Abstract
Direct lineage conversion offers a new strategy for tissue regeneration and disease modelling. Despite recent success in directly reprogramming fibroblasts into various cell types, the precise changes that occur as fibroblasts progressively convert to the target cell fates remain unclear. The inherent heterogeneity and asynchronous nature of the reprogramming process renders it difficult to study this process using bulk genomic techniques. Here we used single-cell RNA sequencing to overcome this limitation and analysed global transcriptome changes at early stages during the reprogramming of mouse fibroblasts into induced cardiomyocytes (iCMs)1,2,3,4. Using unsupervised dimensionality reduction and clustering algorithms, we identified molecularly distinct subpopulations of cells during reprogramming. We also constructed routes of iCM formation, and delineated the relationship between cell proliferation and iCM induction. Further analysis of global gene expression changes during reprogramming revealed unexpected downregulation of factors involved in mRNA processing and splicing. Detailed functional analysis of the top candidate splicing factor, Ptbp1, revealed that it is a critical barrier for the acquisition of cardiomyocyte-specific splicing patterns in fibroblasts. Concomitantly, Ptbp1 depletion promoted cardiac transcriptome acquisition and increased iCM reprogramming efficiency. Additional quantitative analysis of our dataset revealed a strong correlation between the expression of each reprogramming factor and the progress of individual cells through the reprogramming process, and led to the discovery of new surface markers for the enrichment of iCMs. In summary, our single-cell transcriptomics approaches enabled us to reconstruct the reprogramming trajectory and to uncover intermediate cell populations, gene pathways and regulators involved in iCM induction.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Ieda, M. et al. Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell 142, 375–386 (2010)
Jayawardena, T. M. et al. microRNA-mediated in vitro and in vivo direct reprogramming of cardiac fibroblasts to cardiomyocytes. Circ. Res. 110, 1465–1473 (2012)
Qian, L. et al. In vivo reprogramming of murine cardiac fibroblasts into induced cardiomyocytes. Nature 485, 593–598 (2012)
Song, K. et al. Heart repair by reprogramming non-myocytes with cardiac transcription factors. Nature 485, 599–604 (2012)
Dal-Pra, S., Hodgkinson, C. P., Mirotsou, M., Kirste, I. & Dzau, V. J. Demethylation of H3K27 is essential for the induction of direct cardiac reprogramming by miR combo. Circ. Res. 120, 1403–1413 (2017)
Ma, H., Wang, L., Yin, C., Liu, J. & Qian, L. In vivo cardiac reprogramming using an optimal single polycistronic construct. Cardiovasc. Res. 108, 217–219 (2015)
Mathison, M. et al. In vivo cardiac cellular reprogramming efficacy is enhanced by angiogenic preconditioning of the infarcted myocardium with vascular endothelial growth factor. J. Am. Heart Assoc. 1, e005652 (2012)
Mohamed, T. M. et al. Chemical enhancement of in vitro and in vivo direct cardiac reprogramming. Circulation 135, 978–995 (2017)
Zhao, Y. et al. High-efficiency reprogramming of fibroblasts into cardiomyocytes requires suppression of pro-fibrotic signalling. Nat. Commun. 6, 8243 (2015)
Zhou, H., Dickson, M. E., Kim, M. S., Bassel-Duby, R. & Olson, E. N. Akt1/protein kinase B enhances transcriptional reprogramming of fibroblasts to functional cardiomyocytes. Proc. Natl Acad. Sci. USA 112, 11864–11869 (2015)
Ifkovits, J. L., Addis, R. C., Epstein, J. A. & Gearhart, J. D. Inhibition of TGFβ signaling increases direct conversion of fibroblasts to induced cardiomyocytes. PLoS ONE 9, e89678 (2014)
Liu, Z. et al. Re-patterning of H3K27me3, H3K4me3 and DNA methylation during fibroblast conversion into induced cardiomyocytes. Stem Cell Res. 16, 507–518 (2016)
Muraoka, N. et al. MiR-133 promotes cardiac reprogramming by directly repressing Snai1 and silencing fibroblast signatures. EMBO J. 33, 1565–1581 (2014)
Wang, L. et al. Stoichiometry of Gata4, Mef2c, and Tbx5 influences the efficiency and quality of induced cardiac myocyte reprogramming. Circ. Res. 116, 237–244 (2015)
Zhou, Y. et al. Bmi1 is a key epigenetic barrier to direct cardiac reprogramming. Cell Stem Cell 18, 382–395 (2016)
Shin, J. et al. Single-cell RNA-Seq with waterfall reveals molecular cascades underlying adult neurogenesis. Cell Stem Cell 17, 360–372 (2015)
