine and organoids of mice, which enabled the identification and characterization of previously unknown subtypes of intestinal epithelial cell and their gene signatures. We found unexpected diversity in hormone-secreting enteroendocrine cells and constructed the taxonomy of newly identified subtypes, and distinguished between two subtypes of tuft cell, one of which expresses the epithelial cytokine Tslp and the pan-immune marker CD45, which was not previously associated with non-haematopoietic cells. We also characterized the ways in which cell-intrinsic states and the proportions of different cell types respond to bacterial and helminth infections: Salmonella infection caused an increase in the abundance of Paneth cells and enterocytes, and broad activation of an antimicrobial program; Heligmosomoides polygyrus caused an increase in the abundance of goblet and tuft cells. Our survey highlights previously unidentified markers and programs, associates sensory molecules with cell types, and uncovers principles of gut homeostasis and response to pathogens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007)
von Moltke, J., Ji, M., Liang, H. E. & Locksley, R. M. Tuft-cell-derived IL-25 regulates an intestinal ILC2-epithelial response circuit. Nature 529, 221–225 (2016)
Barriga, F. M. et al. Mex3a marks a slowly dividing subpopulation of Lgr5+ intestinal stem cells. Cell Stem Cell 20, 801–816.e7 (2017)
Basak, O. et al. Induced quiescence of Lgr5+ stem cells in intestinal organoids enables differentiation of hormone-producing enteroendocrine cells. Cell Stem Cell 20, 177–190.e4 (2017)
Grün, D. et al. Single-cell messenger RNA sequencing reveals rare intestinal cell types. Nature 525, 251–255 (2015)
Yan, K. S. et al. Non-equivalence of Wnt and R-spondin ligands during Lgr5+ intestinal stem-cell self-renewal. Nature 545, 238–242 (2017)
Yan, K. S. et al. Intestinal enteroendocrine lineage cells possess homeostatic and injury-inducible stem cell activity. Cell Stem Cell 21, 78–90.e6 (2017)
Zheng, G. X. et al. Haplotyping germline and cancer genomes with high-throughput linked-read sequencing. Nat. Biotechnol. 34, 303–311 (2016)
Rosvall, M. & Bergstrom, C. T. Maps of random walks on complex networks reveal community structure. Proc. Natl Acad. Sci. USA 105, 1118–1123 (2008)
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323.e30 (2016)
Amir, E.-a. D. et al. viSNE enables visualization of high dimensional single-cell data and reveals phenotypic heterogeneity of leukemia. Nat. Biotechnol. 31, 545–552 (2013)
Kowalczyk, M. S. et al. Single-cell RNA-seq reveals changes in cell cycle and differentiation programs upon aging of hematopoietic stem cells. Genome Res. 25, 1860–1872 (2015)
Garabedian, E. M., Roberts, L. J., McNevin, M. S. & Gordon, J. I. Examining the role of Paneth cells in the small intestine by lineage ablation in transgenic mice. J. Biol. Chem. 272, 23729–23740 (1997)
Gribble, F. M. & Reimann, F. Enteroendocrine cells: chemosensors in the intestinal epithelium. Annu. Rev. Physiol. 78, 277–299 (2016)
Howitt, M. R. et al. Tuft cells, taste-chemosensory cells, orchestrate parasite type 2 immunity in the gut. Science 351, 1329–1333 (2016)
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protocols 9, 171–181 (2014)
van der Meer-van Kraaij, C. et al. Dietary modulation and structure prediction of rat mucosal pentraxin (Mptx) protein and loss of function in humans. Genes Nutr. 2, 275–285 (2007)
Du Clos, T. W. Pentraxins: structure, function, and role in inflammation. ISRN Inflamm. 2013, 379040 (2013)
Katz, J. P. et al. The zinc-finger transcription factor Klf4 is required for terminal differentiation of goblet cells in the colon. Development 129, 2619–2628 (2002)
Duboc, H., Taché, Y. & Hofmann, A. F. The bile acid TGR5 membrane receptor: from basic research to clinical application. Dig. Liver Dis. 46, 302–312 (2014)
Overton, H. A., Fyfe, M. C. & Reynet, C. GPR119, a novel G protein-coupled receptor target for the treatment of type 2 diabetes and obesity. Br. J. Pharmacol. 153 (Suppl. 1), S76–S81 (2008)
Coifman, R. R. et al. Geometric diffusions as a tool for harmonic analysis and structure definition of data: diffusion maps. Proc. Natl Acad. Sci. USA 102, 7426–7431 (2005)
