Abstract
X chromosome inactivation (XCI) silences transcription from one of the two X chromosomes in female mammalian cells to balance expression dosage between XX females and XY males. XCI is, however, incomplete in humans: up to one-third of X-chromosomal genes are expressed from both the active and inactive X chromosomes (Xa and Xi, respectively) in female cells, with the degree of ‘escape’ from inactivation varying between genes and individuals1,2. The extent to which XCI is shared between cells and tissues remains poorly characterized3,4, as does the degree to which incomplete XCI manifests as detectable sex differences in gene expression5 and phenotypic traits6. Here we describe a systematic survey of XCI, integrating over 5,500 transcriptomes from 449 individuals spanning 29 tissues from GTEx (v6p release) and 940 single-cell transcriptomes, combined with genomic sequence data. We show that XCI at 683 X-chromosomal genes is generally uniform across human tissues, but identify examples of heterogeneity between tissues, individuals and cells. We show that incomplete XCI affects at least 23% of X-chromosomal genes, identify seven genes that escape XCI with support from multiple lines of evidence and demonstrate that escape from XCI results in sex biases in gene expression, establishing incomplete XCI as a mechanism that is likely to introduce phenotypic diversity6,7. Overall, this updated catalogue of XCI across human tissues helps to increase our understanding of the extent and impact of the incompleteness in the maintenance of XCI.
Similar content being viewed by others
Main
Mammalian female tissues consist of two mixed cell populations, each with either the maternally or paternally inherited X chromosome marked for inactivation. To overcome this heterogeneity, assessments of human XCI have often been confined to the use of artificial cell systems1 or to samples that have skewed XCI1,2, that is, preferential inactivation of one of the two X chromosomes; this is common in clonal cell lines but rare in karyotypically normal, primary human tissues8 (Extended Data Fig. 1 and Supplementary Note). Others have used bias in DNA methylation3,4,9 or in gene expression5,10 between males and females as a proxy for XCI status. Surveys of XCI are powerful in engineered model organisms, for example, mouse models with completely skewed XCI11, but the degree to which these discoveries are generalizable to human XCI remains unclear given marked differences in XCI initiation and the extent of escape across species7. Here we describe a systematic survey of the landscape of human XCI using three complementary RNA sequencing (RNA-seq)-based approaches (Fig. 1) that together enable the assessment of XCI from individual cells to population level across a diverse range of human tissues.
Previous expression-based surveys of XCI1,2 have established the incomplete and variable nature of XCI, but these studies have been limited in the tissue types and samples assessed. To investigate the landscape of XCI across human tissues, we combined three approaches: (1) sex biases in expression using population-level GTEx data across 29 tissue types; (2) allelic expression in 16 tissue samples from a female GTEx donor with fully skewed XCI, and (3) validation using scRNA-seq by combining allelic expression and genotype phasing. WGS, whole-genome sequencing; WES, whole-exome sequencing.
Given the limited accessibility of most human tissues, particularly in large sample sizes, no global investigation into the impact of incomplete XCI on X-chromosomal expression has been conducted in datasets spanning multiple tissue types. We used the Genotype-Tissue Expression (GTEx) project12,13 dataset (v6p release), which includes high-coverage RNA-seq data from diverse human tissues, to investigate male–female differences in the expression of 681 X-chromosomal genes that encode proteins or long non-coding RNA in 29 adult tissues (Extended Data Table 1), hypothesizing that escape from XCI should typically result in higher female expression of these genes. Previous work5,10,14 has indicated that some of the genes that escape XCI (hereafter referred to as escape genes) show female bias in expression, but our analysis benefits from a larger set of profiled tissues and individuals, as well as the high sensitivity of RNA-seq.
To confirm that male–female expression differences reflect incomplete XCI, we assessed the enrichment of sex-biased expression in known XCI categories using 561 genes with previously assigned XCI status, defined as escape (n = 82), variable escape (n = 89) or inactive (n = 390) (Fig. 1 and Supplementary Table 1). Sex-biased expression is enriched in escape genes compared to both inactive genes (two-sided paired Wilcoxon rank-sum test, P = 3.73 × 10−9) and variable escape genes (P = 3.73 × 10−9) (Fig. 2b and Extended Data Fig. 2), with 74% of escape genes showing significant (false-discovery rate (FDR) q < 0.01) male–female differences in at least one tissue (Fig. 2a, Extended Data Figs 3, 4 and Supplementary Table 2). In line with two active X-chromosomal copies in females, escape genes in the non-pseudoautosomal, that is, the X-specific, region (nonPAR) predominantly show female-biased expression across tissues (52 out of 67 assessed genes, binomial P = 6.46 × 10−6). However, genes in the pseudoautosomal region PAR1, are expressed more highly in males (14 out of 15 genes, binomial P = 9.77 × 10−6) (Fig. 2a), suggesting that combined Xa and Xi expression in females fails to reach the expression arising from X and Y chromosomes in males (discussed below).
a, Male–female expression differences in reported XCI-escaping genes (n = 82) across 29 GTEx tissues. Definitions for the abbreviations can be found in Extended Data Table 1. b, Proportion of significantly biased (FDR <1%) genes in each tissue by reported XCI status. c, Proportion of tissues where the bias direction is shared with the reported XCI status. Genes expressed in at least five tissues are included. d, Sex bias pattern of nine genes not classified as full escape genes that follow a similar profile to established escape genes. e, Chromatin state enrichment between escape and inactive genes in the Roadmap Epigenomics31 female samples.