Welch, J. D., Hartemink, A. J. & Prins, J. F. SLICER: inferring branched, nonlinear cellular trajectories from single cell RNA-seq data. Genome Biol. 17, 106 (2016)
Polo, J. M. et al. A molecular roadmap of reprogramming somatic cells into iPS cells. Cell 151, 1617–1632 (2012)
Vaseghi, H. R. et al. Generation of an inducible fibroblast cell line for studying direct cardiac reprogramming. Genesis 54, 398–406 (2016)
Fu, X. D. & Ares, M. Jr. Context-dependent control of alternative splicing by RNA-binding proteins. Nat. Rev. Genet. 15, 689–701 (2014)
Park, J. W., Jung, S., Rouchka, E. C., Tseng, Y. T. & Xing, Y. rMAPS: RNA map analysis and plotting server for alternative exon regulation. Nucleic Acids Res. 44, W333–W338 (2016)
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014)
Wang, L. et al. Improved generation of induced cardiomyocytes using a polycistronic construct expressing optimal ratio of Gata4, Mef2c and Tbx5. J. Vis. Exp. 105, e53426 (2015)
He, A., Kong, S. W., Ma, Q. & Pu, W. T. Co-occupancy by multiple cardiac transcription factors identifies transcriptional enhancers active in heart. Proc. Natl Acad. Sci. USA 108, 5632–5637 (2011)
Waldron, L. et al. The cardiac TBX5 interactome reveals a chromatin remodeling network essential for cardiac septation. Dev. Cell 36, 262–275 (2016)
Zheng, X. et al. Prolyl hydroxylation by EglN2 destabilizes FOXO3a by blocking its interaction with the USP9x deubiquitinase. Genes Dev. 28, 1429–1444 (2014)
Qian, L., Berry, E. C., Fu, J. D., Ieda, M. & Srivastava, D. Reprogramming of mouse fibroblasts into cardiomyocyte-like cells in vitro. Nat. Protoc. 8, 1204–1215 (2013)
Wu, A. R. et al. Quantitative assessment of single-cell RNA-sequencing methods. Nat. Methods 11, 41–46 (2014)
Treutlein, B. et al. Reconstructing lineage hierarchies of the distal lung epithelium using single-cell RNA-seq. Nature 509, 371–375 (2014)
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015)
Brennecke, P. et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods 10, 1093–1095 (2013)
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007)
Chen, C. et al. Removing batch effects in analysis of expression microarray data: an evaluation of six batch adjustment methods. PLoS ONE 6, e17238 (2011)
Han, H. et al. MBNL proteins repress ES-cell-specific alternative splicing and reprogramming. Nature 498, 241–245 (2013)
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014)
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015)
Shen, S. et al. rMATS: robust and flexible detection of differential alternative splicing from replicate RNA-seq data. Proc. Natl Acad. Sci. USA 111, E5593–E5601 (2014)
Pinto, A. R. et al. An abundant tissue macrophage population in the adult murine heart with a distinct alternatively-activated macrophage profile. PLoS ONE 7, e36814 (2012)
Whitfield, M. L. et al. Identification of genes periodically expressed in the human cell cycle and their expression in tumors. Mol. Biol. Cell 13, 1977–2000 (2002)
Gooding, C. & Smith, C. W. Tropomyosin exons as models for alternative splicing. Adv. Exp. Med. Biol. 644, 27–42 (2008)
Terenzi, F. & Ladd, A. N. Conserved developmental alternative splicing of muscleblind-like (MBNL) transcripts regulates MBNL localization and activity. RNA Biol. 7, 43–55 (2010)
Dube, D. K., McLean, M. D., Dube, S. & Poiesz, B. J. Translational control of tropomyosin expression in vertebrate hearts. Anat. Rec. (Hoboken) 297, 1585–1595 (2014)
Acknowledgements
We thank UNC AAC Core, HTSF Core, Flow Core for technical support. This study was supported by NIH HG06272 to J.F.P., NIH BD2K Fellowship (T32 CA201159) and NIH F31 Fellowship (HG008912) to J.D.W., NIH/NHLBI R00 HL109079 and American Heart Association (AHA) 15GRNT25530005 to J.L., AHA 13SDG17060010, Ellison Medical Foundation (EMF) AG-NS-1064-13, and NIH/NHLBI R01HL128331 to L.Q., and gifts from H. McAllister and C. Sewell.
Author information
Authors and Affiliations
Contributions
L.Q., Z.L. and L.W. conceived and designed the study. Z.L. and L.W. designed and performed single-cell RNA-seq. Z.L., L.W., Y.Z. and C.Y. prepared samples for microarray and bulk RNA-seq. L.W., Y.Z., H.M., H.R.V., C.Y., S.Y., J.B.W., S.A. and M.Z. performed other experiments. Z.L., J.D.W. and J.F.P. performed data analysis and modelling. W.S. helped with statistical analysis. Z.L, J.D.W., J.L. and L.Q. wrote the manuscript, with extensive input from all authors. J.L. and L.Q. provided funding and overall supervision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Experimental design, analysis pipeline, quality control and normalization for single-cell RNA-seq.
