Abstract
Rare genetic variants are abundant in humans and are expected to contribute to individual disease risk1,2,3,4. While genetic association studies have successfully identified common genetic variants associated with susceptibility, these studies are not practical for identifying rare variants1,5. Efforts to distinguish pathogenic variants from benign rare variants have leveraged the genetic code to identify deleterious protein-coding alleles1,6,7, but no analogous code exists for non-coding variants. Therefore, ascertaining which rare variants have phenotypic effects remains a major challenge. Rare non-coding variants have been associated with extreme gene expression in studies using single tissues8,9,10,11, but their effects across tissues are unknown. Here we identify gene expression outliers, or individuals showing extreme expression levels for a particular gene, across 44 human tissues by using combined analyses of whole genomes and multi-tissue RNA-sequencing data from the Genotype-Tissue Expression (GTEx) project v6p release12. We find that 58% of underexpression and 28% of overexpression outliers have nearby conserved rare variants compared to 8% of non-outliers. Additionally, we developed RIVER (RNA-informed variant effect on regulation), a Bayesian statistical model that incorporates expression data to predict a regulatory effect for rare variants with higher accuracy than models using genomic annotations alone. Overall, we demonstrate that rare variants contribute to large gene expression changes across tissues and provide an integrative method for interpretation of rare variants in individual genomes.
Similar content being viewed by others
Main
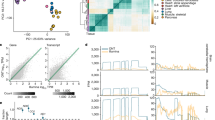
Our analysis focused on individuals with extremely high or extremely low expression of a particular gene compared with the population, using the GTEx v6p release data, which include RNA-sequencing data for 449 individuals and 44 tissues. We refer to these individuals as gene expression outliers. The GTEx data enable the identification of both single-tissue and multi-tissue expression outliers (Fig. 1a), with the latter defined by consistent extreme expression across many tissues (see Methods). To account for broad environmental and technical confounders, we removed hidden factors estimated by PEER (probabilistic estimation of expression residuals)13 from each tissue before outlier discovery (Extended Data Figs 1, 2 and Supplementary Tables 1, 2).
a, A multi-tissue outlier. The individual has extreme expression values for the gene AKR1C4 in multiple tissues (red arrows) and the most extreme median expression value across tissues. b, Outlier expression sharing between tissues, as measured by the proportion of single-tissue outliers that have a |Z-score| ≥ 2 with the same effect direction for the corresponding genes in each replication tissue. Tissues are hierarchically clustered by gene expression. c, Estimated replication rate of multi-tissue outliers in a constant held-out set of tissues for different sets of discovery tissues.
We identified a single-tissue expression outlier for ≥99% of expressed genes in each tissue and a multi-tissue outlier for 4,919 out of 18,380 genes that were tested (27%). Each individual was a single-tissue outlier for a median of 83 genes per tissue and a multi-tissue outlier for a median of 10 genes. Single-tissue outliers that were found in one tissue replicated in other tissues at rates of up to 33%, with higher rates among related tissues (Fig. 1b and Extended Data Fig. 3). The replication rate for multi-tissue outliers was much higher and increased with the number of tissues used for discovery (Fig. 1c).
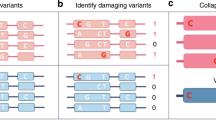
We investigated the influence of rare genetic variation on extreme expression levels, focusing on the individuals of European ancestry with whole-genome sequencing data (1,144 multi-tissue outliers). Multi-tissue outliers were strongly enriched for nearby rare variants. The enrichment was most pronounced for structural variants, as previously described14, and greater for short insertions and deletions (indels) than for single-nucleotide variants (SNVs) (Fig. 2a and Extended Data Fig. 4). Because most rare variants occur as heterozygotes, expression outliers driven by rare variants in cis should exhibit allele-specific expression (ASE). Both single-tissue and multi-tissue outliers were significantly enriched for ASE compared to non-outliers (see Methods; two-sided Wilcoxon rank-sum tests, each nominal P < 2.2 × 10−16; Fig. 2c). For underexpression outliers with exonic rare variants, the rare allele was generally underexpressed with respect to the common allele and conversely so for overexpression outliers, consistent with the rare variant causing the effect (two-sided Wilcoxon rank-sum tests, each nominal P < 4.0 × 10−8; Extended Data Fig. 5a). The enrichment for rare variants and ASE was stronger for multi-tissue outliers than for single-tissue outliers (Fig. 2b, c and Extended Data Fig. 6a), especially at higher Z-score thresholds.
a, Enrichment of SNVs, indels and structural variants (SVs) within 10 kb of the transcription start site (TSS) among outliers. For each frequency stratum, we calculated enrichment as the relative risk of having a nearby rare variant given the outlier status (see Methods). Lines indicate 95% Wald confidence intervals of the relative risk estimates. b, Rare SNV enrichments at increasing Z-score thresholds. Text labels indicate the number of outliers at each threshold. c, ASE at increasing Z-score thresholds. ASE is measured as the magnitude of the difference between the reference-allele ratio and the null expectation of 0.5. The non-outlier category is defined in the Methods.
To characterize the properties of rare variants that correlated with large changes in gene expression, we assessed the enrichment of different classes of variants in outliers compared to non-outliers (Supplementary Table 3a). Outliers were enriched, in order of significance, for structural variants, variants near splice sites, introducing frameshifts, at start or stop codons, near the transcription start site and in conserved regions (Fig. 3a). Variants in coding regions contributed disproportionately to outlier expression; enrichments weakened for all variants types (SNVs, indels and structural variants) when excluding exonic regions (Extended Data Fig. 6b). Additionally, 90% of stop-gain and frameshift variants were predicted to trigger nonsense-mediated decay in outliers (see Methods), suggesting a biological mechanism for these cases.
We considered rare variants in the gene body and within 10 kb of the gene (200 kb for structural variants and enhancers). a, Enrichment of disjoint variant classes among outliers calculated as the log odds ratio with 95% Wald confidence intervals. b, Enrichment of functional annotations for rare SNVs. c, Proportion of genes with an outlier potentially explained by each rare variant class. d, Distribution of median Z-scores for each variant class. e, For each variant class, distribution of ASE (see Methods) averaged across tissues. Grey lines mark the median values among non-outliers.
We also tested the relationship between outlier gene expression and functional annotations. Multi-tissue outliers were strongly enriched for variants in promoter or CpG-rich regions and had variants with higher conservation15,16,17,18 and CADD (combined annotation-dependent depletion)19 scores than non-outliers. We observed weaker enrichment in enhancers and transcription-factor-binding sites (Fig. 3b and Extended Data Fig. 7). Combining all classes of variation, other than non-conserved, non-coding, rare variants (excluded as less likely candidates for causal effects), we observed that 58% of underexpression and 28% of overexpression outliers had rare variants near the relevant gene, compared to 8% for non-outliers (Fig. 3c). Overexpression outliers were more common overall, potentially because detection of underexpression outliers for very low expression genes is inherently limited (Extended Data Fig. 5b). Overexpression outliers were also less enriched for functionally annotated rare variants (Extended Data Fig. 5c). Some variant classes had strong directionality concordant with their expected impact: duplications caused overexpression, whereas deletions, start- and stop-codon variants and frameshifts coincided with underexpression (Fig. 3d). We also observed strong ASE for outliers carrying all classes of variants, except non-conserved variants (Fig. 3e).
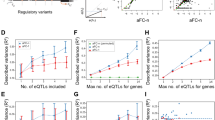
We hypothesized that functional, large-effect rare variants have been under recent selective pressure. As expected, we found that rare promoter variants of outliers were significantly less frequent in the UK10K cohort of 3,781 individuals3 than rare promoter variants of non-outliers for the same genes (two-sided Wilcoxon rank-sum test, P = 0.0060; Fig. 4a). Additionally, genes intolerant to loss-of-function and missense mutations were depleted of both multi-tissue outliers and multi-tissue expression quantitative trait loci (eQTLs; Fisher’s exact test, all P < 2 × 10−15; Fig. 4b and Extended Data Fig. 8a). We observed a similar depletion in two curated disease gene lists—genes involved in heritable cardiovascular disease and genes in the guidelines of the American College of Medical Genetics and Genomics for incidental findings20—but not in broader gene lists (Fig. 4c and Extended Data Fig. 8b, c). Genes with a multi-tissue outlier were more likely to have a multi-tissue eQTL (two-sided Wilcoxon rank-sum test, P < 2.2 × 10−16; Extended Data Fig. 8d, e), suggesting that rare and common regulatory variation influence similar genes. However, we found evidence that genes with outliers were more constrained than genes with multi-tissue eQTLs, because genes with outliers had less missense and loss-of-function variation (Tukey’s range test, missense Z-score P = 0.0070, probability of loss-of-function intolerance score P = 0.032; Fig. 4b and Extended Data Fig. 8a). This suggests that outlier expression analysis can yield unique insights into constraints on gene regulation.
a, Distributions of UK10K minor allele frequencies for promoter SNVs in outlier and non-outlier individuals at genes with multi-tissue outliers. b, Odds ratio of being intolerant to loss-of-function variants for genes with multi-tissue outliers, genes with shared eQTLs (eGenes), genes reported in the genome-wide association study (GWAS) catalogue and Online Mendelian Inheritance in Man (OMIM) genes. c, Odds ratio of a gene having a multi-tissue outlier for each of eight sets of genes involved in complex traits or diseases (gene lists are described in the Methods; DDG2P: Developmental Disorders Genotype-to-Phenotype). In b and c, lines represent 95% confidence intervals (Fisher’s exact test).