Sex bias of escape genes is often shared across tissues; these genes show a higher number of tissues with sex-biased expression than genes in other XCI categories (Fig. 2a and Extended Data Fig. 2c), a result that is not driven by differences in the breadth of expression of escape and inactive genes (Extended Data Fig. 2e). Also, the direction of sex bias across tissues is consistent (Fig. 2a, c and Extended Data Fig. 2b). Together, these observations indicate that there is global and tight control of XCI, that potentially arises from early lockdown of the epigenetic marks regulating XCI. Previous reports have identified several epigenetic signatures associated with XCI escape in humans and mice15; in agreement with these discoveries we show that escape genes are enriched in chromatin states that are related to active transcription (Fig. 2e).
Although sex bias on the X chromosome is broadly specific to escape genes, some genes show unexpected patterns. Eight genes with some previous evidence for inactivation show >90% concordance in effect direction and significant sex bias (Fig. 2d and Supplementary Table 3), suggesting that variable escape can also have considerable population-level effects. For example, CHM demonstrates such concordance in sex bias and escape at this gene is confirmed when using single-cell RNA-seq (scRNA-seq; see below). One gene (RP11-706O15.3) without an assigned XCI status shows a similar sex bias pattern to escape genes. RP11-706O15.3 resides between escape and variable escape genes PRKX and NLGN4X (Fig. 2d), consistent with known clustering of escape genes1,2. Some escape genes show more heterogeneous sex bias, for example, ACE2 (Fig. 2a and Supplementary Discussion). Many such genes lie in the evolutionarily older region of the chromosome16, in Xq, where escape genes also show higher tissue-specificity and lower expression levels (Extended Data Fig. 5), characteristics that have been linked with higher protein evolutionary rates17,18.
Although sex bias serves as a proxy for XCI status, it provides only an indirect measurement of XCI. We identified a GTEx female donor with an unusual degree of skewing of XCI (Fig. 3a), in whom the same copy of chromosome X was silenced in approximately 100% of cells across all tissues, but with no X-chromosomal abnormality detected by whole-genome sequencing (Extended Data Fig. 6 and Supplementary Note), providing an opportunity to analyse allele-specific expression (ASE) across 16 tissues to investigate XCI. This approach is analogous to previous surveys in mouse11 or in human cell lines with skewed XCI2, but extends the assessment to a larger number of tissues and avoids biases arising from genetic heterogeneity between tissue samples.
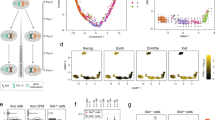
a, Distribution of the skewness of XCI in GTEx female samples (n = 62, v3 release). Each data point shows the mean skew in XCI across tissue samples per individual. b, Classification of X-chromosomal genes (n = 186) into full or incomplete and tissue-shared or heterogeneous XCI based on the analysis of ASE patterns across tissues. Error bars show the 95% credible interval. c–e, Examples of genes where the ASE-based assessment of XCI status match previously reported assignments (TSR2, inactive; XIST, escape; ZBED1, escape). Note that XIST is only expressed monoallelically from Xi, which is unusual for an escape gene. f, KAL1 shows strong evidence for tissue-specific escape. g–k, Genes without previous or conclusive evidence for escape from XCI that are classified as incompletely inactivated in this sample. In c–k asterisks indicate that the Xi expression in the given tissue was significant at FDR <1% (one-sided binomial test) and error bars show the 95% confidence interval.
Analysis of the X-chromosomal allelic counts (Supplementary Tables 4–6) from this GTEx donor highlights the incompleteness and consistency of XCI across tissues (Fig. 3b). Approximately 23% of the 186 X-chromosomal genes that were assessed show expression from both alleles, indicative of incomplete XCI, matching previous estimates of the extent of escape1,2. For 43% of the genes that were expressed from both alleles in this sample, Xi expression is of a similar magnitude between tissues, therefore supporting the observation of a general global and tight control of XCI. However, suggesting some tissue dependence in XCI, the rest of the genes that were expressed from both alleles show variability in Xi expression, including a subset of genes (5.8% of all genes) that appear biallelic in only one of the multiple tissues assayed. While tissue-specific escape is common in mouse11, limited evidence exists for such a pattern in human tissues other than for neurons3,4,9. In our data, one of the genes with the strongest evidence for tissue-specific escape is KAL1 (Fig. 3f and Supplementary Table 6), the causal gene for X-linked Kallmann syndrome. We show that KAL1 shows biallelic expression exclusively in the lung (Fig. 3f), in line with the strong female bias detected specifically in lung expression in the analysis described above and in Fig. 2a, suggesting that tissue differences in escape can directly translate into tissue-specific sex biases in gene expression. The predictions of XCI status in this sample not only align with previous assignments (Fig. 3c–f and Supplementary Table 7, for example, TSR2, XIST and ZBED1) but also suggest five new incompletely inactivated genes (Fig. 3g–k and Supplementary Table 5), three of which act in a tissue-specific manner. For instance, CLIC2, which in previous studies was shown to either be subject to2 or variably escape from1 XCI, shows considerable Xi expression only in skin tissue. Such specific patterns illustrate the need to assay multiple tissue types to fully uncover the diversity in XCI.