a, Experimental workflow. Hearts were isolated from P1.5 neonatal mice and cells were dissociated by enzymatic digestion. Thy1+ cells were then purified by MACS and plated overnight. The adherent cells (cardiac fibroblasts) were then transduced with retroviruses encoding the reprogramming factors Mef2c, Gata4 and Tbx5, DsRed or were left untransduced (mock). On day 3 after transduction, cells were trypsinized and Live/Dead stained. Additionally, for some experiments designed to examine the relative mouse RNA abundance in cells receiving different treatments, mock and M + G + T cells were labelled with a green cell tracker (CFSE), FACS-sorted for live cells, and then mixed at a designated ratio with FACS-sorted live DsRed+ cells from a parallel DsRed transduction. The single-cell suspension was loaded onto a medium size chip (10–17 μm) and single cells were captured on a Fluidigm C1 machine. Bright field, and for some experiments, fluorescent images, were taken for all capture sites. Individual cDNA libraries for each cell were prepared in situ by reverse transcription with pre-amplification after adding RNA spike-in. Bright field and/or fluorescent images for each capture site were examined and libraries from nests with 0 or multiple cells were excluded from downstream analysis. Illumina libraries were then prepared for each cell, pooled, quality-checked and sequenced on Hiseq 2500. b, Design of the seven independent single-cell RNA-seq experiments, including treatment, RNA spike-ins and Fluidigm chips used. c, Data analysis pipeline. Barcodes were trimmed from RNA-seq raw reads and the quality of these reads was confirmed with FASTQC. High-quality reads were mapped to the mm10 genome with TopHat2 and counted with Htseq count. Outliers were removed as described in d. The raw counts were normalized first to technical and biological size factors within each experiment using DEseq and then to experiment size factors calculated based on relative mouse mRNA abundance in cells receiving different treatments (h). Residual batch effects between experiments receiving the same treatment were removed using ComBat. Cell grouping and modelling were then performed using the normalized gene counts with PCA, hierarchical clustering, SLICER and more. The most important three quality control steps are labelled in red in a and c. The above strict quality control criteria ensured that only high-quality and biologically meaningful data from healthy single cells were analysed. d, For each of the seven single-cell experiments, the percentage of reads mapped to spike-in in each cell was plotted against percentage of reads mapped to mouse genome (left) or mouse mRNA (right) in that cell. Cells outside of the red circles are outliers. e, For each of the five single-cell experiments that contained ERCC spike-in, mean count numbers of each ERCC spike-in were plotted against their concentration in lysis mix A (see Fluidigm’s protocol for details). Linear regression coefficients (R value) and corresponding P values (two-sided, α = 0.05) are shown. The results showed a dynamic range (~105) of ERCC concentration that covers the full spectrum of mouse gene expression levels. The high R values indicate a strong correlation between hypothetical molecular concentrations and measured gene counts in our experiments. f, Squared coefficients of variation (CV2) were plotted against mean expression of ERCC spike-ins (left) or mouse genes (right) for experiments containing ERCC spike-in. g, DsRed counts in experiments E3 and E4–E7 plotted against Mef2c counts and/or total mouse mRNA counts after normalization for technical and biological size factors within each experiment (Methods). Cells in the four experiments were classified as DsRed-transduced (E3R, E5R, E6R, E7R), M + G + T-transduced (E5M, E6M) or untransduced cells (E3U, E7U) based on these plots. h, i, Normalization to experiment size factors to account for technical contributions to experiment-to-experiment variations, such as varied capture efficiency, while retain biological variations, such as differences in total mRNA abundance in cells receiving different treatments. h, Median total mouse mRNA counts were calculated for each treatment in each experiment and mean mRNA counts were compared between different experiments (E1, E2 and E4) or different treatments in one experiment (E3, E4–E7) with a two-sided Student’s t-test (α = 0.05). Experiment size factors were calculated based on the ratio of median mRNA counts between different treatments. After normalization to the experiment size factors, the median mRNA count was equal to 1,000,000 cells for uninfected and DsRed-transduced cells and 616,136 cells for M + G + T transduction. i, PCA of two biological replicates (E5 and E6) that have different sequencing depth per cell due to different capture efficiencies before (left) and after (right) normalization to experiment size factors. Top 400 PCA genes were used.
Extended Data Figure 2 Normalization and outlier removal of single-cell RNA-seq data (continued).
a–c, Removal of batch effects using ComBat on non-immune cells (described in Extended Data Fig. 3). PCA of all batches of M + G + T-transduced (a), DsRed-transduced (b) or uninfected (c) cells before (left) and after (right) ComBat normalization. d–g, After ComBat normalization, outliers of each treatment group were further removed by examination of the mean gene expression level of each cell (box plot, d) and PCA (e). Uninfected cells are shown as an example in d and e. A total of 454 healthy non-immune cells were further analysed. f, Pairwise comparison of mean mouse gene expression between different experiments and treatment conditions. Correlation coefficient calculated by linear regression is shown. g, Heat map coloured by correlation coefficient in f. The strongest correlation was seen within each treatment group. DsRed-infected and uninfected cells also showed strong intertreatment correlation. M + G + T-transduced cells showed relatively low correlation with DsRed-transduced and uninfected cells.