Basak, O. et al. Mapping early fate determination in Lgr5+ crypt stem cells using a novel Ki67-RFP allele. EMBO J. 33, 2057–2068 (2014)
Beuling, E. et al. GATA factors regulate proliferation, differentiation, and gene expression in small intestine of mature mice. Gastroenterology 140, 1219–1229.e2 (2011)
Furness, J. B., Rivera, L. R., Cho, H. J., Bravo, D. M. & Callaghan, B. The gut as a sensory organ. Nat. Rev. Gastroenterol. Hepatol. 10, 729–740 (2013)
Worthington, J. J., Reimann, F. & Gribble, F. M. Enteroendocrine cells-sensory sentinels of the intestinal environment and orchestrators of mucosal immunity. Mucosal Immunol. (2017)
Habib, A. M., Richards, P., Rogers, G. J., Reimann, F. & Gribble, F. M. Co-localisation and secretion of glucagon-like peptide 1 and peptide YY from primary cultured human L cells. Diabetologia 56, 1413–1416 (2013)
Gershon, M. D. & Tack, J. The serotonin signaling system: from basic understanding to drug development for functional GI disorders. Gastroenterology 132, 397–414 (2007)
Klok, M. D., Jakobsdottir, S. & Drent, M. L. The role of leptin and ghrelin in the regulation of food intake and body weight in humans: a review. Obes. Rev. 8, 21–34 (2007)
Karra, E., Chandarana, K. & Batterham, R. L. The role of peptide YY in appetite regulation and obesity. J. Physiol. 587, 19–25 (2009)
Gerbe, F. & Jay, P. Intestinal tuft cells: epithelial sentinels linking luminal cues to the immune system. Mucosal Immunol. 9, 1353–1359 (2016)
Gerbe, F. et al. Intestinal epithelial tuft cells initiate type 2 mucosal immunity to helminth parasites. Nature 529, 226–230 (2016)
Bezençon, C. et al. Murine intestinal cells expressing Trpm5 are mostly brush cells and express markers of neuronal and inflammatory cells. J. Comp. Neurol. 509, 514–525 (2008)
Biton, M. et al. Epithelial microRNAs regulate gut mucosal immunity via epithelium–T cell crosstalk. Nat. Immunol. 12, 239–246 (2011)
de Lau, W. et al. Peyer’s patch M cells derived from Lgr5+ stem cells require SpiB and are induced by RankL in cultured “miniguts”. Mol. Cell. Biol. 32, 3639–3647 (2012)
Mabbott, N. A., Donaldson, D. S., Ohno, H., Williams, I. R. & Mahajan, A. Microfold (M) cells: important immunosurveillance posts in the intestinal epithelium. Mucosal Immunol. 6, 666–677 (2013)
Terahara, K. et al. Comprehensive gene expression profiling of Peyer’s patch M cells, villous M-like cells, and intestinal epithelial cells. J. Immunol. 180, 7840–7846 (2008)
Peterson, L. W. & Artis, D. Intestinal epithelial cells: regulators of barrier function and immune homeostasis. Nat. Rev. Immunol. 14, 141–153 (2014)
Loonen, L. M. et al. REG3γ-deficient mice have altered mucus distribution and increased mucosal inflammatory responses to the microbiota and enteric pathogens in the ileum. Mucosal Immunol. 7, 939–947 (2014)
Eckhardt, E. R. et al. Intestinal epithelial serum amyloid A modulates bacterial growth in vitro and pro-inflammatory responses in mouse experimental colitis. BMC Gastroenterol. 10, 133 (2010)
Martinez Rodriguez, N. R. et al. Expansion of Paneth cell population in response to enteric Salmonella enterica serovar Typhimurium infection. Infect. Immun. 80, 266–275 (2012)
Artis, D. et al. RELMbeta/FIZZ2 is a goblet cell-specific immune-effector molecule in the gastrointestinal tract. Proc. Natl Acad. Sci. USA 101, 13596–13600 (2004)
Vassen, L., Okayama, T. & Möröy, T. Gfi1b:green fluorescent protein knock-in mice reveal a dynamic expression pattern of Gfi1b during hematopoiesis that is largely complementary to Gfi1. Blood 109, 2356–2364 (2007)
Su, L. et al. Coinfection with an intestinal helminth impairs host innate immunity against Salmonella enterica serovar Typhimurium and exacerbates intestinal inflammation in mice. Infect. Immun. 82, 3855–3866 (2014)
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012)
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007)
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012)
Brennecke, P. et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods 10, 1093–1095 (2013)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011)
Buja, A. & Eyuboglu, N. Remarks on parallel analysis. Multivariate Behav. Res. 27, 509–540 (1992)