The emergence of scRNA-seq methods19 presents an opportunity to directly assess XCI without the complication of cellular heterogeneity in bulk tissue samples (Fig. 1), as demonstrated recently in mouse studies20,21,22,23 and in human fibroblasts24 and preimplantation development25. To directly profile XCI in human samples, we examined scRNA-seq data in combination with deep genotype sequences from 940 immune-related cells from four females: 198 cells from lymphoblastoid cell lines (LCLs) sampled from three females of African (Yoruba) ancestry, and 742 blood dendritic cells from a female of Asian ancestry26 (Fig. 1 and Extended Data Table 2). We used ASE to distinguish the expression coming from each of the two X-chromosomal haplotypes in a given cell (Supplementary Table 4). Because the inference of allele-specific phenomena in single cells is complicated by widespread monoallelic expression21,27,28,29, besides searching for X-chromosomal sites with biallelic expression (Extended Data Fig. 7), we leveraged genotype phase information to detect sites for which the expressed allele was discordant with the active X chromosome in that cell.
Only 129 (78%) out of the 165 assayed genes (41–98 per sample) were fully inactivated in these data whereas the rest showed incomplete XCI in one or more samples (Fig. 4a, b and Supplementary Tables 8, 9); this is mostly consistent with previous assignments of XCI status to these genes (Fig. 4a and Supplementary Table 10). For instance, single-cell data reveal consistent expression from both X-chromosomal alleles for eleven genes in PAR1, in line with their known escape from XCI (for example, ZBED1, Fig. 4c), and replicate the known expression of XIST exclusively from Xi (Fig. 4d).
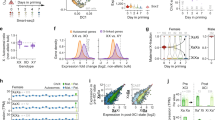
a, Proportion of genes demonstrating full and partial XCI in the ASE analysis in scRNA-seq data, and the concordance with previously reported XCI status. b–l, Examples of genes with different XCI patterns in scRNA-seq: previously reported inactive gene (b), known escape gene in PAR1 (c), escape gene with known exclusive expression from Xi (d), new candidates for escape genes that demonstrate incomplete XCI in only a subset of samples (e–k), and a known escape gene that shows escape of varying degrees in the three samples (Pearson’s χ2 test for equal proportions, P = 3.80 × 10−7) (l). b–l, x axis labels are sample identifiers. Asterisk above a bar indicates that the proportion of Xi expression, that is, blue bar, in a given sample is significantly greater than the expected baseline (FDR <1%, one-sided binomial test). Error bars show the 95% confidence interval.
We next assessed whether our approach could extend the spectrum of escape from XCI. For seven genes that have previously been reported as inactivated, the data from single cells pointed to incomplete XCI (Fig. 4e–k and Supplementary Table 11), including FHL1, which was also highlighted as a candidate escape gene in the GTEx ASE analysis (Fig. 4e), and ATP6AP2, which displays predominantly female-biased expression across GTEx tissues (Fig. 4h). Both of these genes demonstrate significant Xi expression in only a subset of the scRNA-seq samples, a pattern that is consistent with variable escape1,2. Between-individual variability exists not only in the presence but also in the degree of expression from Xi (for example, MSL3, Fig. 4l). Highlighting the capacity of scRNA-seq to provide information beyond bulk RNA-seq, we identify examples where Xi expression varies considerably between the two X-chromosomal haplotypes within an individual (for example, ASMTL; Supplementary Table 12), suggesting cis-acting variation as one of the determinants for the level of Xi expression3. As a further layer of heterogeneity in Xi expression, we find a unique pattern for TIMP1. For this gene, the level of Xi expression across cells is not significant, but exclusive to a subset of cells that express the gene biallelically (Extended Data Fig. 7), pointing to cell-to-cell variability in escape.
Using the ASE estimates from the scRNA-seq and GTEx analyses to infer the magnitude of the incompleteness of XCI, we find that expression from Xi at escape genes rarely reaches levels equal to expression from Xa, Xi expression remaining on average at 33% of Xa expression. However, there is a lot of variability along the chromosome (Extended Data Fig. 8a and Supplementary Discussion), as has previously been demonstrated in specific tissue types1,2. Balanced expression dosage between males and females in PAR1 requires full escape from XCI, however, Xi expression remains below Xa expression also in this region (mean Xi to Xa ratio is around 0.80), pointing to partial spreading of XCI beyond nonPAR. In further support that the consistent male bias in PAR1 expression (Fig. 2a) is due to the incompleteness of escape, we observe no systematic up- or downregulation of Y chromosome expression in PAR1 (Extended Data Fig. 8b and Supplementary Discussion). As another consequence of the partial Xi expression, several of the X–Y homologous genes in nonPAR30 become male-biased when expression from the Y chromosome counterpart is accounted for (Extended Data Fig. 8c).