Extended Data Figure 3 Single-cell analysis identified a sub-population of immune-like cells.
a–c, Data from 513 control or reprogramming single cardiac fibroblasts and bulk RNA-seq data of neonatal cardiac fibroblasts and cardiomyocytes were analysed with PCA. To identify groups of related cells and genes, the top 400 genes with highest weight in the first three principal components were then analysed by unsupervised hierarchical clustering (a) and PCA (b, c). Representative genes in each of the four gene clusters identified by hierarchical clustering are listed to the right of hierarchical clustering heat map. Notably, in addition to cardiomyocyte, fibroblast and cell-cycle genes, immune response genes were identified as the other major gene cluster. b, PCA loading plot showing four major gene clusters. c, PCA score plot. Both hierarchical clustering and PCA results showed that bulk cardiac fibroblast and cardiomyocyte data were very close in distance and both of them were clustered together with single cells expressing high levels of immune genes. The dashed line separates immune-like single cells and other single cells. d, Markers of the major immune cell lineages. e, Violin plots of the expression of major immune cell lineage markers in bulk cardiac fibroblasts, bulk cardiomyocytes, immune-like single cells and other single cells. f, Expression of the macrophage marker Cd14 and the dendritic cell marker Cd11c (also known as Itgax) in each immune-like cell showed that 42 cells express macrophage markers and 3 express dendritic cell markers, with 4 cells expressing both. These data suggest that the immune-like cells are probably cardiac resident immune cells38, which also express cardiac fibroblast markers, such as Thy1. Although follow-up work to delineate the potential of these immune cells to be reprogrammed into iCMs will be of great interest, it is not the focus of this study. Therefore, for all subsequent analyses of the single-cell data, we focused on the non-immune cardiac fibroblasts.
Extended Data Figure 4 Cell grouping, representative gene expression, and number of genes detected in single-cell RNA-seq data and population-based gene expression profiling during reprogramming.
Related to Fig. 1a–h. a, PCA scree plot showing variance of top 10 PCs. Related to Fig. 1a–c. b, Violin plots showing the expression levels of representative cardiac, fibroblast and cell-cycle genes in the seven cell groups identified by hierarchical clustering and PCA (Fig. 1a–c). c, Gene ontology analysis from Fig. 1a showing the P value of each gene ontology term. d–g, Determination of the proliferation status of each single cell using genes periodically expressed during the cell cycle that were identified in a previous report39. d, A 3D LLE plot calculated based on the expression of these cell cycle genes in each of the 454 cells. e, Frequency of cells in LLE component 3. The dark red plane in d and the red dotted line in e indicate the threshold for proliferating (Pro) and non-proliferating (NP) cells. f, g, PCA plots as in Fig. 1c, but colour- and shape-coded for proliferating and non-proliferating (f) or CCA and CCI (g) cells. h, i, tSNE plots of all single cells colour- and shape-coded by hierarchical clustering and PCA cell groups (h) or proliferating and non-proliferating cell groups (i). The cells that were grouped as intermediate fibroblasts, pre-iCMs and iCMs constituted 30.6% (77 out of 252), 24.6% (62 out of 252) and 44.8% (113 out of 252) of all cells transduced with M + G + T, respectively. In contrast to previous population- and marker-based studies, our single-cell RNA-seq data suggests that the fate conversion from fibroblast to iCM occurs rapidly (approximately 3 days) with nearly 45% of the cardiac fibroblasts exhibiting transcriptomic signatures indicative of a cardiac fate. j, Live fluorescent images of day-5 MGT-transduced cardiac fibroblasts showing co-expression of α-MHC–GFP and Thy1 (surface labelling). Double-positive cells are labelled with an asterisk (*). All images were taken at 40× magnification. Scale bar, 100 μm. k, α-MHC–GFP+Thy1+ and α-MHC–GFP−Thy1+ cells were FACS-sorted from day-7 MGT-transduced cardiac fibroblasts and expression of representative cardiac (Myl4 and Actc1) and fibroblast (Col3a1 and Postn) markers were determined by qRT–PCR. Day-7 mock-transduced cells were included as control. Data are mean ± s.d. n = 4 samples. One-way ANOVA followed by Bonferroni correction (two-sided), **P < 0.01, *** P < 0.001; ns, not significant. Myl4 and Actc1 expression increased 80–100-fold and reached approximately the same level as Gapdh in α-MHC–GFP+Thy1+ cells compared to mock transduction. Expression level of the fibroblast marker Postn was maintained at a high level in GFP+Thy1+ cells. For the other fibroblast marker, Col3a1, although its relative expression in GFP+Thy1+ cells was decreased compared to mock-transduced and GFP−Thy1+ cells, but its absolute expression was still high compared to Gapdh (around 1.4-fold of Gapdh). The data strongly support the existence of cardiomyocyte- and fibroblast-marker double-positive pre-iCM and suggest that the pre-iCM state represents an intermediate cell population that is transitioning from intermediate fibroblast to iCM or that is locked between intermediate fibroblast and iCM during reprogramming. l, To determine whether iCMs may be differentiated from rare cardiac stem/progenitor cells, we plotted the expression of cardiac stem/progenitor cell markers in each of the hierarchical clustering and PCA single-cell groups using violin plots. All of these markers were nearly undetectable in fibroblasts, intermediate fibroblasts, pre-iCMs and iCMs, suggesting the direct conversion from cardiac fibroblast to iCM without going through a stem/progenitor cell stage. m, Distribution of gene expression levels in single cells. Data are mean ± s.e.m. n = 454 cells. The limit of gene detection was set to 1 based on this plot. n, Distribution of