van der Maaten, L. Accelerating t-SNE using tree-based algorithms. J. Mach. Learn. Res. 15, 3221–3245 (2014)
van der Maaten, L. & Hinton, G. Visualizing Data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008)
Zeisel, A. et al. Brain structure. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015)
Haghverdi, L., Buettner, F. & Theis, F. J. Diffusion maps for high-dimensional single-cell analysis of differentiation data. Bioinformatics 31, 2989–2998 (2015)
Ester, M ., Kriegel, H.-P ., Sander, J. & Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise. In Proc. 2nd Int. Conf. Knowledge, Discovery and Data Mining (KDD-96) (eds Simoudis, E . et al.) 226–231 (AAAI, 1996)
Levine, J. H. et al. Data-driven phenotypic dissection of AML reveals progenitor-like cells that correlate with prognosis. Cell 162, 184–197 (2015)
Rodriguez, A. & Laio, A. Machine learning. Clustering by fast search and find of density peaks. Science 344, 1492–1496 (2014)
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015)
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995)
Zhang, H.-M. et al. AnimalTFDB: a comprehensive animal transcription factor database. Nucleic Acids Res. 40, D144–D149 (2012)
Ng, A., Eisenberg, J. M. & Heath, R. Human leucine-rich repeat proteins: a genome-wide bioinformatic categorization and functional analysis in innate immunity. Proc. Natl Acad. Sci. 108, 4631–4638 (2011)
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 11, R14 (2010)
Ichimura, A., Hirasawa, A., Hara, T. & Tsujimoto, G. Free fatty acid receptors act as nutrient sensors to regulate energy homeostasis. Prostaglandins Other Lipid Mediat. 89, 82–88 (2009)
Rubin, D. B. The Bayesian bootstrap. Ann. Stat. 9, 130–134 (1981)
Kobayashi, A. et al. Identification of novel genes selectively expressed in the follicle-associated epithelium from the meta-analysis of transcriptomics data from multiple mouse cell and tissue populations. DNA Res. 19, 407–422 (2012)
Datta, R. et al. Identification of novel genes in intestinal tissue that are regulated after infection with an intestinal nematode parasite. Infect. Immun. 73, 4025–4033 (2005)
Acknowledgements
We thank L. Gaffney for help with figure preparation, the Broad Flow Cytometry Facility (P. Rogers, S. Saldi and C. Otis), C. Hafemeister and R. Satija for use of the ‘How Many Cells’ tool, S. Riesenfeld and A. Dixit for statistical advice, and T. Tickle for help with the Single Cell Portal. This study was supported by the Klarman Cell Observatory at the Broad Institute, NIH RC2DK114784 (A.R. and R.J.X.), HHMI (A.R.), Food Allergy Science Initiative (FASI) at the Broad Institute (A.R. and R.J.X.), and a Broadnext10 award (A.R. and R.J.X.). M.B. is supported by a postdoctoral fellowship from the Human Frontiers Science Program (HFSP). R.J.X. is supported by NIH DK43351, DK097485 and the Helmsley Charitable Trust.
Author information
Authors and Affiliations
Contributions
A.L.H., M.B. and N.R. contributed equally to this study; M.B., R.J.X. and A.R. co-conceived the study; M.B., N.R., A.L.H., R.J.X. and A.R. designed experiments and interpreted the results; N.R. and M.B. carried out all experiments; G.B., T.M.D., M.R.H., S.B., D.D., M.Z. and R.R. assisted with experiments; A.L.H. designed and performed computational analysis with assistance from R.H.H., K.S., C.S., Y.K., I.T. and A.R.; M.R.H. and W.S.G. assisted with tuft and follicle-associated epithelium experiments; M.Z. and H.N.S. assisted with pathogen infections; S.B. and O.Y. assisted with epithelial cell sorting; D.D. and O.R.-R. assisted with scRNA-seq; and A.L.H., M.B., N.R., R.J.X. and A.R. wrote the manuscript, with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
A.R. is a member of the scientific advisory board of ThermoFisher, Syros Pharmaceuticals and Driver Group. R.J.X. is a consultant at Novartis, Janssen and Celgene. A.H., M.B., N.R., R.H., K.S., C.S., O.R., R.X. and A.R. are co-inventors on a provisional patent application filed by the Broad Institute relating to this manuscript.
Additional information
Reviewer Information Nature thanks L. Vermeulen and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Identifying intestinal epithelial cell types in scRNA-seq data by unsupervised clustering.
