Abstract
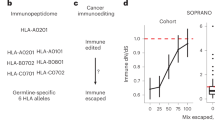
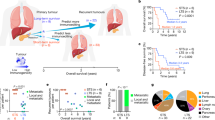
Checkpoint blockade immunotherapies enable the host immune system to recognize and destroy tumour cells1. Their clinical activity has been correlated with activated T-cell recognition of neoantigens, which are tumour-specific, mutated peptides presented on the surface of cancer cells2,3. Here we present a fitness model for tumours based on immune interactions of neoantigens that predicts response to immunotherapy. Two main factors determine neoantigen fitness: the likelihood of neoantigen presentation by the major histocompatibility complex (MHC) and subsequent recognition by T cells. We estimate these components using the relative MHC binding affinity of each neoantigen to its wild type and a nonlinear dependence on sequence similarity of neoantigens to known antigens. To describe the evolution of a heterogeneous tumour, we evaluate its fitness as a weighted effect of dominant neoantigens in the subclones of the tumour. Our model predicts survival in anti-CTLA-4-treated patients with melanoma4,5 and anti-PD-1-treated patients with lung cancer6. Importantly, low-fitness neoantigens identified by our method may be leveraged for developing novel immunotherapies. By using an immune fitness model to study immunotherapy, we reveal broad similarities between the evolution of tumours and rapidly evolving pathogens7,8,9.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Topalian, S. L., Drake, C. G. & Pardoll, D. M. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer Cell 27, 450–461 (2015)
Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015)
Gubin, M. M., Artyomov, M. N., Mardis, E. R. & Schreiber, R. D. Tumor neoantigens: building a framework for personalized cancer immunotherapy. J. Clin. Invest. 125, 3413–3421 (2015)
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014)
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015)
Rizvi, N. A. et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015)
Łuksza, M. & Lässig, M. A predictive fitness model for influenza. Nature 507, 57–61 (2014)
Wang, S. et al. Manipulating the selection forces during affinity maturation to generate cross-reactive HIV antibodies. Cell 160, 785–797 (2015)
Nourmohammad, A., Otwinowski, J. & Plotkin, J. B. Host–pathogen coevolution and the emergence of broadly neutralizing antibodies in chronic infections. PLoS Genet. 12, e1006171 (2016)
Le, D. T. et al. Mismatch-repair deficiency predicts response of solid tumors to PD-1 blockade. Science 357, 409–413 (2017)
Tumeh, P. C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014)
Topalian, S. L. et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N. Engl. J. Med. 366, 2443–2454 (2012)
Herbst, R. S. et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature 515, 563–567 (2014)
De Henau, O. et al. Overcoming resistance to checkpoint blockade therapy by targeting PI3Kγ in myeloid cells. Nature 539, 443–447 (2016)
Ayers, M. et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017)
McGranahan, N. et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351, 1463–1469 (2016)
Anagnostou, V. et al. Evolution of neoantigen landscape during immune checkpoint blockade in non-small cell lung cancer. Cancer Discov. 7, 264–276 (2017)
Abelin, J. G. et al. Mass spectrometry profiling of HLA-associated peptidomes in mono-allelic cells enables more accurate epitope prediction. Immunity 46, 315–326 (2017)
Andreatta, M. & Nielsen, M. Gapped sequence alignment using artificial neural networks: application to the MHC class I system. Bioinformatics 32, 511–517 (2016)
Vita, R. et al. The immune epitope database (IEDB) 3.0. Nucleic Acids Res. 43, D405–D412 (2015)
Murugan, A., Mora, T., Walczak, A. M. & Callan, C. G. Jr. Statistical inference of the generation probability of T-cell receptors from sequence repertoires. Proc. Natl Acad. Sci. USA 109, 16161–16166 (2012)
Birnbaum, M. E. et al. Deconstructing the peptide-MHC specificity of T cell recognition. Cell 157, 1073–1087 (2014)