the number of genes detected in all, CCI or CCA single cells. Comparison of the distributions in CCI and CCA cells using a two-sample Kolmogorov–Smirnov test resulted in a one-sided P value of 5.248 × 10–11, suggesting that the number of genes in CCI is significantly smaller than in CCA. On the basis of this result, only CCI cells were used in o. o, Distribution of the number of genes detected in each CCI cell group. Analysis using a one-sided, two-sample Kolmogorov–Smirnov test (P values: 0.00521 for intermediate fibroblasts versus fibroblasts, 0.00481 for pre-iCMs versus intermediate fibroblasts and 1.104 × 10−6 for iCMs versus pre-iCMs) suggests that the number of genes expressed decreased when the cells adopted the iCM fate. This observation demonstrates a dynamic re-patterning of transcription machinery during reprogramming and is consistent with the hierarchical clustering analysis and experimental evidence that pre-iCMs co-expressed both cardiac and fibroblast markers, further indicating that the pre-iCM state constitutes an intermediate population during iCM reprogramming. p–v, Population-based gene expression profiling of reprogramming cardiac fibroblasts at day 0, 3, 5, 7, 10 and 14. p, q, Results from PCA analysis using all genes were similar to those using the top 400 genes (r–v). p, Scree plot of the top 10 PCs. q, A 3D PCA score plot. r–v, Analyses using the top 400 PCA genes. Related to Fig. 1g, h. r, Scree plot of top 10 PCs. s, PCA score plot using PC1 and PC3. t, Hierarchical clustering identified four major gene clusters: gradually upregulated during reprogramming (red, mainly cardiac genes), downregulated in MGT-transduced compared to LacZ-transduced (blue, mainly extracellular matrix (ECM) genes) and gradually upregulated (light grey) or downregulated (dark grey) in both LacZ and MGT cells (culture or viral effects, mainly ECM and immune-response genes). The results are consistent with the expression of representative genes selected from single-cell data (Fig. 1h), showing gradually increased expression of cardiomyocyte markers during reprogramming, first increased and then decreased expression of cell-cycle genes in both MGT and LacZ cells, and significantly lower fibroblast markers in MGT compared to LacZ cells at each time point. u, Heat map showing loading (weight) of the genes in t in PC1, 2 and 3. Upregulated (cardiac) genes are highly weighted in PC1, and the other three gene clusters are highly weighted in PC2 and PC3. The results are consistent with s and Fig. 1g. v, Gene ontology analysis of the four gene clusters in t, showing gene ontology terms and their corresponding P values (listed on the right).
Extended Data Figure 5 Inhibition of cell proliferation or cell-cycle synchronization promotes iCM reprogramming.
Related to Fig. 1i–p. a, Comparison of the ratio of CCA:CCI cells in the three treatment groups: uninfected, DsRed-infected and M + G + T-infected. Analysis using a χ2 test suggests that proliferation states were not significantly different among the treatment groups at day 3. b, c, Knockdown (KD) efficiency of shRNAs (b) or overexpression (OE) levels (c) of cell cycle-related genes were determined by qRT–PCR on day-4 lentiviral transduced cells. shNT, non-targeting control shRNA. Data are mean ± s.d. n = 3 samples. d–i, Cell-cycle staging of cardiac fibroblasts simultaneously transduced with reprogramming factors and shRNA (e, g) or overexpression (f, h) constructs by propidium iodide (PI) staining. e, f, Flow cytometry histogram of propidium iodide staining intensity. g–i, Percentages of cells in G0/G1, S or G2/M phases were calculated based on e and f. i, Summary of g and h. j–m, Measurement of DNA synthesis in cardiac fibroblasts simultaneously transduced with reprogramming factors and shRNA (k) or overexpression (l) constructs by EdU incorporation assay followed by flow cytometry. dMFI, difference (Δ) in median EdU fluorescence intensity between EdU+ cells and EdU− cells. m, Summary of k and l. Constructs that markedly decreased or increased cell proliferation were labelled in red in g–i, k–m and were used for experiments in n–s. n–s, The effect of manipulation of cell proliferation through knockdown or overexpression of cell-cycle-related genes on iCM reprogramming. Reprogramming factors were introduced by lentiviral vectors instead of retroviral vectors to avoid retroviral infection bias of proliferating cells. Cardiac fibroblasts were simultaneously transduced with lentiviral Mef2c, Gata4 and Tbx5 (n–p) or inducible MGT (iMGT, q–s) and lentiviral knockdown or overexpression constructs that markedly decreased (o, r) or increased (p, s) cell proliferation. Percentages of α-MHC–GFP+ and cTnT+ cells were quantified by flow cytometry. t–z, The effect of large T antigen transduction on iCM reprogramming. Cardiac fibroblasts were simultaneously transduced with reprogramming factors and lentiviral large T antigen. After 10 days, α-actinin+cTnT+ cells were immunostained, imaged and quantified by counting randomly selected 20× fields from multiple repeated experiments (u–x). Both percentages of positive cells per field (v, x) and numbers of positive cells per field (w) were quantified. Percentages of cells showing a sarcomere structure in α-actinin+ cells were also quantified (y–z). The percentage of α-actinin+ cells that show sarcomere structures decreased from 50% to 0% upon large T transduction and accelerated proliferation. u, y, Representative images at 40× magnification with Hoechst nuclear staining. Scale bars, 100 μm. o–z, Data are mean ± s.e.m. o–s, n = 4 samples. v, w, z, n = 20 images. x, n = 10 images. b, c, w–z, Two-sided Student’s t-test. o–s, One-way ANOVA followed by Bonferroni correction (two-sided). P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 6 Heterogeneity of the isolated cardiac fibroblasts (Thy1+ non-immune, non-cardiomyocyte cardiac cells) and stepwise suppression of non-cardiomyocyte lineages during iCM reprogramming.