Related to Fig. 1. a, b, Quality metrics for scRNA-seq data. Shown are distributions of the number of reads per cell (left), the number of genes detected with non-zero transcript counts per cell (centre) and the fraction of reads mapping to the mm10 mouse transcriptome per cell (right) in the droplet-based 3′ scRNA-seq data (a) and the plate-based full-length scRNA-Seq data (b). c–f, Agreement across batches. c, Contribution of batches to each cluster. Each pie chart shows the batch composition (colour-coded legend) of each detected cluster (labelling and number of cells are marked above each pie chart) in the droplet-based 3′ scRNA-seq dataset (n = 6 mice). All ten replicates contribute to all clusters, and no major batch effect is observed. d, Cell type proportions across batches. Shown is the proportion of detected cells in each major cell type in the droplet-based 3′ scRNA-seq dataset in each of ten batches (points; n = 6 mice). Grey bar, mean; error bars, s.e.m. e, Agreement in expression profiles across mice. Box and whisker plot shows the Pearson correlation coefficients in average expression profiles (average log2(TPM + 1)) for cells in each cluster, across all pairs of mice. Black bar, median value; box edges, 25th and 75th percentiles; whiskers, a further 1.5 times the interquartile range. Clusters with additional subtypes (such as tuft and EEC cells) show more variation, as expected. f, Scatter plots comparing the average log2(TPM + 1) gene expression values between two scRNA-seq experiments from the droplet-based 3′ scRNA-seq dataset (top), between two scRNA-seq experiments from the plate-based full-length scRNA-seq dataset (centre), or between the average of a plate-based full-length scRNA-seq and a population control (bottom). Pearson correlation is marked top left. g, Additional quality control metrics and cluster annotation on the basis of the expression of known cell type markers. t-SNE visualization of 7,216 single cells is shown, where individual points correspond to single cells. Cells are coloured by, from top left to bottom right, their assignment to clusters using a k-nearest-neighbour graph-based algorithm (Methods; legend shows the cluster type) (identical to Fig. 1b), mean expression (log2(TPM + 1)) of several known marker genes for a particular cell type or state (indicated above each plot), the mouse from which they originate (see legend), the number of reads per cell, the number of genes detected per cell, or the number of transcripts as measured by UMIs per cell.
Extended Data Figure 2 Identification and characterization of intestinal epithelial cell types in plate-based full-length scRNA-seq data by unsupervised clustering.
Related to Fig. 1. a, Quality control metrics and cluster annotation on the basis of the expression of known cell type markers. t-SNE visualization of 1,522 single cells (n = 8 mice) is shown, where individual points correspond to single cells. Cells are coloured by, from top left to bottom right, their assignment to clusters, mean expression (log2(TPM + 1)) of several known marker genes for a particular cell type or state (indicated above each plot; same as in Extended Data Fig. 1g), the mouse from which they originate (see legend) and its genotype, the FACS gate used to sort them (see legend), the number of reads per cell, or the number of genes detected per cell. b, Cell-type-specific signatures. Heatmap shows the relative expression level (row-wise Z scores) of genes (rows) in cell-type-specific signatures (same genes as in Fig. 1c, with the exception of enterocyte markers), across the individual post-mitotic intestinal epithelial cells (columns) in the full-length scRNA-seq data. Colour code marks the cell type and their associated signatures. c, Mptx2 is a novel Paneth cell marker. t-SNE of the cells from the droplet-based 3′ scRNA-seq (left; as in Fig. 1b) and plate-based full-length scRNA-seq (right; as in a) datasets is shown, coloured by expression (log2(TPM + 1)) of the mucosal pentraxin Mptx2. d, Cell-type-enriched GPCRs. Heatmap shows the relative expression (row-wise Z scores) of genes encoding GPCRs (rows) that are significantly (FDR < 0.001; Mann–Whitney U test; Methods) upregulated or downregulated in the cells (columns) of a given cell type (top; colour coded as in a) compared to all other cells, in the plate-based full-length scRNA-seq data. e, Cell-type-specific leucine-rich repeat proteins (LRRs). Heatmap depicts the mean relative expression (column-wise Z score of mean log2(TPM + 1) values) of genes (columns) encoding leucine-rich repeat proteins that are significantly (FDR < 0.001; Mann–Whitney U test) upregulated or downregulated in a given cell type (rows) compared to all other cells, in the plate-based full-length scRNA-seq data. f, Cell-type transcription factors and GPCRs. Average relative expression (Z score of mean log2(TPM + 1); colour scale) of the top ten transcription factors (left) and GPCRs (right) (columns) enriched in each cell type (rows).