Lehmann, J., Libchaber, A. & Greenbaum, B. D. Fundamental amino acid mass distributions and entropy costs in proteomes. J. Theor. Biol. 410, 119–124 (2016)
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015)
Tanne, A. et al. Distinguishing the immunostimulatory properties of noncoding RNAs expressed in cancer cells. Proc. Natl Acad. Sci. USA 112, 15154–15159 (2015)
Vétizou, M. et al. Anticancer immunotherapy by CTLA-4 blockade relies on the gut microbiota. Science 350, 1079–1084 (2015)
Dubin, K. et al. Intestinal microbiome analyses identify melanoma patients at risk for checkpoint-blockade-induced colitis. Nat. Commun. 7, 10391 (2016)
Strønen, E. et al. Targeting of cancer neoantigens with donor-derived T cell receptor repertoires. Science 352, 1337–1341 (2016)
Johnson, D. B. et al. Fulminant myocarditis with combination immune checkpoint blockade. N. Engl. J. Med. 375, 1749–1755 (2016)
Hofmann, L. et al. Cutaneous, gastrointestinal, hepatic, endocrine, and renal side-effects of anti-PD-1 therapy. Eur. J. Cancer 60, 190–209 (2016)
Deshwar, A. G. et al. PhyloWGS: reconstructing subclonal composition and evolution from whole-genome sequencing of tumors. Genome Biol. 16, 35 (2015)
Stormo, G. D. Modeling the specificity of protein–DNA interactions. Quant. Biol. 1, 115–130 (2013)
Yu, W. et al. Clonal deletion prunes but does not eliminate self-specific αβ CD8+ T lymphocytes. Immunity 42, 929–941 (2015)
Legoux, F. P. et al. CD4+ T cell tolerance to tissue-restricted self antigens is mediated by antigen-specific regulatory T cells rather than deletion. Immunity 43, 896–908 (2015)
Paul, S. et al. HLA class I alleles are associated with peptide-binding repertoires of different size, affinity, and immunogenicity. J. Immunol. 191, 5831–5839 (2013)
Mason, D. A very high level of crossreactivity is an essential feature of the T-cell receptor. Immunol. Today 19, 395–404 (1998)
Sewell, A. K. Why must T cells be cross-reactive? Nat. Rev. Immunol. 12, 669–677 (2012)
Henikoff, S. & Henikoff, J. G. Amino acid substitution matrices from protein blocks. Proc. Natl Acad. Sci. USA 89, 10915–10919 (1992)
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015)
Nielsen, M., Lundegaard, C., Lund, O. & Keşmir, C. The role of the proteasome in generating cytotoxic T-cell epitopes: insights obtained from improved predictions of proteasomal cleavage. Immunogenetics 57, 33–41 (2005)
Acknowledgements
We thank N. Bhardwaj, C. Callan, S. Cocco, Y. Elhanati, J. Finnegan, D. Krotov, M. Lässig, S. Leach, S. Leibler, A. Libchaber, R. Monasson, A. Nourmohammad, V. Roudko, Z. Sethna, A. Snyder-Charen, P. Sulc, D. T. Ting and the members of the Chan, Greenbaum and Wolchok laboratories for discussions; M. Lässig for suggestions regarding the biophysical model and comments on the manuscript; A. Snyder-Charen and D. T. Ting for their reading of the manuscript. Research was supported by a Stand Up To Cancer–American Cancer Society Lung Cancer Dream Team Translational Research Grant (SU2C-AACR-DT17-15) (M.D.H., T.M., T.A.C. and J.D.W.), a Stand Up To Cancer–National Science Foundation–Lustgarten Foundation Convergence Dream Team Grant (V.P.B., A.S., J.D.W., and B.D.G.), a Phillip A. Sharp Innovation in Collaboration Award from Stand Up To Cancer (B.D.G. and J.D.W.), the Janssen Research & Development LLC (M.Ł.), the STARR Cancer Consortium (T.A.C.), the Pershing Square Sohn Cancer Research Alliance (T.A.C.), the NIH R01 CA205426 (N.A.R. and T.A.C.), the V Foundation (V.P.B., A.S., J.D.W. and B.D.G), the Lustgarten Foundation (V.P.B., A.S., J.D.W. and B.D.G.), the National Science Foundation (NSF) 1545935 (B.D.G. and J.D.W.), the Swim Across America (V.P.B., T.M. and J.D.W.), Ludwig Institute for Cancer Research, the Parker Institute for Cancer Immunotherapy, the NCI K12 Paul Calabresi Career Development Award for Clinical Oncology K12CA184746-01A1 (V.P.B.). The work was also supported in part by the MSKCC Core Grant (P30 CA008748).