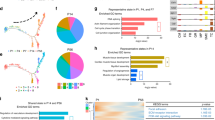
Related to Fig. 2. a–c, Limited transcriptome change by retrovirus transduction. To determine whether introduction of viruses could influence cellular identities of cardiac fibroblasts, molecular features of the uninfected and DsRed-transduced cells were compared and only 25 genes were differentially expressed (ANOVA, P < 0.05), many of which were related to immune response (data not shown), suggesting that uninfected and virus-infected cardiac fibroblasts shared very similar gene expression profiles. a, PCA of the control cells from experiment E3 as shown in Fig. 2b, but colour-coded by treatment. The results showed that uninfected and DsRed-transduced cells were indistinguishable by PCA, suggesting limited global transcriptome changes by retroviral transduction. b, Violin plots showing the expression of representative cardiomyocyte, fibroblast and cell-cycle genes in uninfected- and DsRed-transduced cardiac fibroblasts. Retroviral transduction does not affect the expression of these genes. c, Same as a, but with all control cardiac fibroblasts from experiments E3, E5R, E6R and E7. On the basis of the results from a–c, we concluded that retroviral transduction does not influence cellular identities of cardiac fibroblasts and therefore we analysed control cardiac fibroblasts containing both uninfected and DsRed-transduced cells together in Fig. 2a, b. d, Gene ontology analysis from Fig. 2a showing the P value of each gene ontology term. e, Violin plots showing the expression of additional non-cardiomyocyte lineage markers in cardiac fibroblasts. Related to Fig. 2a. f, PCA analysis of control cardiac fibroblasts from all four experiments (E3, E5R, E6R and E7) with cells colour-coded by non-cardiomyocyte lineage groups. Related to Fig. 2b. g, Immunostaining of Thy1 and α-SMA, or Thy1 and CD31 of day-7 explant cardiac fibroblast cultures. Images were taken at 20× magnification. Scale bars, 200 μm. h, i, Representative flow cytometry plots (h) and quantification (i) of α-SMA+ and CD31+ cells in Thy1+ cells. There were 72.6% α-SMA+ and 9% Cd31+ cardiac fibroblasts, consistent with the single-cell RNA-seq data in Fig. 2a, showing a high percentage of cells expressing myofibroblast/smooth muscle markers and a low percentage of cells expressing endothelial markers. Data are mean ± s.d. j, Violin plots showing the expression levels of additional lineage markers. Related to Fig. 2c. k, Same as j, but using cells from experiments E4–E7. These experiments were performed using the redesigned Fluidigm medium chip as a repeat of experiments E1–E3 (Fig. 2c), which used the original Fluidigm medium chip. l, m, Tracking of protein expression of a myofibroblast/smooth muscle cell marker (α-SMA) by co-staining with α-MHC–GFP in cardiac fibroblasts during reprogramming at days 5, 7, 10, 14 and 21. l, Representative images at 40× magnification with Hoechst nuclear staining. Scale bar, 100 μm. m, Quantification of α-MHC–GFP+ and α-SMA high, low or negative cells. Data are mean ± s.e.m. The results showed that as reprogramming proceeded, protein expression of Thy1, SM22α (Fig. 2d–f) and α-SMA in α-MHC–GFP+ cells decreased over time, with no Thy1highSM22αhighα-SMAhigh cells and around 50–60% of Thy1−SM22α−α-SMA− cells on day 21 of reprogramming.
Extended Data Figure 7 Identification of regulatory pathways involved in iCM reprogramming and screening of a shRNA library against major splicing factors during iCM induction.
a–d, Three clusters of genes that significantly related to, and showed similar trends during, the reprogramming process were identified by nonlinear regression (see Methods). The number of genes included in each cluster is shown in parentheses. The solid line in each plot shows the overall trend of the cluster, and the grey colour indicates the 2D density of gene trends passing through each region of the plot. b–d, Gene ontology analysis of genes in the three clusters showing gene ontology terms with FDR < 0.05. e, f, Screening of a shRNA library of splicing factors for key regulators of iCM reprogramming. icMEFs were induced by doxycycline to express MGT and at the same time were transduced with lentiviruses encoding shRNA targeting various splicing factors. On day 3 after transduction, knockdown efficiency was determined by qRT–PCR (e; n = 6 samples from two independent experiments) and α-MHC–GFP+ cells were quantified by flow cytometry (f; n = 3 samples, data representative of three independent experiments). Data are mean ± s.d. Knockdown of Ptbp1 led to the highest fold increase in percentage of α-MHC–GFP+ cells compared to shNT. g, h, Ptbp1 expression in freshly isolated cardiac fibroblasts and cardiomyocytes was determined by qRT–PCR (g; mean ± s.e.m., n = 8 samples from two independent experiments) or western blotting (h). e, g, Two-sided Student’s t-test. **P < 0.01, ***P < 0.001.