Next, we sought to prioritize rare variants in each individual genome by their predicted impact on gene expression. We developed RIVER (RNA-informed variant effect on regulation), a Bayesian statistical model that jointly analyses genome and transcriptome data from the same individual to estimate the probability that a variant has regulatory impact (https://bioconductor.org/packages/release/bioc/html/RIVER.html, see Methods). RIVER uses a generative model that assumes that genomic annotations (Supplementary Table 3b) determine the prior probability that a variant is a functional regulatory variant, in terms of influence on gene expression, which in turn affects whether nearby genes are likely to display outlier levels of expression (Fig. 5a). RIVER does not require a labelled set of functional/non-functional variants; rather it derives its power from identifying expression patterns that coincide with predictive genomic annotations.
a, RIVER probabilistic graphical model (see Methods). b, Predictive power of RIVER compared to an L2-regularized logistic regression model using only genomic annotations. Accuracy was assessed using held-out individuals, who shared the same rare SNVs as observed individuals (AUCs compared with DeLong’s approach29). c, Distribution of RIVER scores (shades of blue) as a function of expression and genomic annotation scores. The distributions of variant categories across expression and genomic annotation scores are shown as histograms aligned opposite the corresponding axes.
We trained RIVER on the GTEx v6p cohort, and evaluated the model on held-out pairs of individuals who shared the same rare variants. We then computed the RIVER score (the posterior probability of having a functional regulatory variant) for one individual, using both expression and genomic data, and assessed the accuracy with respect to the expression levels of the second individual that had been held out (see Methods). Incorporating expression data significantly improved prediction compared with a model that uses genomic annotations alone (area under the curve (AUC) of 0.64 and 0.54, respectively, P = 3.5 × 10−4; Fig. 5b and Extended Data Fig. 9a, b), and RIVER learned, unsupervised, to prioritize variants supported by both genomic annotations and extreme expression levels across tissues (Fig. 5c and Extended Data Fig. 9c). ASE was also enriched among the top RIVER hits compared with the genomic annotation model (Extended Data Fig. 9d). Finally, even after accounting for the most informative genomic annotations or summary scores, personal expression data were highly informative of rare variant effects (average log odds ratio, 2.76; Extended Data Fig. 9e, f).
RIVER can be used to predict regulatory effects on gene expression of disease-associated variants and aid in prioritization of rare variants in disease studies. To investigate this potential, we evaluated 27 pathogenic variants from ClinVar21 present in 21 GTEx donors (Fig. 5c and Extended Data Fig. 10a). Overall, pathogenic variants had RIVER scores that were higher than background variants (two-sided Wilcoxon rank-sum test, P = 3.3 × 10−9; Extended Data Fig. 10b–d), and the six that were probably regulatory variants (those not annotated as missense or as an indel within a coding region) scored in the 99.9th percentile. Several cases, which we evaluated in detail, illustrated that rare disease-causing variants can have a regulatory impact evident from RNA-sequencing data, even from healthy individuals that have those variants (in whom the variants are often heterozygous; Extended Data Fig. 10e, f). Note that RIVER trained on healthy cohorts, such as GTEx, can then be directly applied to new cohorts that include disease samples.
To experimentally validate a subset of the variants that were identified through outlier analysis, we used CRISPR–Cas9-mediated genome editing22,23. In K562 cells, we tested six SNVs and matched controls in transcribed regions of genes with an outlier (see Methods and Extended Data Fig. 11a, b), and compared the allelic ratios between mRNA and genomic DNA (gDNA), which was used as an internal control. All variants that were tested were SNVs in underexpression outliers and were therefore expected to decrease expression. Two variants were excluded owing to low cDNA and gDNA total reads counts. The four remaining SNVs in outliers all showed lower proportions of the alternate (installed) allele in the cDNA compared to the gDNA, confirming that these variants decreased expression (Extended Data Fig. 11c).
In summary, by combining data across multiple tissues, we curated a set of gene expression outliers that replicated at higher rates and showed stronger enrichment of rare variants than those from any single tissue. We found that rare structural variants, frameshift indels, coding variants and variants near the transcription start site were most likely to have large effects on expression. However, our ability to characterize the genetic basis of multi-tissue outliers remains incomplete. Outliers without an underlying rare variant in our analysis may be due to variants in more distal regions or in annotations we did not consider, or may be attributable to residual technical or environmental effects.
Although variant interpretation remains challenging, RIVER demonstrates the value of incorporating personal gene expression data to examine the consequences of rare variants that may be uncertain based on the sequence alone. Our results suggest that a general approach can be applied to studies that supplement genome sequencing with other molecular phenotypes, such as methylation24,25,26 and histone modification27,28. We anticipate that such integrative approaches will be essential for effective interpretation of genome-wide genetic variation on a personalized level.
Methods
Study population
All human subjects were deceased donors. Informed consent was obtained for all donors via next-of-kin consent to permit the collection and banking of de-identified tissue samples for scientific research. The research protocol was reviewed by Chesapeake Research Review Inc., Roswell Park Cancer Institute’s Office of Research Subject Protection, and the institutional review board of the University of Pennsylvania. We used the RNA-seq, allele-specific expression, and whole-genome sequencing (WGS) data from the v6p release of the GTEx project. The generation of these data are described in the supplementary information of ref. 12.
Correction for technical confounders
We restricted our expression analyses to the 449 individuals and 44 tissues for which sex and the top three genotype principal components, which capture major population stratification, were available. For each tissue, we log2-transformed all expression values (log2(RPKM + 2)), where RPKM is the number of reads per kilobase of transcript per million mapped reads. We then standardized the expression of each gene to prevent shrinkage of outlier expression values caused by quantile normalization. To remove unmeasured batch effects and other confounders, for each tissue separately, we estimated hidden factors using PEER13 on the transformed expression values. In each tissue, we defined expressed genes and corrected for the same number of PEER factors as in the GTEx eQTL analyses (see supplementary information of ref. 12). We regressed out the PEER factors, the top three genotype principal components and sex (where appropriate) from the transformed expression data for each tissue using the following linear model:

where Yg is the transformed expression of a given gene g, μg is the mean expression level for the gene, Pn is the nth PEER factor, G1, G2, G3 are the top three genotype principal components, and S is the sex covariate. We assumed the residual vector εg follows the multivariate normal distribution εg ~ N(0, σ2I). Finally, we standardized the expression residuals εg for each gene, which yielded Z-scores.
To better understand the effect of PEER correction on the removal of technical and biological confounders, we compared the PEER factors in each tissue separately to pre-collected sample and subject covariates. We considered the subset of covariates with >50 observations in at least 31 tissues, where we first selected covariates with more than one unique entry in each tissue. For categorical covariates, we only considered categories with more than 20 observations. For each PEER factor and each covariate, we fit a linear model with the PEER factor as the response and the covariate as the predictor. From this model, we computed the proportion of that PEER factor’s variance explained by the covariate as the adjusted R2:

where p and n are the number of parameters and samples, respectively, and

SST and SSR refer to the total and residual sums of squares, respectively.
To quantify the degree to which each covariate was captured by the combination of all PEER factors, genotype principal components and sex (where appropriate) for each tissue, we considered the expression component regressed out from the uncorrected data:

For each covariate, we then fit a linear model with Wg as the response and the covariate as the predictor. We assessed the proportion of the variance of Wg explained by each covariate by computing the adjusted R2 for the covariate across all genes. We used the formula above, but summed across all genes to compute SST and SSR.