Extended Data Figure 3 Regional variation in Paneth cell subtypes and stem cell markers.
a, Paneth cell subsets. t-SNE of 10,396 single cells (points) was obtained using a large cell-enriched protocol (Methods), coloured by cluster annotation (n = 2 mice). b, Paneth cell subset markers. Shown is the expression (row-wise Z score; colour scale) of genes specific (FDR < 0.05; Mann–Whitney U test; log2(fold change) > 0.5) to each of the two Paneth cell subsets (average of 724.5 cells per subtype, down-sampled to 500 for visualization) shown in a. c, Two Paneth subsets reflect regional diversity. Shown is the expression of the same genes (rows) as in b in Paneth cells from each of three small intestinal regions (average of 176.3 cells per region; columns; Fig. 2a). 11 of 11 Paneth-1 markers are enriched in the ileal Paneth cells, whereas 7 of 10 Paneth-2 markers are enriched in duodenal or jejunal Paneth cells (FDR < 0.05; Mann–Whitney U test). d, Validation of regional enterocyte markers. Shown is smFISH of Lct (red) and Fabp6 (white) in the duodenum (proximal; left) and ileum (distal; right). Dotted line, boundary between crypt and villi; green and yellow arrows, proximal and distal enterocytes, respectively; scale bars, 50 μm. e, Regional variation of intestinal stem cells. Expression (row-wise Z score) of genes specific to stem cells from each intestinal region (FDR < 0.05; Mann–Whitney U test; log2(fold change) > 0.5). On average, 1,226.3 cells were obtained from each of the three regions, down-sampled to 500 for visualization (columns).
Extended Data Figure 4 Differentiation from stem cells to mature enterocytes.
a-d. Diffusion-map embedding of 5,282 cells (points) progressing through stages of enterocyte differentiation (Methods). a, b, Cells are coloured by their cluster assignment (Fig. 1b). Diffusion components 1 and 3 (DC-1 and DC-3) are associated with the transition from stem cells to progenitors (a), whereas DC-2 distinguishes between proximal and distal enterocyte fate commitment (b). c, d, Cells are coloured by the expression (log2(TPM + 1)) of known and newly identified transcription factors associated with stages of differentiation (c), or with proximal or distal enterocyte differentiation (d). e, Transcription factors that are differentially expressed between proximal and distal cell fate. Heatmap shows the mean expression level of 44 transcription factors differentially expressed between the proximal and distal (rows) enterocyte clusters of Fig. 1b (FDR < 0.05; Mann–Whitney U test). f, Newly identified regional stem cell markers (Extended Data Fig. 3e) identify distinct populations in diffusion-map space. Shown are close-ups of the stem-cell region in diffusion space (b, inset square), coloured by expression level (log2(TPM + 1)) of pan-stem cell marker Lgr5 (left), proximal stem cell marker Gkn3 (centre) or distal stem cell marker Bex1 (right). Dashed line helps to visualize separation of stem cells on the basis of region-specific markers.
Extended Data Figure 5 Heterogeneity within EEC cells.
Related to Fig. 3. a, EEC subset discovery and regional location. Shown is the t-SNE of the 533 EEC cells identified from the droplet-based datasets for whole small intestine (SI) and regional samples (colour legend; n = 8 mice; Methods). b, Agreement in hormone detection rates between droplet-based 3′ and full-length scRNA-seq. Scatter plot shows the detection rate (fraction of cells with non-zero expression of a given transcript) for a set of known EEC hormones, transcription factors and marker genes (see legend) in EEC cells from the full-length dataset, and from the droplet-based 3′ dataset. Linear fit (dashed line) and 95% confidence interval (shaded) are also shown. c, Expression of key genes across subset clusters. t-SNE plot shows cells coloured by their assignment to the 12 clusters (top left; identical to Fig. 3a) or by the expression (log2(TPM + 1)) of markers of immature EEC cells (Neurog3), genes encoding gut hormones (Sct, Sst, Cck, Gcg, Ghrl, GIP, Nts, PYY) or markers of enterochromaffin cells (Tac1, Reg4). d, Co-expression of gastrointestinal hormones by individual cells. Left, heatmap shows the expression of canonical gut hormone genes (rows) in each of 533 individual EEC cells (columns), coloured on the basis of their assignment to the clusters in Fig. 3a (top). Right, heatmap shows for each cluster (columns) the percentage of cells (inset text) in which the transcript for each hormone (rows) is detected.
Extended Data Figure 6 Classification and specificity of EEC subsets.