By combining diverse types and analyses of high-throughput RNA-seq data, we have systematically assessed the incompleteness and heterogeneity in XCI across 29 human tissues (Supplementary Table 13). We establish that scRNA-seq is suitable for surveys of human XCI and present the first steps towards understanding the cellular-level variability in the maintenance of XCI. Our phasing-based approach enables the full use of low-coverage scRNA-seq, however, because any single individual and cell type is only informative for restricted number of genes, larger datasets with more diverse cell types and conditions are required to fully profile XCI. We have therefore used the multi-tissue GTEx dataset to explore XCI in a larger number of X-chromosomal genes and to assess the tissue heterogeneity and impacts of XCI on gene expression differences between the sexes.
These analyses show that incomplete XCI is mostly shared between individuals and tissues, and extend previous surveys by pinpointing several examples of variability in the degree of XCI escape between cells, chromosomes, and tissues. In addition, our data demonstrate that escape from XCI results in sex-biased expression of at least 60 genes, potentially contributing to sex-specific differences in health and disease (Supplementary Discussion). As a whole, these results highlight the between-female and male–female diversity introduced by incomplete XCI, the biological implications of which remain to be fully explored.
Methods
GTEx data
The GTEx project12 collected tissue samples from 554 postmortem donors (187 females, 357 males; age range, 20–70), carried out RNA-seq on 8,555 tissue samples and generated genotyping data for up to 449 donors (GTEx analysis v6p release). More detailed methods can be found in ref. 13. All GTEx data, including RNA, genome and exome sequencing data, used in the analyses described here are available through dbGaP under accession number phs000424.v6.p1, unless otherwise stated. Summary data and details on data production and processing are also available from the GTEx Portal (http://gtexportal.org).
Single-cell samples
For the human dendritic cells samples profiled, the healthy donor (ID: 24A) was recruited from the Boston-based PhenoGenetic project, a resource of healthy subjects that are re-contactable by genotype32. The donor was a female Asian individual from China, 25 years of age at the time of blood collection. She was a non-smoker, had a normal BMI (height: 168.7 cm; weight: 56.45 kg; BMI: 19.8), and normal blood pressure (108/74). The donor had no family history of cancer, allergies, inflammatory disease, autoimmune disease, chronic metabolic disorders or infectious disorders. She provided written informed consent for the genetic research studies and molecular testing, as previously reported26.
Daughters of three parent–child Yoruba trios from Ibadan, Nigeria (that is, YRI trios), collected as part of the International HapMap Project, were chosen for single-cell profiling, both to maximize heterozygosity and due to availability of parental genotypes enabling phasing. DNA and LCLs were ordered from the NHGRI Sample Repository for Human Genetic Research (Coriell Institute for Medical Research): LCLs from B lymphocytes for the three daughters (catalogue numbers: GM19240, GM19199 and GM18518) and DNA extracted from LCLs for all members of the three trios (catalogue numbers for DNA: NA19240, NA19238, NA19239, NA19199, NA19197, NA19198, NA18518, NA18519 and NA18520). These YRI samples are referred to by their family IDs: Y014, Y035 and Y117.
Clinical muscle samples
To assess whether PAR1 genes are equally expressed from X and Y chromosomes, a combination of skeletal muscle RNA-seq data and trio genotyping data from eight male patients with muscular dystrophy, sequenced as part of an unrelated study, was used. Patient cases with available muscle biopsies were referred from clinicians starting April 2013 until June 2016. All patients included for RNA-seq had previously available trio whole-exome sequencing (WES) data, with one sample having additional trio whole-genome sequencing (WGS). Muscle biopsies were shipped frozen from clinical centres by liquid nitrogen dry shipping and, where possible, frozen muscle was sectioned on a cryostat and stained with haematoxylin and eosin to assess muscle quality as well as the presence of overt freeze–thaw artefacts.
Genotyping
The GTEx v6p release includes WGS data for 148 donors, including GTEX-UPIC. WGS libraries were sequenced on the Illumina HiSeqX or Illumina HiSeq2000. WGS data was processed through a Picard-based pipeline, using base quality score recalibration and local realignment at known indels. BWA-MEM aligner was used for mapping reads to the human genome build 37 (hg19). Single-nucleotide polymorphisms (SNPs) and indels (insertions and deletions) were jointly called across all 148 samples and additional reference genomes using HaplotypeCaller v.3.1 of GATK. Default filters were applied to SNP and indel calls using the variant quality score recalibration (VQSR) approach of GATK. An additional hard filter InbreedingCoeff ≤−0.3 was applied to remove sites that VQSR failed to filter.
WGS for one of the clinical muscle samples was performed on 500 ng to 1.5 μg of genomic DNA using a PCR-Free protocol that substantially increases the uniformity of genome coverage. These libraries were sequenced on the Illumina HiSeq X10 with 151-bp paired-end reads and a target mean coverage of >30×, and were processed similarly to the above description.
The Y117 trio (sample IDs NA19240 (daughter), NA19238 (mother), and NA19239 (father)) was whole-genome-sequenced as part of the 1000 Genomes Project as described previously33. The VCF file containing the WGS-based genotypes for SNPs (YRI.trio.2010_09.genotypes.vcf.gz) was downloaded from the FTP site of the project. The genotype coordinates (in human genome build 36) in the original VCF were converted to hg19 using the liftover script (liftOverVCF.pl) and chain files provided as part of the GATK package.