To assess the impact of PEER correction on rare variant enrichment, we also tried removing either the top five PEER factors for each tissue or no PEER factors. We then performed multi-tissue outlier calling and tested the enrichment of rare and common variants in the two partially corrected datasets (see ‘Enrichment of rare and common variants near outlier genes’).
Single-tissue and multi-tissue outlier discovery
Single-tissue and multi-tissue outlier calling was restricted to autosomal lincRNA and protein-coding genes. For each tissue, an individual was called a single-tissue outlier for a particular gene if that individual had the largest absolute Z-score and the absolute value was at least 2. For each gene, the individual with the most extreme median Z-score taken across tissues was identified as a multi-tissue outlier for that gene provided the absolute median Z-score was at least 2. Therefore, each gene had at most one single-tissue outlier per tissue and one multi-tissue outlier. Under this definition an individual could be an outlier for multiple genes. In addition, we only tested for multi-tissue outliers among individuals with expression measurements for the gene in at least five tissues. To reduce cases where non-genetic factors may cause widespread extreme expression, we removed eight individuals that were multi-tissue outliers for 50 or more genes from all downstream analyses, including before single-tissue outlier discovery. Removing these individuals with extreme expression across many genes improved our rare variant enrichments, but the precise threshold mattered less (Extended Data Fig. 2g). We chose the threshold of 50 to strike a balance between removing extreme individuals while not excluding a large proportion of our cohort.
Replication of expression outliers
We calculated the proportion of single-tissue outliers discovered in one tissue that had |Z-score| ≥ 2 with the same direction of effect for the same gene in the replication tissue. Since certain groups of tissues were sampled in a specific subset of individuals, we evaluated the extent to which replication was influenced by the size and the overlap of the discovery and replication sets. We repeated the replication analysis with the discovery and replication in exactly 70 overlapping individuals for each pair of tissues with enough samples and compared the replication patterns to those obtained by using all individuals. To estimate the extent to which individual overlap biased replication estimates, for each pair of tissues with sufficient samples, we defined three disjoint groups of individuals: 70 individuals with data for both tissues, 69 distinct individuals with data in the first tissue, and 69 distinct individuals with data in the second tissue. We discovered outliers in the first tissue using the shared set of individuals then tested for replication using the same individuals in the second tissue. Then, for each gene, we added the identified outlier to the distinct set of individuals and tested the replication again in the second tissue. We repeated the process running the discovery in the second tissue and the replication in the first one. We compared the replication rates when using the same or different individuals for the discovery and replication.
We assessed the confidence of our multi-tissue outliers using cross-validation. We separated the tissue expression data randomly into two groups: a discovery set of 34 tissues and a replication set of 10 tissues. For t = 10, 15, 20, 25, and 30, we randomly sampled t tissues from the discovery set and performed outlier calling as described above. Owing to incomplete tissue sampling, the number of tissues supporting each outlier is at least five but less than t. We computed the replication rate as the proportion of outliers in the discovery set with |median Z-score| ≥ 1 or 2 in the replication set. We set no restriction on the number of tissues required for testing in the replication set. To calculate the expected replication rate, we randomly selected individuals in the discovery set with at least five tissues that expressed the gene and computed the replication rate. We repeated this process 10 times for each discovery set size.
Quality control of genotypes and rare variant definition
We restricted our rare variant analyses to individuals of European descent, as they constituted the largest homogenous population within our dataset. We considered only autosomal variants that passed all filters in the VCF (those marked as PASS in the Filter column). Minor allele frequencies (MAFs) within the GTEx data were calculated from the 123 individuals of European ancestry with WGS data (average coverage 30×). The MAF was the minimum of the reference and the alternate allele frequency where the allele frequencies of all alternate alleles were summed together. Rare variants were defined as having MAF ≤ 0.01 in GTEx, and for SNVs and indels we also required MAF ≤ 0.01 in the European population of the 1000 Genomes Project Phase 3 data30. To ensure that population structure among the individuals of European descent was unlikely to confound our results, we verified that the allele frequency distribution of rare variants included in our analysis (within 10 kb of a protein-coding or lincRNA gene, see below) was similar for the five European populations in the 1000 Genomes Project (Extended Data Fig. 4d).
Enrichment of rare and common variants near outlier genes
We assessed the enrichment of rare SNVs, indels and structural variants near outlier genes. Proximity was defined as within 10 kb of the transcription start site for most analyses. For Fig. 3 and Extended Data Figs 5, 7, 8, we included all variants within 10 kb of the gene, including the gene body, to also capture coding variants. In Fig. 3 and Extended Data Figs 5, 8, we extended the window to 200 kb for enhancers and structural variants. For each gene with an outlier, we chose the remaining set of individuals tested for outliers at the same gene as non-outlier controls. We only considered genes that had both an outlier and at least one control. We stratified variants of each class into four minor allele frequency bins (0–1%, 1–5%, 5–10%, 10–25%) to compare the relative enrichments of rare and common variants. We also assessed the enrichment of SNVs at different Z-score cutoffs. Enrichment was defined as the ratio of the proportion of outliers with a variant whose frequency lies within the range to the corresponding proportion for non-outliers. This enrichment analysis is equivalent to the relative risk of having a nearby rare variant given outlier status. We used the asymptotic distribution of the log relative risk to obtain 95% Wald confidence intervals. Within our set of European individuals, we observed some individuals with minor admixture that had relatively more rare variants than the rest (Extended Data Fig. 1b). We confirmed that inclusion of these admixed individuals did not substantially affect our results (Extended Data Fig. 1c). We also calculated rare variant enrichments when restricting to variants outside protein-coding and lincRNA exons in the Gencode v.19 annotation (extending internal exons by 5 bp to capture canonical splice regions).
To measure the informativeness of variant annotations, we used logistic regression to model outlier status as a function of the feature of interest; this yielded log odds ratios with 95% Wald confidence intervals. Note that for the feature enrichment analysis in Fig. 3b and Extended Data Fig. 7, we required that outliers and their gene-matched non-outlier controls have at least one rare variant near the gene. We standardized all features, including binary features, to facilitate comparison between features of different scale. We also calculated the proportion of overexpression outliers, underexpression outliers and non-outliers with a rare variant near the gene (within 10 kb for SNVs and indels and 200 kb for structural variants). To each outlier instance, we assigned at most one of the 12 rare variant classes that we considered (Supplementary Table 3a). If an outlier had rare variants from multiple classes near the relevant genes, we selected the class that was most significantly enriched among outliers.
Annotation of variants
We obtained structural variant annotations from ref. 14 and computed features for rare SNVs and indels using three primary data sources: Roadmap Epigenomics31, CADD v.1.2 (ref. 19) and VEP v.80 (ref. 32). Promoter and enhancer annotation tracks were obtained from the Roadmap Epigenomics Project (http://www.broadinstitute.org/~meuleman/reg2map/HoneyBadger2_release/). We mapped 28 unique tissues in the GTEx project to 19 tissue groups in the Roadmap Project. Using these annotations, for each individual, we assessed whether each SNV or indel overlapped a promoter or enhancer region in at least one of the 19 Roadmap tissue groups. Features, including conservation15,16,17,18, transcription factor binding and deleteriousness, were extracted from the full annotation tracks of the CADD v.1.2 release (downloaded 15 May 2015; http://cadd.gs.washington.edu/download). Finally, we obtained protein-coding and transcription-related annotations from VEP and LOFTEE. This information was provided in the GTEx v6p VCF file (described in ref. 12). Stop-gain and frameshift variants annotated as high-confidence loss-of-function variants by LOFTEE were assumed to trigger nonsense-mediated decay. We generated gene-level features described in Supplementary Table 3.
Allele-specific expression (ASE)
We only considered sites with at least 30 total reads and at least five reads supporting each of the reference and alternate alleles. To minimize the effect of mapping bias, we filtered out sites that showed mapping bias in simulations33, that were in low mappability regions (ftp://hgdownload.cse.ucsc.edu/gbdb/hg19/bbi/wgEncodeCrgMapabilityAlign50mer.bw) or that were rare variants or within 1 kb of a rare variant in the given individual (the variants were extracted from the GTEx exome-sequencing data described in ref. 12). The first two filters were provided in the GTEx ASE data release. The third filter was applied to eliminate potential mapping artefacts that mimic genetic effects from rare variants. We measured ASE at each testable site as the absolute deviation of the reference-allele ratio from 0.5. For each gene, all testable sites in all tissues were included. We compared ASE in single-tissue and multi-tissue outliers at different Z-score thresholds to non-outliers using two-sided Wilcoxon rank-sum tests. To obtain a matched background, we only included a gene in the comparison when ASE data existed for both the outlier individual and at least one non-outlier. In the case of single-tissue outliers, we also required the tissue to match between the outlier and the non-outlier. All individuals that were neither multi-tissue outliers for the given gene nor single-tissue outliers for the gene in the corresponding tissue were included as non-outliers.