Related to Fig. 3. a, b, Relationships between EEC subsets. a, Dendrogram shows the relationship between EEC clusters as defined by hierarchical clustering of mean expression profiles of all of the cells in a subset (Methods). Estimates for the significance of each split are derived from 100,000 bootstrap iterations using the R package pvclust (■P < 0.1, *P < 0.05, **P < 0.01, P < 0.001; χ2 test). b, Heatmap shows cell–cell similarities (Pearson’s r) between the 11 significant principal component scores (P < 0.05; Methods) across the 533 EEC cells (rows, columns). Rows and columns are ordered using cluster labels obtained using unsupervised clustering (Methods). c. Subset specificity of gut hormones and related genes. Scatter plot shows each the specificity of each gene to its marked cell subset (defined as the proportion of cells not in a given subset that do not express a given gene) and its sensitivity in that subset (defined as the fraction of cells of a given type that do express the gene (Methods). Subsets are colour coded as in the legend. Genes are assigned to the subset where they are most highly expressed on average. Genes were chosen on the basis of their known annotation as gut hormones (Cck, Gal, Gcg, Ghrl, GIP, Iapp, Nucb2, Nts, Pyy, Sct, Sst), enterochromaffin markers (Tph1, Tac1) and canonical EEC markers (Chga, Chgb). d, GPCRs enriched in different EEC subtypes. Heatmap shows the expression levels (row-wise Z score) averaged across the cells in each of the EEC subtypes (columns) of 11 GPCR-encoding genes (rows) that are differentially expressed (FDR < 0.25; Mann–Whitney U test) in one of the EEC subtypes. The free fatty acid receptors (Ffar) 1 and 4 show specific expression patterns: Ffar1 is highest in SIN cells and is also expressed by the Cck-expressing subsets previously termed I cells (SIL-P, SILA and SIK-P), whereas Ffar4 is highest in the GIP-expressing subsets (SIK and SIK-P). These receptors are known to induce the expression of GIP and Gcg to maintain energy homeostasis64. Ffar2 was expressed by some progenitors and by enterochromaffin cells, but absent from GIP-expressing cells, whereas the oleoylethanolamide receptor Gpr119, which is important for food intake and glucose homeostasis21, is most highly expressed in SILA cells.
Extended Data Figure 7 Characterization of tuft cell heterogeneity and identification of Tslp and the haematopoietic lineage marker Ptprc (CD45) in a subset of tuft cells.
Related to Fig. 4. a, Tuft-1 and tuft-2 cells. Shown is t-SNE visualization of 102 tuft cells (points; n = 8 mice) from the plate-based full-length scRNA-seq dataset (Extended Data Fig. 2a), labelled by their subclustering into tuft-1 (orange) and tuft-2 (brown) subtypes. b, Gene signatures for tuft-1 and tuft-2 cells. Heatmap shows the relative expression (row-wise Z scores) of the tuft-1 and tuft-2 marker genes (rows; orange and brown, respectively) across single cells from the plate-based dataset (columns) assigned to tuft-1 and tuft-2 cell clusters (orange and brown, respectively). The top 25 genes are shown for each subtype (all FDR < 0.01 and log2(fold change) > 0.1 in both plate- and droplet-based datasets). c, Tuft-2 signature genes are enriched in immune functions. Shown are the significantly enriched (Methods; FDR < 0.1; −log10(Q value)) gene ontology terms in the gene signature for the tuft-2 subset. d, Expression of neuron- and inflammation-related genes in tuft-1 and tuft-2 subsets, respectively. Plot shows for each gene (y axis) its differential expression (x axis) between Tuft-1 and Tuft-2 cells. Bar indicates Bayesian bootstrap65 estimates of log2(fold change); hinges and whiskers indicate 25% and 95% confidence intervals, respectively. e, Il33 is not detected in tuft cells. Distribution of expression of Il33 in cell subsets in full-length scRNA-seq. (*FDR < 0.1; Mann–Whitney U test). f, g, Tuft-2 cells are enriched for Tslp. f, Combined smFISH and immunofluorescence of Tslp (green) with DCLK1 (red). Scale bars, 10 μm. g, Relative quantification of mRNA expression by qPCR of Alpi, Tslp and Dclk1 (tuft cell markers) from tuft-1, tuft-2 or randomly selected EpCAM+ single cells identified from 96-well plate-based full-length scRNA-seq (16 cells per group) (*P < 0.05, **P < 0.005; t test). h, Validation of CD45 expression in tuft-2 cells. Immunofluorescence assay showing co-expression of the tuft cell marker DCLK1 and of CD45 (left) and CD45 (right, with increased brightness); yellow boxes show three representative tuft cells. Scale bars, 200 μm. i, Isolation of tuft-2 cells based on CD45 expression using FACS. Shown is t-SNE of 332 EpCAM+/CD45+ FACS-sorted single cells (points; n = 3 pooled mice), coloured by unsupervised clustering (top left), the expression of the Tuft cell marker Dclk1 (top right), or the signature scores for tuft-1 and tuft-2 cells (bottom left and right, respectively).
Extended Data Figure 8 Microfold cells from RANKL-treated intestinal organoids and in vivo.