WES was performed using Illumina’s capture Exome (ICE) technology (Y035, Y014, 24A) or Agilent SureSelect Human All Exon Kit v.2 exome capture (clinical muscle samples) with a mean target coverage of >80×. WES data was aligned with BWA, processed with Picard, and SNPs and indels were jointly called with other samples using GATK HaplotypeCaller package v.3.1 (24A, clinical muscle samples) or v.3.4 (Y035, Y014). Default filters were applied to SNP and indel calls using the VQSR approach. A modified version of the Ensembl variant effect predictor was used for variant annotation for all WES and WGS data. For trio WES or WGS data the genotypes of the proband were phased using the PhaseByTransmission tool of the GATK toolkit.
Single-cell data preparation and sequencing
For profiling of healthy dendritic cells (DCs), peripheral blood mononuclear cells (PBMCs) were first isolated from fresh blood within 2 h of collection, using Ficoll–Paque density gradient centrifugation as previously described34. Single-cell suspensions were stained as per the manufacturer’s recommendations with an antibody panel designed to enrich for all known blood DC population for single-cell sorting and scRNA-seq profiling26. A total of 24 single cells from four loosely gated populations were sorted per 96-well plate, with each well containing 10 μl of lysis buffer. A total of eight plates were analysed by scRNA-seq.
All LCL cell lines were cultured according to Coriell’s recommendations (medium: RPMI 1640, 2 mM l-glutamine, 15% fetal bovine serum (all three from ThermoFisher Scientific)) in T25 tissue culture flask with 10–20 ml medium at 37 °C in 5% carbon dioxide. Cells were split upon reaching a cell density of approximately 300,000–400,000 viable cells per ml. All three lymphoblast cultures were split once before single-cell sorting. Cells were washed with 1× PBS, the pellet was resuspended and stained with DAPI (Biolegend) for viability according to the manufacturer’s recommendations.
All single live cells (for both DCs and LCL cell lines) were sorted into a 96-well full-skirted Eppendorf plate chilled to 4 °C, that were pre-prepared with 10 μl TCL buffer (Qiagen) supplemented with 1% β-mercaptoethanol (lysis buffer), using a BD FACS Fusion instrument. Single-cell lysates were sealed, vortexed, spun down at 300g at 4 °C for 1 min, immediately placed on dry ice and transferred for storage at −80 °C.
The Smart-Seq2 protocol was performed on single-sorted cells as described35,36, with some modifications as described in ref. 26 (Supplementary Methods). A total of 768 single DCs isolated from a healthy Asian female individual, along with 96 single cells from GM19240, 48 single cells from GM19199 and 48 single cells from GM18518 were profiled. In brief, single-cell lysates were thawed on ice, purified and reverse-transcribed using Maxima H Minus Reverse Transcriptase. PCR was performed with KAPA HiFi HotStart ReadyMix (KAPA Biosystems) and purified with Agencourt AMPureXP SPRI beads (Beckman-Coulter). The concentration of amplified cDNA was measured on the Synergy H1 Hybrid Microplate Reader (BioTek) using High-Sensitivity Qubit reagent (Life Technologies) and the size distribution of select wells was checked on a High-Sensitivity Bioanalyzer Chip (Agilent). The expected concentration was around 0.5−2 ng μl−1 with a size distribution that sharply peaked around 2 kb.
Library preparation was carried out using the Nextera XT DNA Sample Kit (Illumina) with custom indexing adapters, allowing up to 384 libraries to be simultaneously generated in a 384-well PCR plate (note that DCs were processed in a 384-well plate whereas LCLs were processed in 96-well plate format). The concentration of the final pooled libraries was measured using the High-Sensitivity DNA Qubit (Life Technologies) and the size distribution was measured on a High-Sensitivity Bioanalyzer Chip (Agilent). The expected concentration of the pooled libraries was 10–30 ng μl−1 with a size distribution of 300–700 bp. For the DCs, we created pools of 384 cells, whereas 96 LCL samples were pooled at the time. We sequenced one library pool per lane as paired-end 25-bp reads on a HiSeq2500 (Illumina). Barcodes used for indexing are listed in the Supplementary Methods.
RNA-seq in GTEx
RNA sequencing was performed using a non-strand-specific RNA-seq protocol with polyA selection of RNA using the Illumina TruSeq protocol with sequence coverage goal of 50 million 76-bp paired-end reads as has been previously described in detail12. The RNA-seq data, except for GTEX-UPIC, was aligned with TopHat v.1.4.1 to the UCSC human genome release version hg19 using the Gencode v.19 annotations as the transcriptome reference. Gene level read counts and reads per kilobase per million reads (RPKMs) were derived using the RNA-SeQC tool37 using the Gencode v.19 transcriptome annotation. The transcript model was collapsed into a gene model as described previously12. Read count and RPKM quantification include only uniquely mapped and properly paired reads contained within exon boundaries.
RNA-seq alignment to personalized genomes
For the four single-cell samples and for GTEX-UPIC RNA-seq, data were processed using a modification of the AlleleSeq pipeline38,39 to minimize reference allele bias in alignment. A diploid personal reference genome for each of the samples was generated with the vcf2diploid tool38 including all heterozygous biallelic single-nucleotide variants identified in WES or WGS either together with (YRI samples) or without (GTEX-UPIC, 24A) maternal and paternal genotype information. The RNA-seq reads were then aligned to both parental references using STAR40 v.2.4.1a in a per-sample two-pass mode (GTEX-UPIC and YRI samples) or v.2.3.0e (24A) using hg19 as the reference. The alignments were combined by comparing the quality of alignment between the two references: for reads aligning uniquely to both references the alignment with the higher alignment score was chosen and reads aligning uniquely to only one reference were kept as such.