In cases where outliers had rare coding variants in the gene, if the rare variants were causing the extreme expression in cis, we expected to see ASE at the rare variant matching the direction of the effect. For underexpression outliers, we expected the (rare) minor allele to be underexpressed compared to the major allele. For overexpression outliers, we expected the minor allele to be overexpressed. To test this, we used the same filters as above, but looked exclusively at rare variants (instead of excluding them). We measured ASE as the minor-allele ratio: the number of reads supporting the minor allele over the total number of reads.
We also used ASE to evaluate the performance of both the genomic annotation model and RIVER (see below) by testing the association between allelic imbalance and model predictions using Fisher’s exact test. Here, we defined allelic imbalance as the top 10% of the median absolute deviation, across tissues, of the reference-allele ratio from 0.5.
Allele frequency measurements in UK10K
UK10K3 VCF files of whole-genome cohorts were downloaded from https://www.ebi.ac.uk. We merged the Avon Longitudinal Study of Parents and Children (ALSPAC) EGAS00001000090 and the Department of Twin Research and Genetic Epidemiology (TWINSUK) EGAS00001000108 datasets for a total of 3,781 individuals. We counted the occurrence of all rare GTEx SNVs in Roadmap Epigenomics-annotated promoter regions among the UK10K samples. GTEx variants absent from the UK10K cohorts were assigned a count of 0.
Definition of multi-tissue eGenes
We defined multi-tissue eGenes using two approaches. For the tissue-by-tissue approach, we obtained lists of significant eGenes (q value ≤ 0.05) for each of the 44 tissues from the GTEx v6p release. The second approach used cis-eQTLs with shared effects across tissues estimated by the RE2 model of the Meta-Tissue software34, as described in ref. 12. We chose, for each gene, the variant with the lowest nominal P value from the RE2 model. We then determined the number of tissues in which this variant-gene pair showed a cis-eQTL effect (m value ≥ 0.9 (ref. 34)). For each of the 18,380 genes tested for multi-tissue outliers, we calculated the number of tissues in which the gene appeared as a significant eGene (tissue-by-tissue approach) or had a shared eQTL effect (Meta-Tissue approach). To show that the enrichment of outlier genes as multi-tissue eGenes was not confounded by gene expression level, using the Meta-Tissue results, we stratified genes tested for multi-tissue outliers into RPKM deciles and repeated the comparison between genes with and without a multi-tissue outlier. When comparing the enrichment for eGenes among constrained and disease gene lists, we classified the top n Meta-Tissue eGenes (ranked by nominal P value from the RE2 model) as multi-tissue eGenes and considered the remaining genes as background. We selected n to match the number of multi-tissue outliers in the comparison.
Evolutionary constraint of genes with multi-tissue outliers
We obtained gene-level estimates of evolutionary constraint from the Exome Aggregation Consortium35 (http://exac.broadinstitute.org/,ExACreleasev.0.3). We intersected the 17,351 autosomal lincRNA and protein-coding genes with constraint data from ExAC with the 18,380 genes tested for multi-tissue outliers from GTEx, yielding 14,379 genes for further analysis (3,897 and 10,482 genes with and without a multi-tissue outlier, respectively). We examined three functional constraint scores from the ExAC database: synonymous Z-score, missense Z-score and probability of loss-of-function intolerance (pLI). Synonymous- and missense-intolerant genes were defined as those with corresponding Z-scores above the 90th percentile. We defined loss-of-function intolerant genes as those with a pLI score above 0.9, following the guidelines provided by ExAC. We calculated odds ratios and 95% confidence intervals for the enrichment of genes with multi-tissue outliers in these lists using a Fisher’s exact test. We repeated this analysis for three other gene sets: 19,182 multi-tissue eGenes from GTEx v6p defined using Meta-Tissue, 9,480 reported GWAS genes from the NHGRI-EBI catalogue36 (http://www.ebi.ac.uk/gwas, accessed 30 November 2015) and 3,576 OMIM genes (http://omim.org/, accessed 26 May 2016).
We tested for a difference in the mean constraint for genes with multi-tissue outliers and genes with multi-tissue eQTLs using ANOVA. For each constraint score in ExAC, we treated the score for each gene as the response and the status of the gene as having a multi-tissue outlier and/or a multi-tissue eQTL as a categorical predictor with four classes. After fitting the model, we performed a Tukey’s range test to determine whether there was a significant difference in the mean constraint between genes with a multi-tissue outlier but no multi-tissue eQTL and genes with a multi-tissue eQTL but no multi-tissue outlier.
Overlap of genes with multi-tissue outliers and disease genes
We examined the enrichment of genes with multi-tissue outliers in eight disease gene lists: the GWAS catalogue and OMIM (described above), as well as ClinVar (6,279 genes; http://www.ncbi.nlm.nih.gov/clinvar/), OrphaNet (3,451 genes; http://www.orpha.net/), ACMG20 (58 genes; http://www.ncbi.nlm.nih.gov/clinvar/docs/acmg/), Developmental Disorders Genotype-to-Phenotype37 (DDG2P; 1,693 genes; http://www.ebi.ac.uk/gene2phenotype/), and two curated gene lists of 86 cardiovascular disease genes and 55 cancer genes (described below). We computed odds ratios and 95% confidence intervals using a Fisher’s exact test to compare each disease gene list to the genes with multi-tissue outliers and repeated the comparison for genes with multi-tissue eQTLs.
Heritable cancer predisposition and heritable cardiovascular disease gene lists were curated by local experts in clinical and laboratory-based genetics in the two respective areas (Stanford Medicine Clinical Genomics Service, Stanford Cancer Center’s Cancer Genetics Clinic and Stanford Center for Inherited Cardiovascular Disease). Genes were included if both the clinical and laboratory-based teams agreed there was sufficient published evidence to support using variants in these genes in clinical decision making.
For each of the eight disease gene lists above and for genes with multi-tissue outliers or multi-tissue eQTLs, we computed the number of variants (SNVs and indels within 10 kb and structural variants within 200 kb of the gene, including the gene body) at each gene in the 123 individuals of European ancestry with WGS data. For each gene list and for each MAF bin (0–1%, 1–5%, 5–10%, 10–25%), we compared the mean number of variants near genes in the list to the mean number near all other annotated autosomal protein-coding and lincRNA genes using a two-sided t-test.
The RIVER integrative model for predicting regulatory effects of rare variants
RIVER (RNA-informed variant effect on regulation) is a hierarchical Bayesian model that predicts the regulatory effects of rare variants by integrating gene expression with genomic annotations. The RIVER model consists of three layers: a set of nodes G = G1,..., GP in the topmost layer representing P observed genomic annotations over all rare variants near a particular gene; a latent binary variable F in the middle layer representing the unobserved functional regulatory status of the rare variants; and one binary node E in the final layer representing expression outlier status of the nearby gene. We model each conditional probability distribution as follows:




with parameters β and θ and hyper-parameters λ and C.
Because F is unobserved, the RIVER log-likelihood objective over instances n = 1, …, N  is non-convex. We therefore optimize model parameters using Expectation–Maximization38 (EM) as follows:
is non-convex. We therefore optimize model parameters using Expectation–Maximization38 (EM) as follows:
In the E-step, we compute the posterior probabilities (ωn(i)) of the latent variables Fn given current parameters and observed data. For example, at the ith iteration, the posterior probability of Fn = 1 for the nth instance is



In the M-step, at the ith iteration, given the current estimates ω(i), the parameters (β(i + 1)*) are estimated as

where λ is an L2 penalty hyper-parameter derived from the Gaussian prior on β.
The parameter θ gets updated as:

where I is an indicator operator, t is the binary value of expression En, s is the possible binary values of Fn, and C is a pseudo count derived from the Beta prior on θ. The E and M steps are applied iteratively until convergence.