Author information
Authors and Affiliations
Contributions
M.Ł. and B.D.G. designed the mathematical model, analysed data and wrote the manuscript with critical comments from all the authors. N.R., V.M., V.P.B., M.D.H., A.S., N.A.R., T.M., A.J.L., T.A.C. and J.D.W. contributed to data acquisition and analysis. M.Ł., T.A.C., J.D.W. and B.D.G. contributed to study conception and design. M.Ł., N.R., V.M., V.P.B., M.D.H., A.S., N.A.R., T.M., A.J.L., T.A.C., J.D.W. and B.D.G. interpreted the data and provided a critical reading of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
M.Ł. has consulted for Merck. V.P.B. has received research funding from Bristol-Myers Squibb. A.J.L. is on the board of directors for Adaptive Biotechnologies and has consulted for Jansen Pharmaceuticals and Merck. T.A.C. is a co-founder of Gritstone Oncology and holds equity. T.A.C. receives grant funding from Bristol Myers Squibb. N.A.R is co-founder and shareholder of Gritstone Oncology. M.D.H. has consulted for Genentech, BMS, Merck, AstraZeneca, Janssen and Novartis. B.D.G. has consulted for Merck.
Additional information
Reviewer Information Nature thanks R. Simon, S. van den Burg and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Inferred MHC binding affinities of mutant versus wild-type peptides.
Neoantigens used in this study are nine-residue-long peptides with affinities predicted to be less than 500 nM by NetMHC3.4 (ref. 19) (Supplementary Information). We plot predicted affinities of mutant peptides compared to the predicted affinities of their wild-type peptides. A single point mutation can lead to a predicted affinity difference of up to four orders of magnitude.
Extended Data Figure 2 Positions 2 and 9 have a subset of neoantigens with a lower predictive value.
The violin plots represent the data density at a given value on the vertical axis. a, Neoantigens with mutations at position 2 or 9 tend to have wild-type peptides with larger predicted affinities. In particular, this is magnified if the corresponding wild-type residue is non-hydrophobic. b, Those biases are reflected by a wider distribution of amplitudes (Methods, equation (7)) for wild-type peptides with non-hydrophobic residues at positions 2 and 9. c, Shannon entropy of amino acid diversity by position in neoantigens, shown for all distinct HLA types and computed based on neoantigens across all datasets. Positions 2 and 9 have a lower entropy than other residues. Other sites have the same entropy as the overall proteome23 and are therefore unconstrained. Five HLA profiles with non-canonical entropies are highlighted in the plot. These HLA types contributed only five informative neoantigens across all datasets and are therefore not treated differentially in our model.
Extended Data Figure 3 Survival analysis score landscape as a function of model parameters.
a–c, The landscape of log-rank test scores as the function of the parameters of the TCR-binding model (a and 1 / k), shown for the consistent choice of τ = 0.09; colours represent the significance level of the long-rank test. The regions of high scores are similar across all three datasets. The point corresponding to consistent parameters (a = 26 and k = 4.87) is marked by a black dot in each plot. d–f, log-rank score for the fitness model at consistent binding-function parameters, plotted as a function of τ. Dashed vertical lines are at τ = 0.09, thin solid lines mark the score values corresponding to significance of P = 0.05, P = 0.01 and P = 0.005 (n = 103 (a, d), n = 64 (b, e), n = 34 (c, f)).
Extended Data Figure 4 Alignments to IEDB epitopes.