RNA-seq of clinical muscle samples
Patient RNA samples derived from primary muscle were sequenced using the GTEx sequencing protocol12 with sequence coverage of 50 million or 100 million 76-bp paired-end reads. RNA-seq reads were aligned using STAR40 2-pass version v.2.4.2a using hg19 as the reference genome. Junctions were filtered after first pass alignment to exclude junctions with less than 5 uniquely mapped reads supporting the event and junctions found on the mitochondrial genome. The value for unique mapping quality was assigned to 60 and duplicate reads were marked with Picard MarkDuplicates (v.1.1099).
Catalogue of X-inactivation status
To compare results from the ASE and GTEx analyses with previous observations on genic XCI status we collated findings from two earlier studies1,2 that represent systematic expression-based surveys into XCI. Each study catalogues hundreds of X-linked genes and together the data span two tissue types.
Carrel and Willard1 surveyed in total 624 X-chromosomal transcripts expressed in primary fibroblasts in nine cell hybrids each containing a different human Xi. In order to find the gene corresponding to each transcript, the primer sequences designed to test the expression of the transcripts in the original study were aligned to reference databases based on the Gencode v.19 transcriptome and hg19 using in-house software (unpublished) (Supplementary Methods). In total 553 transcripts primer pairs were successfully matched to X-chromosomal Gencode v.19 reference mapping together with 470 unique X-chromosomal genes (Supplementary Methods). These 470 genes were split into three XCI status categories (escape, variable, inactive) based on the level of Xi expression (that is, the number of cell lines expressing the gene from Xi) resulting in 75 escape, 51 variable escape and 344 inactive genes.
Cotton et al.2 surveyed XCI using allelic imbalance in clonal or near-clonal female LCL and fibroblast cell lines and provided XCI statuses for 508 genes (68 escape, 146 variable escape, 294 subject genes). The data were mapped to Gencode v.19 using the reported gene names and their known aliases (Supplementary Methods), resulting in a list of XCI statuses for 506 X-chromosomal genes.
The results were combined by retaining the XCI status in the original study where possible (that is, same status in both studies or gene unique to one study) and for genes where the reported XCI statuses were in conflict the following rules were applied: (1) a gene was considered ‘escape’ if it was called escape in one study and variable in the other; (2) ‘variable escape’ if classified as escape and inactive; and (3) ‘inactive’ if classified as inactive in one study and variable escape in the other. The final combined list of XCI statuses consisted of 631 X-chromosomal genes including 99 escape, 101 variable escape and 431 inactive genes.
Analysis of sex-biased expression
Differential expression analyses were conducted to identify genes that are expressed at significantly different levels between male and female samples using 29 GTEx v6p tissues with RNA-seq and genotype data available from more than 70 individuals after excluding samples flagged in QC and sex-specific, outlier (that is, breast tissue) and highly correlated tissues14. Only autosomal and X-chromosomal protein-coding or long non-coding RNA genes in Gencode v.19 were included, and all lowly expressed genes were removed (Extended Data Table 1 and Supplementary Methods).
Differential expression analysis between male and female samples was conducted following the voom-limma pipeline41,42,43 available as an R package through Bioconductor (https://bioconductor.org/packages/release/bioc/html/limma.html) using the gene-level read counts as input. The analyses were adjusted for age, three principal components inferred from genotype data using EIGENSTRAT44, sample ischaemic time, surrogate variables45,46 built using the sva R package47, and the cause of death classified into five categories based on the four-point Hardy scale (Supplementary Methods).
To control the FDR, the qvalue R package was used to obtain q values applying the adjustment separately for the differential expression results from each tissue. The null hypothesis was rejected for tests with q values below 0.01.
XY homologue analysis
A list of Y-chromosomal genes with functional counterparts in the X chromosome, that is, X–Y gene pairs, was obtained from ref. 30, which lists 19 ancestral Y chromosome genes that have been retained in the human Y chromosome. After excluding two of the genes (MXRA5Y and OFD1Y), which were annotated as pseudogenes in ref 30, and a further four genes (SRY, RBMY, TSPY and HSFY) that according to ref. 30 have clearly diverged in function from their X-chromosomal homologues, the remaining 13 Y-chromosomal genes were matched with their X-chromosome counterparts using gene-pair annotations given in ref. 30 or by searching for known paralogues of the Y-chromosomal genes. To test for completeness of dosage compensation of the X–Y homologous genes, the sex-bias analysis in GTEx data was repeated replacing the expression of the X-chromosomal counterpart with the combined expression of the X and Y homologues.
Chromatin state analysis
To study the relationship between chromatin states and XCI, we used chromatin state calls from the Roadmap Epigenomics Consortium31. Specifically, we used the chromatin state annotations from the core 15-state model, publicly available at http://egg2.wustl.edu/roadmap/web_portal/chr_state_learning.html#core_15state. We followed our previously published method48 to calculate the covariate-corrected percentage of each gene body assigned to each chromatin state. After pre-processing, we filtered down to the 399 inactive and 86 escape genes on the X chromosome and down to 38 female epigenomes.