RIVER application to the GTEx cohort
As input, RIVER requires a set of genomic features G and a set of corresponding expression outlier observations E, each over instances of individual and gene pairs. Using the variant annotations described above, we generated site-level genomic features for the 116 European individuals with GTEx WGS data that had fewer than 50 multi-tissue outliers. We then collapsed these features for all rare SNVs within 10 kb of each transcription start site to generate the gene-level features that are described in Supplementary Table 3b. This produced a matrix of genomic features G of size (116 individuals × 1,736 genes) × (112 genomic features), where we standardized features before use. For the values of E, we defined any individual with |median Z-score| ≥ 1.5 as an outlier if expression was observed in at least five tissues; the remaining individuals were labelled as non-outliers for the gene. We used this more lenient threshold in order to obtain a sufficiently large set of outliers for robust training and testing. In total, we extracted 48,575 instances where an individual had at least one rare variant within 10 kb of the transcription start site of a gene.
To train and evaluate RIVER on the GTEx cohort, we used the 3,766 instances of individual and gene pairs where two individuals had the same rare SNVs near a particular gene. We held out those instances and trained RIVER parameters with the remaining instances. RIVER requires two hyper-parameters λ and C. To select λ, we first applied an L2-regularized multivariate logistic regression with features G and response variable E, selecting λ with the minimum squared error via tenfold cross-validation (we selected λ = 0.01). We selected C = 50, informed simply by the total number of training instances available, as validation data were not available for extensive cross-validation. Initial parameters for EM were set to θ = (P(E = 0 | F = 0), P(E = 1 | F = 0), P(E = 0 | F = 1), P(E = 1 | F = 1)) = (0.99, 0.01, 0.3, 0.7) and β from the multivariate logistic regression above, although different initializations did not significantly change the final parameters (Extended Data Fig. 9b).
The 3,766 held-out pairs of instances were used to create a labelled evaluation set. For one of the two individuals from each pair, we estimated the posterior probability of a functional rare variant P(F | G, E, β, θ). The outlier status of the second individual, whose data were not observed either during training or prediction, was then treated as a ‘label’ of the true status of functional effect F. Using this labelled set, we compared the RIVER score to the posterior P(F | G, β) estimated from the plain L2-regularized multivariate logistic regression model with genomic annotations alone. We produced receiver operating characteristic curves and computed areas under the curve (AUCs) for both models, testing for significant differences using DeLong’s method29. This analysis relied on outlier status reflecting the consequences of rare variants. Indeed, pairs of individuals who shared rare variants tended to have highly similar outlier status even after regressing out effects of common variants (Kendall’s τ rank correlation, P < 2.2 × 10−16). We repeated this evaluation, varying the median Z-score threshold used to define outliers, and we also compared RIVER to individual features that were strongly enriched among outliers as well as PolyPhen39 and SIFT40.
Supervised model integrating expression and genomic annotation
To assess the information gained by incorporating gene expression data in the prediction of functional rare variants, we applied a simplified supervised approach to a limited dataset. We used the instances where two individuals had the same rare SNVs to create a labelled training set where the outlier status of the second individual was used as the response variable. We then trained a logistic regression model with only two features: (1) the outlier status of the first individual and (2) a single genomic feature value, such as CADD or deleterious annotation of genetic variants using neural networks (DANN). We estimated parameters from the entire set of rare-variant-matched pairs using logistic regression to determine the log odds ratio and corresponding P value of expression status as a predictor. While this approach was not amenable to training a full predictive model over all genomic annotations jointly given the limited number of instances, it provided a consistent estimate of the log odds ratio of outlier status. We tested five genomic predictors: CADD19, DANN41, transcription-factor-binding site annotations, PhyloP scores15 and one aggregated feature: the posterior probability from a multivariate logistic regression model learned with all genomic annotations.
RIVER assessment of pathogenic ClinVar variants
We downloaded variants from the ClinVar database21 (accessed 04 May 2015) and searched for these disease variants within the set of rare variants segregating in the GTEx cohort. Any disease variant reported as pathogenic, likely pathogenic or a risk factor for disease was considered pathogenic. We further categorized the pathogenic variants as likely regulatory if they were annotated as splice-site variants, synonymous or nonsense, whereas missense variants were considered unlikely to have a regulatory effect. To explore RIVER scores for those pathogenic variants, all instances were used for training RIVER. We then computed a posterior probability P(F | G, E, β, θ) for each instance coinciding with a pathogenic ClinVar variant.
Stability of estimated parameters with different parameter initializations
We tried several different initialization parameters for β and θ to explore how this affected the estimated parameters. We initialized a noisy β by adding K% Gaussian noise compared to the mean of β with fixed θ (for K = 10, 20, 50 100, 200, 400, 800). For θ, we fixed P(E = 1 | F = 0) and P(E = 0 | F = 0) as 0.01 and 0.99, respectively, and initialized (P(E = 1 | F = 1), P(E = 0 | F = 1)) as (0.1, 0.9), (0.4, 0.6) and (0.45, 0.55) instead of (0.3, 0.7) with β fixed. For each parameter initialization, we computed Spearman rank correlations between parameters from RIVER using the original initialization and the alternative initializations. We also investigated how many instances within top 10% of posterior probabilities from RIVER under the original settings were replicated in the top 10% of posterior probabilities under the alternative initializations (replication accuracy in Extended Data Fig. 9b).
Validation of large-effect rare variants using CRISPR–Cas9 genome editing
To select rare, coding SNVs for validation by CRISPR–Cas9 editing, we first restricted to the (gene, individual, variant) tuples identified in multi-tissue outliers without a rare structural variant or a rare indel within 200 kb or 10 kb of the gene, respectively. We considered the 116 rare SNVs with a coding consequence for the corresponding gene as annotated by VEP32; coding annotations included stop gained, stop lost, splice acceptor variant, splice donor variant, start lost, missense variant, splice region variant, stop retained variant, synonymous variant, coding sequence variant and 5′/3′ UTR variant. Using RNA-seq data from ENCODE, we further restricted our variant list to the 59 SNVs occurring in genes with an average FPKM (fragments per kilobase per million reads) of at least 10 in K562 cells (ENCODE experiment accession numbers ENCSR000AEL and ENCSR000AEN)42. Finally, we filtered for rare, coding SNVs in (gene, individual) pairs with |median Z-score| > 4 and a RIVER score above the 99.5th percentile. These filters yielded a final set of 13 rare SNVs from which we chose the six exonic SNVs for testing.
As controls, we selected SNVs present within the same cDNA amplicon region as the corresponding outlier SNV (see details on targeted sequencing below). We first searched for coding SNVs present within these regions in the GTEx cohort that did not occur in the outlier individual. If no SNV could be found satisfying these criteria, we expanded our search for SNVs using the ExAC database (ExAC release v.0.3)35. If multiple possible control variants existed for an outlier SNV, we ranked the controls by CADD score19 and prioritized synonymous variants.
Sequences of single-guide RNAs (sgRNAs) used in the study are listed in Extended Data Fig. 11b. For each variant, a sgRNA and two donor oligonucleotides (with the reference and alternative alleles) were designed such that the PAM was located as close to the variant as possible. The donors were 99 bp long centred on the variant being installed. The variants were installed into K562 cells as previously described22,23. The K562 cells were those generated previously23 and were regularly tested for mycoplasma infection. sgRNAs were expressed in the pGH020 (Addgene plasmid 85405) expression vector. For each donor oligonucleotide, K562 cells constitutively expressing a Cas9–BFP fusion protein were electroporated with 3 μg of sgRNA plasmid DNA and 1 μl of 100 μM donor oligonucleotide using the T-016 program on a Lonza Nucleofector 2b. After electroporation, cells were allowed to recover for five days. Cells electroporated with the reference and alternative allele donor oligonucleotides were mixed in a 1:1 ratio and grown together for three more days to control for differences in culturing conditions. We included cells electroporated with the reference allele to ensure that any changes in expression we observed were not due to the editing process itself. Because the editing efficiency is not 100% and varies between loci, we expected fewer than half the cells to carry the alternative allele and for this proportion to vary by locus. One to two million cells were collected for RNA and genomic DNA extraction.
Genomic DNA (gDNA) was extracted using the QiaAmp DNA mini kit (Qiagen). Total RNA was extracted using QiaShredder and RNeasy Mini kit (Qiagen). Subsequently, 6 μg of RNA was converted into cDNA using AMV reverse transcriptase (Promega). cDNA was purified and concentrated with the PCR Purification Kit (Qiagen). PCR primers were designed to generate 300–400-bp amplicons including the variant in either the gDNA or cDNA locus. For both gDNA and cDNA samples, 400 ng of DNA was amplified in triplicate (technical replicates) using Phusion High-Fidelty polymerase (Fisher) and the amplicon was purified on a 1% TAE agarose gel. The amplicons were then prepared for sequencing using the Nextera XT kit (Illumina) and sequenced together on a NextSeq 500.