Extended Data Figure 8 Manipulation of Ptbp1 through loss- and gain-of-function during iCM reprogramming.
a, b, Ptbp1 knockdown efficiency of different shRNA clones in day-3 transduced mouse embryonic fibroblasts (MEFs) determined by qRT–PCR (a, mean ± s.e.m., data representative of three independent experiments) or western blotting (b). shPtbp1-271 showed the highest knockdown efficiency (>97%) and was used for subsequent experiments. c–u, Knockdown (shPtbp1, c–p) or overexpression (lentiviral OE-Ptbp1, q–u) of Ptbp1 in neonatal cardiac fibroblasts (neoCF) (c–h, q–u), AdCF (i–l) or AdTTF (m–p) when iCM reprogramming was induced by MGT (except in e–h, where M + G + T was used as a further confirmation). After 10 days (14 days for OE-Ptbp1), expression of cardiac markers was determined by immunostaining followed by imaging and blinded quantification (e, f, i, j, m, n, r, s) or flow cytometry (c, d, g, h, k, l, o, p, t). e, i, m, r, Representative 20× images with Hoechst nuclear staining. Scale bars, 200 μm. f, j, n, s, n = 10–20 images. Data are mean ± s.e.m. c, g, k, o, Representative flow cytometry plots. Percentages of cells are shown. d, h, l, p, t, Quantification of flow cytometry data measured in triplicate. Data are mean ± s.d. q, Ptbp1 overexpression was verified by qRT–PCR (mean ± s.d.). u, Expression levels of representative cardiac (left axis, Tnnt2, Actc1 and Ryr2) and fibroblast (right axis, Col3a1) markers were determined by qRT–PCR (mean ± s.d.). Mock, untransduced cardiac fibroblasts. Where appropriate, a two-sided Student’s t-test or one-way ANOVA followed by Bonferroni correction (two-sided) was performed. *P < 0.05, **P < 0.01, ***P < 0.001; ns, not significant.
Extended Data Figure 9 Splicing re-patterning and transcriptome shift underlying shPtbp1-mediated enhancement of iCM reprogramming, and correlation of Mef2c, Gata4 and Tbx5 expression and reprogramming.
a–n, Splicing analyses of day-3 reprogramming cells upon Ptbp1 silencing. Related to Fig. 3j–q. a, Number of overlapping and non-overlapping alternative splicing events identified between MGT versus LacZ and MGT and shPtbp1 versus MGT and shNT. The minimal overlap suggests that Ptbp1 knockdown caused extensive re-patterning of the splicing landscape during iCM induction. b, Number of exon-skipping events that skip (grey) or include (red) the exon more in MGT and shPtbp1 compared to MGT and shNT. c, Gene ontology analysis of alternative splicing genes between MGT and LacZ. d, e, A total of 138 alternative splicing events (83 genes) between MGT and shPtbp1 and MGT and shNT were reported by rMATS to be the most significant (P value = 0, which was actually <1 × 10−16). d, Top 10 events ranked by junction counts include four events on tropomyosin genes (Tpm1 and Tpm2). Tropomyosins are essential genes for muscle contraction and they are known to undergo extensive alternative splicing40. The most studied tropomyosin alternative splicing events have been mutually exclusive exons and it is interesting to find two exon-skipping events of Tpm1 as the top 2 alternative splicing events upon Ptbp1 knockdown during reprogramming. AS, alternative splicing event; A3SS, alternative 3′ splicing site; ES, exon-skipping event; IR, intron retention. e, Gene ontology analysis of the most significant alternative splicing genes. f–i, Sashimi plots for the rest of the representative alternative splicing events in the blue-labelled gene ontology terms in Fig. 3n. The event shown in Fig. 3o was for Mbnl, which is an essential splicing factor for cardiac function that switches isoforms during heart development41. The event shown in h was an exon-skipping event in Tpm1 exon 3 (exon 2b in older literature), which was also the top event in d. This exon-3-skipped isoform of Tpm1 (Tpm1α) is the one enriched in cardiac and striated muscle cells, regulating the assembly and functionality of actin filament for contraction42. j, Overlap of alternative splicing genes and differentially expressed genes between MGT and shPtbp1 versus MGT and shNT cells. k, Overlap of differentially expressed genes between MGT versus LacZ and MGT and shPtbp1 versus MGT and shNT. l, Percentage of overlapping differentially expressed genes in k, showing the same or opposite direction of changes. m–o, On the basis of the results shown in k, top gene ontology terms for differentially expressed genes between MGT and LacZ (m), overlapping differentially expressed genes (n) and differentially expressed genes (DEG) between MGT and shPtbp1 and MGT and shNT (o) are shown. p–s, Correlation between Mef2c, Gata4 and Tbx5 expression and reprogramming. Related to Fig. 4a. p, Correlation between the total expression of M, G, and T in individual cells and the SLICER-calculated reprogramming progress of each cell. Trend line and the