To compare the chromatin state profiles of the escape and inactive genes in female samples, we used the one-sided Wilcoxon rank-sum test. Specifically, for each chromatin state, we averaged the chromatin state coverage across the 38 female samples for each gene, and compared that average chromatin state coverage for all 86 escape genes to the average chromatin state coverage for all 399 inactive genes. We performed both one-sided tests, to test for enrichment in escape genes, as well as for enrichment in inactive genes.
Next, we performed simulations to account for possible chromatin state biases, such as the fact that the escape and inactive genes are all from the X chromosome. Specifically, we generated 10,000 randomized simulations where we randomly shuffled the escape or inactive labels on the combined set of 485 genes, while retaining the sizes of each gene set. For each of these simulated escape and inactive gene sets, we calculated both one-sided Wilcoxon rank-sum test P values as described above, and then, we calculated a permutation P value for the real gene sets based on these 10,000 random simulations (Supplementary Methods). Finally, we used Bonferroni multiple hypothesis corrections for our significance thresholds to correct for our 30 tests, one for each of 15 chromatin states, and both possible test directions.
Allele-specific expression
For ASE analysis the allele counts for biallelic heterozygous variants were retrieved from RNA-seq data using GATK ASEReadCounter (v.3.6)39. Heterozygous variants that passed VQSR filtering were first extracted for each sample from WES or WGS VCFs using GATK SelectVariants. The analysis was restricted to biallelic SNPs owing to known issues in mapping bias in RNA-seq against indels39. Sample-specific VCFs and RNA-seq BAMs were inputted to ASEReadCounter requiring minimum base quality of 13 in the RNA-seq data (scRNA-seq samples, GTEX-UPIC) or requiring coverage in the RNA-seq data of each variant to be at least 10 reads, with a minimum base quality of 10 and counting only reads with unique mapping quality (MQ = 60) (clinical muscle samples).
For downstream processing of the scRNA-seq and GTEX-UPIC ASE data, we applied further filters to the data to focus on exonic variation only and to conservatively remove potentially spurious sites (Supplementary Methods), for example, sites with non-unique mappability were removed, and furthermore, after an initial analysis of the ASE data, we subjected 22 of the X-chromosomal ASE sites to manual investigation. For GTEX-UPIC the X-chromosomal ASE data was limited to only one site per gene in case of multiple ASE sites, by selecting the site with coverage >7 reads in the largest number of tissues, to have equal representation of each gene for downstream analyses.
Assessing ASE across tissues
For the GTEX-UPIC individual, for whom ASE data from up to 16 tissues per each ASE site was available, we applied the two-sided hierarchical grouped tissue model (GTM*) implemented in MAMBA v.1.0.0 (refs 49, 50) to ASE data. The Gibbs sampler was run for 200 iterations with a burn-in of 50 iterations.
GTM* is a Bayesian hierarchical model that borrows information across tissues and across variants, and provides parameter estimates that are useful for interpreting global properties of variants. It classifies the sites into ASE states according to their tissue-wide ASE profiles and provides an estimate of the proportion of variants in each of the five different ASE states (strong ASE across all tissues (SNGASE), moderate ASE across all tissues (MODASE), no ASE across all tissues (NOASE) and heterogeneous ASE across tissues (HET1 and HET0)).
To summarize the GTM* output in the context of XCI, SNGASE was considered to reflect full XCI, MODASE and NOASE were taken together to represent partial XCI with similar effects across tissues, and HET1 and HET0 were considered to reflect partial yet heterogeneous patterns of XCI across tissues. To combine estimates from two ASE states, we summed the estimated proportions in each class and subsequently calculated the 95% confidence intervals for each remaining ASE state using Jeffreys’ prior.
Determining XCI status in GTEX-UPIC
In addition to the ASE states provided by the above MAMBA analysis, genic XCI status was assessed by comparing the allelic ratios at each X-chromosomal ASE site in each tissue individually. For each ASE site, the alleles were first mapped to Xa and Xi; the allele with lower combined relative expression across tissues was assumed to be the Xi allele. As an exception, at XIST the higher expressing allele was assumed to be the Xi allele. The significance of Xi expression at each ASE observation was tested using a one-sided binomial test, where the hypothesized probability of success was set at 0.025, that is, the fraction of Xi expression from total expression was expected to be significantly greater than 0.025. To account for multiple testing, a FDR correction was applied, using the qvalue R package, to the P values from the binomial test for each of the 16 tissues separately. Observations with q values <0.01 were considered significant, that is, indicative of incomplete XCI at the given ASE site and tissue.
Biallelic expression in single cells
Biallelic expression in individual cells in the X chromosome was assessed only at ASE sites covered by the minimum of eight reads. A site was considered biallelically expressed when (1) allelic expression >0.05 and (2) the one-sided binomial test indicated allelic expression to be at least nominally significantly greater than 0.025. Only genes with at least two observations of biallelic expression across all cells within a sample were counted as biallelic.