Reads were trimmed with cutadapt43 (v.1.13) and aligned using bwa44 (v.0.7.12-r1039) allowing no mismatches (bwa aln –n 0), which excluded any reads with indels created during editing. We used custom reference sequences, one each for the reference and alternate alleles of the targeted cDNA and gDNA amplicon regions. Allele counts at the target locus were computed for each sample using samtools pileup as implemented in the R package Rsamtools45 (v.1.22.0). Only reads with a minimum mapping quality of 20 were considered. Two of the tested loci amplified poorly in preparation for sequencing, and they had extremely low mapping rates and total read counts over the target locus (median read count across replicates <400 compared to 281,000 and 397,000 for gDNA and cDNA, respectively, for the remaining loci). As such, we removed these two loci from further analysis. Finally, to assess the effect of each variant on expression, we tested for a significant difference between the cDNA and gDNA alternate allele proportions with a two-sided t-test. We corrected for multiple testing using the Bonferroni procedure.
Code availability
RIVER is available at https://bioconductor.org/packages/release/bioc/html/RIVER.html. Additionally, the code for running analyses and producing the figures throughout this manuscript is available separately (https://github.com/joed3/GTExV6PRareVariation).
Data availability
The GTEx v6p release genotype and allele-specific expression data are available from dbGaP (study accession phs000424.v6.p1; http://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs000424.v6.p1). Expression data from the v6p release and eQTL results are available from the GTEx portal (http://gtexportal.org).
References
Tennessen, J. A. et al. Evolution and functional impact of rare coding variation from deep sequencing of human exomes. Science 337, 64–69 (2012)
Nelson, M. R. et al. An abundance of rare functional variants in 202 drug target genes sequenced in 14,002 people. Science 337, 100–104 (2012)
The UK10K Consortium. The UK10K project identifies rare variants in health and disease. Nature 526, 82–90 (2015)
Keinan, A. & Clark, A. G. Recent explosive human population growth has resulted in an excess of rare genetic variants. Science 336, 740–743 (2012)
Uricchio, L. H., Zaitlen, N. A., Ye, C. J., Witte, J. S. & Hernandez, R. D. Selection and explosive growth alter genetic architecture and hamper the detection of causal rare variants. Genome Res. 26, 863–873 (2016)
MacArthur, D. G. et al. A systematic survey of loss-of-function variants in human protein-coding genes. Science 335, 823–828 (2012)
Narasimhan, V. M. et al. Health and population effects of rare gene knockouts in adult humans with related parents. Science 352, 474–477 (2016)
Montgomery, S. B., Lappalainen, T., Gutierrez-Arcelus, M. & Dermitzakis, E. T. Rare and common regulatory variation in population-scale sequenced human genomes. PLoS Genet. 7, e1002144 (2011)
Zhao, J. et al. A burden of rare variants associated with extremes of gene expression in human peripheral blood. Am. J. Hum. Genet. 98, 299–309 (2016)
Zeng, Y. et al. Aberrant gene expression in humans. PLoS Genet. 11, e1004942 (2015)
Li, X. et al. Transcriptome sequencing of a large human family identifies the impact of rare noncoding variants. Am. J. Hum. Genet. 95, 245–256 (2014)
The GTEx Consortium. Genetic effects on gene expression across human tissues. https://doi.org/10.1038/nature24277 (2017)
Stegle, O., Parts, L., Piipari, M., Winn, J. & Durbin, R. Using probabilistic estimation of expression residuals (PEER) to obtain increased power and interpretability of gene expression analyses. Nat. Protoc. 7, 500–507 (2012)
Chiang, C. et al. The impact of structural variation on human gene expression. Nat. Genet. 49, 692–699 (2017)
Pollard, K. S., Hubisz, M. J., Rosenbloom, K. R. & Siepel, A. Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res. 20, 110–121 (2010)
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005)
Cooper, G. M. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913 (2005)
Arbiza, L. et al. Genome-wide inference of natural selection on human transcription factor binding sites. Nat. Genet. 45, 723–729 (2013)
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014)
Green, R. C. et al. ACMG recommendations for reporting of incidental findings in clinical exome and genome sequencing. Genet. Med. 15, 565–574 (2013)
Landrum, M. J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D862–D868 (2016)
Hendel, A. et al. Chemically modified guide RNAs enhance CRISPR–Cas genome editing in human primary cells. Nat. Biotechnol. 33, 985–989 (2015)
Hess, G. T. et al. Directed evolution using dCas9-targeted somatic hypermutation in mammalian cells. Nat. Methods 13, 1036–1042 (2016)
Grundberg, E. et al. Global analysis of DNA methylation variation in adipose tissue from twins reveals links to disease-associated variants in distal regulatory elements. Am. J. Hum. Genet. 93, 876–890 (2013)
Gamazon, E. R. et al. Enrichment of cis-regulatory gene expression SNPs and methylation quantitative trait loci among bipolar disorder susceptibility variants. Mol. Psychiatry 18, 340–346 (2013)
Bell, J. T. et al. DNA methylation patterns associate with genetic and gene expression variation in HapMap cell lines. Genome Biol. 12, R10 (2011)
Waszak, S. M. et al. Population variation and genetic control of modular chromatin architecture in humans. Cell 162, 1039–1050 (2015)
Grubert, F. et al. Genetic control of chromatin states in humans involves local and distal chromosomal interactions. Cell 162, 1051–1065 (2015)
DeLong, E. R., DeLong, D. M. & Clarke-Pearson, D. L. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44, 837–845 (1988)
1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015)
The Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015)
McLaren, W. et al. The Ensembl variant effect predictor. Genome Biol. 17, 122 (2016)
Panousis, N. I., Gutierrez-Arcelus, M., Dermitzakis, E. T. & Lappalainen, T. Allelic mapping bias in RNA-sequencing is not a major confounder in eQTL studies. Genome Biol. 15, 467 (2014)
Sul, J. H., Han, B., Ye, C., Choi, T. & Eskin, E. Effectively identifying eQTLs from multiple tissues by combining mixed model and meta-analytic approaches. PLoS Genet. 9, e1003491 (2013)
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016)
Welter, D. et al. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 42, D1001–D1006 (2014)
Deciphering Developmental Disorders Study. Large-scale discovery of novel genetic causes of developmental disorders. Nature 519, 223–228 (2015)
Dempster, A. P., Laird, N. M. & Rubin, D. B. Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. Soc. 39, 1–38 (1977)
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010)
Ng, P. C. & Henikoff, S. Predicting deleterious amino acid substitutions. Genome Res. 11, 863–874 (2001)
Quang, D., Chen, Y. & Xie, X. DANN: a deep learning approach for annotating the pathogenicity of genetic variants. Bioinformatics 31, 761–763 (2015)
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012)
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Morgan, M., Pagès, H., Obenchain, V. & Hayden, N. Rsamtools: Binary alignment (BAM), FASTA, variant call (BCF), and tabix file import. R package v.1.28.0 http://bioconductor.org/packages/release/bioc/html/Rsamtools.html (2017)
Dror, Y. & Freedman, M. H. Shwachman–Diamond syndrome. Br. J. Haematol. 118, 701–713 (2002)
Austin, K. M. et al. Mitotic spindle destabilization and genomic instability in Shwachman–Diamond syndrome. J. Clin. Invest. 118, 1511–1518 (2008)
Schmidt, A. et al. Severely altered guanidino compound levels, disturbed body weight homeostasis and impaired fertility in a mouse model of guanidinoacetate N-methyltransferase (GAMT) deficiency. Hum. Mol. Genet. 13, 905–921 (2004)
Acknowledgements
We thank members of the MacArthur laboratory and the Laboratory, Data Analysis, and Coordinating Center (LDACC) for performing the quality control of the whole genome sequencing data, D. Conrad for help with the structural variant calls, D. A. Knowles for code review, J. T. Leek and C. D. Brown for feedback on the manuscript, and the artists of the graphics that we modified in Fig. 1 (https://pixabay.com/en/man-silhouette-stand-straight-308387/ and http://www.allvectors.com/human-organs/). The Genotype-Tissue Expression (GTEx) project was supported by the Common Fund of the Office of the Director of the National Institutes of Health (NIH). Additional funds were provided by the National Cancer Institute; National Human Genome Research Institute (NHGRI); National Heart, Lung, and Blood Institute; National Institute on Drug Abuse; National Institute of Mental Health; and National Institute of Neurological Disorders and Stroke. Donors were enrolled at the Biospecimen Source Sites funded by Leidos Biomedical, Inc. (Leidos) subcontracts to the National Disease Research Interchange (10XS170) and Roswell Park Cancer Institute (10XS171). The LDACC was funded through a contract (HHSN268201000029C) to The Broad Institute. Biorepository operations were funded through a Leidos subcontract to the Van Andel Institute (10ST1035). Additional data repository and project management were provided by Leidos (HHSN261200800001E). The Brain Bank was supported by a supplement to University of Miami grant DA006227. We are grateful for support from a Hewlett-Packard Stanford Graduate Fellowship (E.K.T.), a doctoral scholarship from the Natural Science and Engineering Council of Canada (E.K.T.), a Lucille P. Markey Biomedical Research Stanford Graduate Fellowship (J.R.D.), the Stanford Genome Training Program (SGTP; NHGRI T32HG000044) (J.R.D., Z.Z.), the National Science Foundation GRFP (DGE-114747) (Z.Z.), the Joseph C. Pistritto Research Fellowship (F.N.D.), NIH training grant T32 GM007057 (B.J.S.), a Mr and Mrs Spencer T. Olin Fellowship for Women in Graduate Study (A.J.S.), the Searle Scholars Program (A.B.), NIH grants 1R01MH109905-01 (A.B.), R01MH101814 (NIH Common Fund; GTEx Program) (A.B. and S.B.M.), R01HG008150 (NHGRI; Non-Coding Variants Program) (A.B., S.B.M.), and NHGRI grants U01HG007436 and U01HG009080 (S.B.M.).