Related to Fig. 5. a–d, Microfold cells in RANKL-treated organoids. a–c, t-SNE of 5,434 single cells (points) from control (left) or RANKL-treated (middle and right) intestinal organoids, or colouring each cell (b, c) by the expression (log2(TPM + 1)) of the canonical microfold cell markers TNF-α-induced protein 2 (Tnfaip2, M-sec; b) and glycoprotein 2 (Gp2; c) (n = 4 pooled wells per treatment condition). d, Expression of microfold cell marker genes35,37,66 in each of the organoid cell clusters. Violin plots show the distribution of expression levels (log2(TPM + 1)) for each of ten previously reported microfold cell marker genes37 (columns), in the cells (points) in each of 13 clusters, including mature microfold cells (red), identified by k-nearest-neighbour clustering of the 5,434 scRNA-seq profiles from organoids. e, f, Microfold cell gene signature in vitro. Heatmaps show for each mature or stem cell cluster of organoid-derived intestinal epithelial cells (columns) the mean expression of genes (rows) for known (grey bars) or newly identified (black bars) microfold cell markers (e) or transcription factors (f), identified as being specific (FDR < 0.05; Mann–Whitney U test) to microfold cells in vitro and in vivo (Methods). g, Congruence of in vitro- and in vivo-derived microfold cell gene signatures. Violin plot shows the distribution of the mean expression of the in vitro-derived signature genes across the in vivo microfold cells (red) and across all other cells derived from the follicle-associated epithelia (grey).
Extended Data Figure 9 Intestinal epithelial cell response to pathogenic stress.
Related to Fig. 6. a, Generalized and pathogen-specific response genes. Volcano plots show for each gene (points) the differential expression and its associated significance (−log10(Q value); likelihood-ratio test) in response to either Salmonella (top) or H. polygyrus (bottom). Genes strongly upregulated in Salmonella (FDR < 10−6) or H. polygyrus (FDR < 5 × 10−3) are highlighted in purple or red, respectively. All highlighted genes are significantly differentially expressed (FDR < 0.05) in both 3′ and full-length scRNA-seq datasets. Left, all genes differentially expressed in the noted pathogen infection versus uninfected controls; middle, the subset differentially expressed in both pathogens versus control; right, the subset differentially expressed in only the noted pathogen, but not the other (Methods). b, Global induction of enterocyte-specific genes across cells during Salmonella infection. Shown is t-SNE of 5,010 single intestinal epithelial cells from control wild-type mice (left) and mice infected with Salmonella (right). Cells are coloured by the expression of the indicated genes, all specific to enterocytes in control mice (Supplementary Tables 2–4) and strongly upregulated by infection (FDR < 10−10 in both 3′ and full-length scRNA-seq datasets). c, Intestinal epithelial cell programs in Salmonella infection. Enriched (–log10(Q value)) gene ontology terms in genes induced in Salmonella-treated intestinal epithelial cells versus control. d, Cell-intrinsic changes after Salmonella infection. Relative expression (row-wise Z scores; colour scale) of 104 genes (top), of which 58 (bottom) are specific to Salmonella infection, significantly upregulated (FDR < 0.05; Mann–Whitney U test; log2(fold change) > 0.1) in enterocytes (columns) from Salmonella infection. Ten representative genes are labelled. e, Upregulation of pro-inflammatory apolipoproteins serum amyloid A 1 and 2 (Saa1 and Saa2) in distal enterocytes under Salmonella infection. Violin plot shows log2(TPM + 1) expression level of Saa1 (top) and Saa2 (bottom) across all post-mitotic cell types from control and Salmonella-treated mice (n = 4 mice; sample identity shown in the legend) (*FDR < 0.01, **FDR < 0.0001; Mann–Whitney U test). f, Upregulation of antimicrobial peptides by Paneth cells after Salmonella infection. Violin plots show log2(TPM + 1) expression levels of genes encoding antimicrobial peptides and the mucosal pentraxin Mptx2 in the cells (points) from control and Salmonella-infected mice (n = 4 mice; sample identity shown in the legend) (*FDR < 0.1, **FDR < 0.01, ***FDR < 0.0001; Mann–Whitney U test). g, Paneth cell numbers detected (using graph clustering; Methods) after Salmonella infection. Frequencies of Paneth cells in each mouse (points) under each condition (see legend) (**FDR < 0.01; Wald test). Error bars, s.e.m.
Extended Data Figure 10 Goblet and tuft cell responses to H. polygyrus show a unique defence mechanism.