correlation coefficient by linear regression are shown (P = 3.9 × 10−78, α = 0.05, two-sided). q, r, Expression levels of Mef2c, Gata4 and Tbx5 in fibroblasts, intermediate fibroblasts, pre-iCMs and iCMs plotted as mean ± s.e.m. (q) or violin plots to show distribution (r). s, Ratios of expression levels of Mef2c, Gata4 and Tbx5 in the four cell groups. Data are mean ± s.e.m. t, u, Spearman correlation between Mef2c, Gata4 and Tbx5 expression and the expression of 178 known and predicted splicing factors34 or 1,602 additional transcription factors12. Genes with correlation coefficient >0.3 or <–0.3 with one or more of Mef2c, Gata4 and Tbx5 were selected and the intercorrelation matrix of the 17 selected splicing factors (t) and the 65 selected transcription factors (u) were calculated and plotted as a heat map. The splicing factors Mbnl1 and Rbms3 are strongly anti-correlated with expression of Mef2c, Gata4 and Tbx5, and Rbm20 is the only factor that is positively correlated with expression Mef2c, Gata4 and Tbx5 (P < 1 × 10−7 by two-sided Spearman correlation, α = 0.05). In u, two sets of genes, A and B, were found to be strongly anti-correlated with Mef2c, Gata4 and Tbx5 expression and that were also strongly co-expressed. These genes include Id1, Id2, Id3, Tcf21 and Foxp1 (P < 1 × 10–15 by two-sided Spearman correlation, α = 0.05) that might serve as ‘secondary’ key factors that further trigger the activation/inhibition of downstream cascades for successful conversion from fibroblasts to iCMs.
Extended Data Figure 10 Putative markers for iCMs and pre-iCMs.
Analysis using an ANOVA identified 7,624 differentially expressed genes among fibroblasts, intermediate fibroblasts, pre-iCMs and iCMs. There were 954 and 285 candidates for negative and positive selection markers of iCMs and 55 candidates for positive markers of pre-iCMs. These candidates were expressed the lowest and highest in iCMs and highest in pre-iCMs, respectively. No gene passed the selection criteria for negative markers of pre-iCMs. Top candidates were selected by largest fold change in expression in the cell population of interest compared to the expression in fibroblasts. a, Violin plots showing the expression of non-surface genes in top 30 candidates for negative markers of iCMs. Related to Fig. 4d. b–e, Top 30 candidates for positive selection markers of iCMs (b, c) or pre-iCMs (d, e). b, d, Fold change in gene expression in iCM/fibroblast (b) or pre-iCM/fibroblast (d). c, e, Violin plots of the same genes in four cell populations. f, Top 30 genes showing largest expression fold change in pre-iCMs and iCMs. g–n, Effect of Cd200 knockdown (g–j) or overexpression (k–n) on iCM reprogramming. Cardiac fibroblasts were untransduced (mock) or simultaneously transduced with MGT and lentiviral shNT or shCd200 or LacZ and OE-Cd200 for 14 days. Knockdown or overexpression efficiency was verified by qRT–PCR (g, k). Data are mean ± s.d. n = 3 samples. Percentages of α-MHC–GFP+, cTnT+ and Cx43+ cells were determined by immunostaining followed by imaging and blinded quantification (h, i, l, m) with representative 20× images in h, l. Scale bars, 200 μm. n = 20 (i) or 10 (m) images. Data are mean ± s.e.m. Percentages of α-MHC–GFP+, cTnT+ and double-positive cells were also quantified by flow cytometry (j, n). Data are mean ± s.d. n = 3 samples. Two-sided Student’s t-test was used. ***P < 0.001; ns, not significant.
Supplementary information
Supplementary Information
This file contains Supplementary Discussions 1–3 and Supplementary Figure 1, the raw images of the western blots.
Supplementary Tables
This file contains Supplementary Tables 1–8.
Source data
Rights and permissions
About this article
Cite this article
Liu, Z., Wang, L., Welch, J. et al. Single-cell transcriptomics reconstructs fate conversion from fibroblast to cardiomyocyte. Nature 551, 100–104 (2017). https://doi.org/10.1038/nature24454
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24454
This article is cited by
-
A review on gene regulatory network reconstruction algorithms based on single cell RNA sequencing
Genes & Genomics (2024)
-
Cardiomyocyte precursors generated by direct reprogramming and molecular beacon selection attenuate ventricular remodeling after experimental myocardial infarction
Stem Cell Research & Therapy (2023)
-
M-type pyruvate kinase 2 (PKM2) tetramerization alleviates the progression of right ventricle failure by regulating oxidative stress and mitochondrial dynamics
Journal of Translational Medicine (2023)
-
Single-cell transcriptomics for the assessment of cardiac disease
Nature Reviews Cardiology (2023)
-
Translational landscape of direct cardiac reprogramming reveals a role of Ybx1 in repressing cardiac fate acquisition
Nature Cardiovascular Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.