Author information
Authors and Affiliations
Consortia
Contributions
X.L., Y.K., E.K.T., J.R.D., A.B. and S.B.M. designed the study, performed analyses and wrote the manuscript. Y.K., F.N.D. and A.B. developed RIVER. G.T.H., A.L. and M.C.B. designed and executed the validation using CRISPR–Cas9. C.C., A.J.S. and I.M.H. provided the set of structural variants. J.D.M. provided the lists of curated cancer and cardiovascular disease genes. Z.Z., B.J.S. and A.G. contributed to analysis and feedback.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks E. Birney, A. Clark and Y. Gilad for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 PEER correction.
a, Adjusted R2 between top 15 PEER factors and top 20 sample (left) and subject (right) covariates in an example tissue, skeletal muscle. Covariates were ranked by the average adjusted R2 across all PEER factors and hierarchically clustered. The corresponding data for all tissues are provided in Supplementary Tables 1, 2. b, Adjusted R2 between the total expression component removed by PEER in each tissue and the top 20 sample (left) and subject (right) covariates. The covariates were ranked by the average adjusted R2 across all tissues, and both axes were hierarchically clustered. White denotes missing values, and tissues are coloured as in Fig. 1. PEER factors captured slightly different covariates across tissues, with a noticeable difference between the brain and other tissues. c, Rare variant enrichments as in Fig. 2a for different levels of PEER correction. The fully corrected data show substantially stronger rare variant enrichments than the two partially corrected datasets.
Extended Data Figure 2 Distribution of the number of genes with a multi-tissue outlier.
a, Distribution of the number of genes for which each individual was a multi-tissue outlier. Each individual was an outlier for a median of 10 genes. Individuals with 50 or more outliers are coloured in grey and were excluded from downstream analyses. b–f, Distribution of the number of genes for which individuals, stratified by common covariates, were multi-tissue outliers. For race and sex, we compared the distributions using an unsigned Wilcoxon rank-sum test, whereas we used Spearman’s ρ to test for association with the remaining covariates. Only age (Spearman’s ρ = 0.10, P = 0.033) and ischaemic time (Spearman’s ρ = 0.18, P = 0.00022) were nominally associated with the number of outlier genes per individual. The association with age fails to achieve significance after correcting for multiple testing using the Bonferroni method. Note that in b we only tested for a significant difference in the distribution of the number of outlier genes between white and black individuals, because there were too few individuals in the other groups. g, Enrichments as shown in Fig. 2a either including all individuals, or excluding individuals that are outliers for 50 (matches Fig. 2a) or 30 genes.
Extended Data Figure 3 Single-tissue outlier replication.
a, Correlation between the replication proportions (see Methods) obtained from all samples and from a subset of 70 overlapping individuals per tissue pair (Pearson’s correlation, P < 2.2 × 10−16). When restricting to 70 individuals, the replication rates decreased more for discovery tissues with larger sample sizes in the full dataset, indicating that replication rates were underestimated for tissues with small sample sizes. b, Correlation between replication in the 70 individuals used for discovery and replication assessed in a set of 70 individuals that included the outlier individual and 69 individuals excluded from the discovery set (Pearson’s correlation, P < 2.2 × 10−16). Replication was higher when computed in the discovery individuals rather than in a distinct set of individuals. c, Single-tissue outlier replication using all individuals, as in Fig. 1b, but data are only shown for pairs with at least 70 overlapping individuals. Tissue pairs with insufficient overlap are in grey. d, For each pair of tissues with sufficient samples, outlier discovery and replication using 70 individuals sampled in both tissues. The replication values decreased compared with replication performed in all individuals (c), particularly for tissues with large sample sizes in the complete dataset. However, the pattern of replication, with more similar tissues having higher replication rates, is maintained. e, For each tissue, the proportion of (individual, gene) outlier pairs where the individual was also a multi-tissue outlier for the gene. This proportion was positively correlated with the tissue sample size (P = 1.4 × 10−10). Points are coloured by tissue as in Fig. 1.
Extended Data Figure 4 Number of rare variants per individual and population structure.
a, The distribution of the number of rare variants of each type for individuals of European descent (reported as white). Certain individuals had many more rare variants than the population median (vertical black line). b, Principal component analysis of all individuals. Individuals are plotted according to their first two genotype principal components (PCs) and coloured by their reported ancestry. White individuals with WGS data, included in a, are coloured in a lighter shade of blue and those with 60,000 or more rare variants are circled in black. The individuals with an excess of rare variants probably had African or Asian admixture. c, Enrichments as in Fig. 2a and excluding individuals with >60,000 rare variants (circled in b), which did not substantially affect the enrichment patterns. d, European population allele frequency distributions in the 1000 Genomes Project of rare SNVs and indels used in our analysis. The rare variants included in our analysis were constrained to have MAF ≤ 0.01 in the 1000 Genomes European super population, but they were also relatively rare in each of the individual European populations.
Extended Data Figure 5 Comparison of overexpression and underexpression outliers.
a, ASE at rare exonic variants. ASE is shown as the ratio of the number of reads supporting the minor allele to the total number of reads at the site. If the rare variant is driving the extreme expression, we expect this ratio to be below 0.5 for underexpression outliers and above 0.5 for overexpression outliers. Rare coding variants were enriched for ASE in the direction of the extreme expression effect (two-sided Wilcoxon rank-sum tests, each nominal P < 4.0 × 10−8). b, Expression level distribution of all genes and genes with overexpression or underexpression outliers. Expression is shown as the log2 of the median (RPKM + 2), where the median was first taken across individuals in each tissue then across expressed tissues for each gene. For genes with low expression, even an RPKM of 0 may not yield a Z-score ≤ −2. Indeed, underexpression outliers were depleted among low expressed genes whereas the opposite was true of overexpression outliers (two-sided Wilcoxon rank-sum test comparing to all genes, P < 2.2 × 10−16 for both overexpression and underexpression). c, Feature enrichments (as in Fig. 3b) shown separately for over and underexpression outliers.
Extended Data Figure 6 Extended rare variant enrichments.
a, For each tissue, rare SNV enrichment in single-tissue outliers compared with non-outliers at the same genes for increasing Z-score thresholds. Enrichments calculated as in Fig. 2. The rare variant enrichments varied between tissues though the overall pattern mirrored that of multi-tissue outliers when combining all the tissues (Fig. 2b). The high variance in the enrichments underscores the noise in single-tissue outlier discovery. b, As in Fig. 2a, enrichment for SNVs, indels and structural variants in outliers compared with the same genes in non-outliers, either including all rare variants or only those outside protein-coding or lincRNA exons in Gencode v.19. The enrichment of rare variants was weaker, but still significant, for all variant types when excluding exonic regions.