The TCR recognition probability for a neoantigen is a sigmoidal function of the alignment scores of a given neoantigen to the IEDB epitopes, evaluated for the set of neoantigens from the cohort of patients from ref. 5, using the consistent set of parameters.
Extended Data Figure 5 Effect of IEDB sequence content on predictive power of the neoantigen fitness model.
Predictions were performed using subsampled IEDB epitope sequences, with subsampling rate varying between 0.1 and 0.9. For each rate, 10,000 iterations were performed to obtain a distribution of log-rank test scores. The violin plots represent the data density at a given value on the vertical axis (n = 10,000). Solid black lines mark the log-rank test score of the prediction for the full set of epitope sequences and grey thick lines mark the median scores of the subsampled data. a–c, Subsampling of the original set of IEDB sequences, supported by positive T-cell assays, shows that the quality of predictions decreases with subsampling rate. Prediction quality is more robust in the datasets from refs 4, 6. d–f, The analogous subsampling procedure was repeated for IEDB sequences that were not supported by positive T-cell assays. For the datasets from refs 4, 5, model performance is substantially decreased.
Extended Data Figure 6 Reshuffling patient HLA types reduces the predictive power of the neoantigen fitness model.
For each cohort, we performed 10 iterations for the analysis of reshuffled patient HLA types, followed by computational neoantigen prediction, fitness model calculation and survival analysis. We report the distribution of log-rank test scores over these iterations: boxes mark 75% confidence intervals and whiskers mark the range of scores (n = 10). The score values for the model of the original (unshuffled) data are marked with blue squares.
Extended Data Figure 7 Cytolytic score improves prediction quality.
a, Kaplan–Meier curves of overall survival shown for our model applied to the dataset from ref. 4 for a subset of n = 40 patients with transcriptional data. Samples were split by the median value of their tumour’s relative population size n(τ) (equation (1)). Error bars represent the standard error due to sample size. b, A model optimized for cytolytic score significantly separates patients (Methods). c, Inclusion of the cytolytic score in our model improves prediction for the subset of 40 patients. The P values from log-rank tests comparing the two Kaplan–Meier curves are shown above each plot. For a and c, we used consistent parameters trained on the three cohorts (Extended Data Fig. 2); for b, the parameter τ is optimized.
Supplementary information
Supplementary Information
This file contains Supplementary Text and Data and additional references.
Supplementary Data 1
Mutations in tumour samples in all 3 cohorts.
Supplementary Data 2
Putative neoantigens predicted with NetMHC 3.4 in all 3 cohorts.
Supplementary Data 3
Overall survival data for patients in the 3 cohorts.
Supplementary Data 4
Reconstructed tumour clones from the highest scoring tree with PhyloWGS and their neoantigens with computed fitness cost.
Supplementary Data 5
IEDB sequences validated by positive T-cell assays (positive) and IEDB sequences not validated by positive T-cell assays (negative).
Supplementary Data 6
Cytolytic score for a subset of 40 Van Allen et al. samples used in Extended Data Fig. 7
Supplementary Data 7
Source code example to compute neoantigen fitness cost.
Rights and permissions
About this article
Cite this article
Łuksza, M., Riaz, N., Makarov, V. et al. A neoantigen fitness model predicts tumour response to checkpoint blockade immunotherapy. Nature 551, 517–520 (2017). https://doi.org/10.1038/nature24473
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24473
This article is cited by
-
Vaccines: a promising therapy for myelodysplastic syndrome
Journal of Hematology & Oncology (2024)
-
Bioinformatic analysis and experimental validation of cuproptosis-related LncRNA as a novel biomarker for prognosis and immunotherapy of oral squamous cell carcinoma
Hereditas (2024)
-
Computational immunogenomic approaches to predict response to cancer immunotherapies
Nature Reviews Clinical Oncology (2024)
-
Mi-2β promotes immune evasion in melanoma by activating EZH2 methylation
Nature Communications (2024)
-
Tissue-specific thresholds of mutation burden associated with anti-PD-1/L1 therapy benefit and prognosis in microsatellite-stable cancers
Nature Cancer (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.