Related to Fig. 6. a, Genes induced significantly in response to infection in a non-cell-type-specific manner. Shown is t-SNE visualization of 9,842 single intestinal epithelial cells (points) from control wild-type mice (left), mice infected with H. polygyrus for 3 or 10 days (middle) and mice infected with Salmonella (right). Cells are coloured by the expression (log2(TPM + 1)) of the indicated genes. Genes were selected as significantly differentially expressed in response to infection in a non-cell-type-specific manner (FDR < 0.001 in both the 3′ scRNA-seq and full-length scRNA-seq datasets). b, c, Identification of the tuft-1 and tuft-2 subsets in the dataset of control, Salmonella- and H. polygyrus-infected cells. b, Violin plots of the distribution of the respective signature scores (left and middle) and the expression of Dclk1 (right; log2 (TPM+1)) in cells (points) in each of the tuft subsets. c, t-SNE mapping of the 409 tuft progenitor, tuft-1 and tuft-2 cells, coloured by the scores for each signature (left and middle) and by their assignment to subtype clusters via k-nearest-neighbour graph clustering (right). d, Induction of antiparasitic genes by goblet cells after helminth infection. Shown is the distribution of expression (log2(TPM+1)) of three antiparasitic immunity genes67 upregulated by goblet cells in response to H. polygyrus infection (FDR < 0.05; Mann–Whitney U test), in control and infected mice. e, Antiparasitic protein secretion by goblet cells during H. polygyrus infection. Immunofluorescence assay of formalin-fixed paraffin-embedded (FFPE) sections of RELMβ (top left; red) and E-cadherin (bottom left; green), and their merged view including DAPI nuclear stain (blue) (right), after 10 days of helminth infection. Arrow, sections of H. polygyrus; scale bars, 200 μm.
Supplementary information
Supplementary Table 1 - Summary of single-cell RNAseq experiments.
This table provides the number (after quality filtering, see Methods) of individual intestinal epithelial cells profiled in each of the in this study. (XLSX 31 kb)
Supplementary Table 2 - Cell-type specific signature genes – droplet-based dataset.
This table provides the lists of genes specific to each of the identified clusters of intestinal epithelial cells, identified using 3’ droplet-based scRNA-seq data (Figure 1b). (XLSX 2008 kb)
Supplementary Table 3 - Cell-type specific signature genes – plate-based dataset.
This table provides the lists of genes specific to each of the identified clusters of intestinal epithelial cells, identified using full-length scRNA-seq data (Extended Data Fig. 2a). (XLSX 1253 kb)
Supplementary Table 4 - Consensus cell-type specific signature genes – both datasets.
This table provides high-confidence lists of genes specific to each subtype of intestinal epithelial cells in both 3’ droplet-based and full-length scRNA-seq datasets. (XLSX 13 kb)
Supplementary Table 5 - Cell-type specific TFs and receptors.
This table provides lists of genes annotated as either transcription factors (TFs), G protein-coupled receptors (GPCRs), or leucine-rich repeat (LRR) proteins, enriched in each subtype of intestinal epithelial cells in full-length plate-based scRNA-seq data. (XLSX 13 kb)
Supplementary Table 6 - Enteroendocrine cell subset signature genes.
This table provides the lists of genes specific to each of the identified subsets of tuft cells, identified using both 3’ droplet-based and full-length scRNA-seq data. (XLSX 1482 kb)
Supplementary Table 7 - Consensus tuft cell subset signature genes.
This table provides the lists of genes specific to each of the identified subsets of tuft cells, identified using both 3’ droplet-based and full-length scRNA-seq data. (XLSX 611 kb)
Supplementary Table 8 - In vitro and in vivo M cell signature genes.
This table provides the lists of genes specific to intestinal microfold (M) cells, using 3’ droplet-based scRNA-seq data from in vitro cells derived from RANKL-treated organoids, and in vivo cells derived from the mouse follicle associated epithelia (FAE). (XLSX 319 kb)
Supplementary Table 9 - Intestinal epithelial response to pathogenic infection.
This table provides estimates of differential gene expression in response to infection with H. polygyrus and Salmonella enterica, for each epithelial cell type, using both full-length and 3’ droplet-based scRNA-seq data. (XLSX 6005 kb)
Supplementary Table 10 - Markers of proximal and distal Paneth cells.
This table provides estimates of differential gene expression between two subsets of Paneth cells identified by clustering and interpreted (post-hoc) as derived from proximal and distal small intestine (Extended Data Fig. 3). (XLSX 1541 kb)
Rights and permissions
About this article
Cite this article
Haber, A., Biton, M., Rogel, N. et al. A single-cell survey of the small intestinal epithelium. Nature 551, 333–339 (2017). https://doi.org/10.1038/nature24489
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24489
This article is cited by
-
Integrated meta-omics reveals the regulatory landscape involved in lipid metabolism between pig breeds
Microbiome (2024)
-
The role of goblet cells in Crohn’ s disease
Cell & Bioscience (2024)
-
Niche-DE: niche-differential gene expression analysis in spatial transcriptomics data identifies context-dependent cell-cell interactions
Genome Biology (2024)
-
LSD1 drives intestinal epithelial maturation and controls small intestinal immune cell composition independent of microbiota in a murine model
Nature Communications (2024)
-
Distal colonocytes targeted by C. rodentium recruit T-cell help for barrier defence
Nature (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.