Extended Data Figure 7 Enrichment of an extended list of functional genomic annotations.
log odds ratios and 95% Wald confidence intervals from logistic regression models of outlier status as a function of each genomic feature. Features were calculated among rare SNVs within 10 kb of the gene. When more than one feature corresponded to the same genomic annotation (for example, the number or the presence of rare variants in a splice region; Supplementary Table 3b), the feature with the highest enrichment is shown. Lighter shading indicates a non-significant log odds ratio (nominal P > 0.05).
Extended Data Figure 8 Evolutionary constraint and regulatory control of multi-tissue outlier genes.
a, Odds ratio of being intolerant to synonymous and missense variants for genes with multi-tissue eQTLs (eGenes), genes with multi-tissue outliers, OMIM and GWAS genes (see Methods). As expected, GWAS and OMIM genes showed no enrichment or depletion for synonymous variation intolerant genes. Genes with multi-tissue outliers and eGenes showed slight depletion for these genes. Genes with multi-tissue outliers and eGenes were strongly depleted for genes intolerant to missense variation compared with OMIM and GWAS genes. b, Comparison of the depletion of disease genes among genes with a multi-tissue outlier and eGenes. Similar to Fig. 4c, bars represent 95% confidence intervals from Fisher’s exact test. c, For each of ten gene lists, the difference in the mean number of variants near genes in the list compared with the mean for all other annotated genes. Results are stratified by minor allele frequency, and bars indicate the 95% confidence interval for the difference from a two-sided t-test. Disease genes had more variants than control genes in general, and the difference was particularly striking for rare variants. This suggests that the depletion of outliers and eQTLs for certain groups of disease genes is not due to less rare variation near these genes. Instead, we hypothesize that the variation around these genes in our healthy cohort is less likely to have large regulatory effects. d, Distribution of the number of tissues with an eQTL for genes with and without outliers. Genes with multi-tissue outliers had eQTLs in more tissues than genes without. This suggests that they are more susceptible to shared regulatory control. This result held for both multi-tissue eQTL definitions (see Methods; Meta-Tissue: 23 versus 3 tissues, Wilcoxon rank-sum test P < 2.2 × 10−16; tissue-by-tissue: 7 versus 3 tissues, P < 2.2 × 10−16). e, This eGene enrichment was robust across different mean expression levels across tissues (two-sided Wilcoxon rank-sum tests, Bonferroni-adjusted P < 1 × 10−11).
Extended Data Figure 9 RIVER performance.
a, Comparison between the predictive power of RIVER and that of the genomic annotation model, as in Fig. 5a, across different Z-score thresholds for outlier calling. Increasing the Z-score threshold improved AUC values, but reduced the number of outlier examples, which led to noisy receiver operating characteristic curves. b, Stability analysis of estimated parameters with different parameter initializations (see Methods). c, Correlations, using Kendall’s τ, between the fraction of tissues with |Z-score| ≥ 2 and the test probabilities from the genomic annotation model (left) and RIVER (right). We calculated test posterior probabilities using tenfold cross-validation and only considered individual and gene pairs with a fraction of tissues with |Z-score| ≥ 2 that was significantly different from 0.05 (one-sided binomial exact test, Benjamini–Hochberg adjusted P < 0.05). d, P values from a one-sided Fisher’s exact test measuring the association between allelic imbalance (see Methods) and the posterior probability of a functional rare variant according to the genomic annotation model and RIVER. The posterior probabilities from RIVER were more strongly associated with allelic imbalance across all four thresholds tested. e, Assessment of the advantage of incorporating gene expression with genomic annotations for predicting outlier status using simplified supervised models (see Methods). All models showed consistent improvement of the log odds ratio of outlier status when incorporating expression. f, Performance of models with 12 individual genomic features compared with the genomic annotation model and RIVER. Some models with single genomic features provided slightly better AUCs compared with the genomic annotation model, but they were not statistically different. On the other hand, RIVER predicted the effects of rare variants significantly better than each of the models that included a single feature.
Extended Data Figure 10 Evaluation of known pathogenic variants using RIVER.
a, The 27 GTEx rare SNVs reported as disease variants in ClinVar. b–d, Relative frequency of the |median Z-score| (b), posterior probabilities from the genomic annotation model (c) and posterior probabilities from RIVER (d) for all (individual, gene) pairs (grey) and 27 pairs with pathogenic variants from ClinVar (orange). P values were computed using two-sided Wilcoxon rank-sum tests. We note that rare indels and structural variants were not found nearby the genes in the individuals carrying these pathogenic variants. e, f, The Z-score and RPKM distributions for SBDS (e) and GAMT (f) were compared with the values from four individuals carrying regulatory pathogenic variation (red asterisks and triangles). The median Z-score and RPKM values across tissues are shown at the top of each plot (black circle). Tissues are coloured as in Fig. 1 and sorted in decreasing order of the difference between the average Z-score of individual(s) with a regulatory pathogenic variant and the median Z-score for the tissue. Three individuals carrying a total of two unique rare variants are shown for SBDS. Both variants are associated with the recessive Shwachman–Diamond syndrome, which causes systemic symptoms that include pancreatic, neurological and haematologic abnormalities46 and can disrupt fibroblast function47. The individuals, being heterozygous for these variants, lacked the disease phenotype. Nonetheless, we saw extreme underexpression of SBDS across almost all tissues in these individuals, including brain tissues, fibroblasts and pancreas. One individual had a rare variant for GAMT associated with cerebral creatine deficiency syndrome 2, shown to cause neurological deficiencies and also lead to low body fat48. The individual had the most extreme underexpression in (subcutaneous) adipose tissue.
Extended Data Figure 11 Validation of large-effect rare variants using CRISPR–Cas9 genome editing.
a, SNVs in outliers and controls assayed for expression effects using CRISPR–Cas9 genome editing. For common SNVs in controls (MAF >1% in the GTEx cohort), the range of median Z-scores and RIVER scores are given for all individuals with the minor allele. Missing values indicate that the variant was absent from our cohort. b, sgRNAs for four SNVs found in outliers and four control SNVs in the same genes. c, Alternate (installed) gDNA and cDNA allele proportions for four rare, coding SNVs in outliers (left) and four matched control SNVs (right). Each gDNA and cDNA sample was sequenced in triplicate (technical replicates). Asterisks denote the Bonferroni-adjusted significance level from a two-sided t-test of the difference between the gDNA and cDNA alternate allele proportions: ·P < 0.05, *P < 0.01, **P < 0.001. Although one control SNV showed a significant difference in the alternate allele proportion between cDNA and gDNA, it displayed an increase rather than a decrease in expression.
Supplementary information
Supplementary Table 1
This table contains the adjusted R2 values between the top PEER factors estimated for each tissue and the sample covariates available for that tissue. Data for each tissue are presented in separate sheets named according to the tissue ID. Example adjusted R2 data for sample covariates for skeletal muscle (tissue ID: Muscle_Skeletal) are presented in Extended Data Figure 1a (left). (XLS 554 kb)
Supplementary Table 2
This table contains the adjusted R2 values between the top PEER factors estimated for each tissue and the subject covariates available for that tissue. Data for each tissue are presented in separate sheets named according to the tissue ID. Example adjusted R2 data for subject covariates for skeletal muscle (tissue ID: Muscle_Skeletal) are presented in Extended Data Figure 1a (right). (XLS 1458 kb)
Supplementary Table 3
This table contains descriptions of the variant features for the enrichment analyses and training of RIVER. Sheet ‘A-Selected features’ describes the disjoint variant classes whose enrichments among outliers are presented in Figure 3 and Extended Data Figure 5c. Sheet ‘B-RIVER features’ describes the gene-level features used for training RIVER. The enrichments of these features among outliers are presented in Extended Data Figure 7. (XLSX 31 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International (CC BY 4.0) licence. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, X., Kim, Y., Tsang, E. et al. The impact of rare variation on gene expression across tissues. Nature 550, 239–243 (2017). https://doi.org/10.1038/nature24267
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24267
This article is cited by
-
Analysis of 3760 hematologic malignancies reveals rare transcriptomic aberrations of driver genes
Genome Medicine (2024)
-
Massively parallel screen uncovers many rare 3′ UTR variants regulating mRNA abundance of cancer driver genes
Nature Communications (2024)
-
High-dimensional phenotyping to define the genetic basis of cellular morphology
Nature Communications (2024)
-
Investigating the role of common cis-regulatory variants in modifying penetrance of putatively damaging, inherited variants in severe neurodevelopmental disorders
Scientific Reports (2024)
-
Haplotype-aware modeling of cis-regulatory effects highlights the gaps remaining in eQTL data
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.