Phasing scRNA-seq data
We assigned each cell to either of two cell populations distinguished by the parental X-chromosome designated for inactivation using genotype phasing. For the YRI samples, where parental genotype data was available, the assignment to the two parental cell populations was unambiguous for all cells where X-chromosomal sites outside PAR1 or frequently biallelic sites were expressed. For 24A, no parental genotype data were available, and we therefore used the correlation structure of the expressed X-chromosomal alleles across the 948 cells to infer the two parental haplotypes using the fact that in individual cells the expressed alleles at the chrX sites subject to full inactivation (that is, the majority chrX ASE sites), are from the X chromosome active in each cell (Supplementary Methods). In other words, while monoallelic expression in scRNA-seq in the autosomes is largely stochastic in origin, in the X chromosome the pattern of monoallelic expression is consistent across cells with the same parental X chromosome active22, unless a gene is expressed also from the inactive X. As such, for the phase inference calculations, we excluded all PAR1 sites and all additional sites that were frequently biallelic, to minimize the contribution of escape genes to the phase estimation. After assigning each informative cell to either of the parental cell populations, the reference and alternate allele reads for each ASE site were mapped to active and inactive allele reads within each sample using the actual or inferred parental haplotypes. The data were first combined per variant by taking the sum of active and inactive counts separately across cells, and further similarly combined per gene, if multiple SNPs per gene were available. For 24A the allele expressed at XIST was assumed the Xi allele, in line with the exclusive Xi expression in the Yoruba samples confirmed using the information on parental haplotypes.
Determining XCI status from scRNA-seq ASE
Before calling XCI status using the Xa and Xi read counts from the phased data aggregated across cells, we excluded all sites without fewer than five cells contributing ASE data at each gene and also all sites with coverage lower than eight reads across cells within each sample. To determine whether the observed Xi expression is significantly different from zero, and therefore indicative of incomplete XCI at the site or gene, we required the Xi to total expression ratio to be significantly (q value <0.01) greater than the hypothesized upper bound for error, 0.025. This threshold was determined using the proportion of miscalled alleles at XIST ASE sites (by definition, XIST should express only alleles from the inactive chrX) in the two YRI samples, which presented with fully skewed XCI, that is, the same active X chromosome across all assessed cells. The median proportion of miscalled XIST alleles was 0, yet one site in one of the samples showed up to 2.5% of other allele calls, and therefore this was chosen as the error margin. FDR correction, conducted using the qvalue R package, was applied to each sample individually. Genes where at least one of the samples showed significant Xi expression were considered partially inactivated, while the remaining were classified as subject to full XCI. Allelic dropout, which is extensive in scRNA-seq19,28, can lead to biases in allelic ratios in individual cells, that is, in our case resulting in false negatives where true escape genes are classified as inactivated, the used approach is based on using aggregate data across several cells and therefore the XCI status estimates are robust to such errors.
ChrX and chrY expression in PAR1
Using the parental origin of each allele reference and alternate allele read counts at PAR1 ASE sites were assigned to X and Y chromosomes (that is, maternally and paternally inherited alleles, respectively). For each sample, the PAR1 ASE data was summarized by gene by taking the sum of X and Y chromosome reads across all informative ASE sites within each gene. Significance of deviation from equal expression was assessed using a two-sided binomial test.
Manual curation of heterozygous variants from ASE analyses
Twenty-two heterozygous variants assessed in chrX ASE analysis were subjected to manual curation because of results in the XCI analysis that were in conflict with previous assignment of the underlying gene to be subject to full XCI. For each sample, BWA-aligned germline BAM files were viewed in IGV using either WGS or WES data. The presence of a number of characteristics called into question the confidence of the variant read alignments and thus the variant itself (Supplementary Methods). Allele balance that deviated significantly from 50:50 was considered suspect and often coincided with the existence of homology between the reference sequence in the region surrounding the variant and another area of the genome, as ascertained using the UCSC browser self-chain track and/or BLAT alignment of variant reads from within IGV. Other sequence-based annotations added to the VCF by HaplotypeCaller were also evaluated in the interests of examining other signatures of ambiguous mapping. The phasing of nearby variants was also considered. If phased variants occurred in the DNA sequencing data that were not assessed in the ASE analysis, those variants were considered suspect.
Data availability
Gene expression and genotype data from the GTEx v6p release are available in dbGaP (study accession phs000424.v6.p1; http://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs000424.v6.p1). Raw RNA-seq data for 24A is available through dbGaP accession number phs001294.v1.p1 (https://www.ncbi.nlm.nih.gov/bioproject/?term=phs001294.v1.p1). The authors declare that all data supporting the findings of this study are available within the paper and its Supplementary Information. Source Data for Figs 2, 3, 4 are provided with the paper.
Change history
07 March 2018
Please see accompanying Corrigendum (http://doi.org/10.1038/nature25993). The Source Data associated with Fig. 2a and d have been replaced. See Supplementary Information to the Corrigendum for the original Source Data for Fig. 2.
References
Carrel, L. & Willard, H. F. X-inactivation profile reveals extensive variability in X-linked gene expression in females. Nature 434, 400–404 (2005)
Cotton, A. M. et al. Analysis of expressed SNPs identifies variable extents of expression from the human inactive X chromosome. Genome Biol. 14, R122 (2013)
Cotton, A. M. et al. Landscape of DNA methylation on the X chromosome reflects CpG density, functional chromatin state and X-chromosome inactivation. Hum. Mol. Genet. 24, 1528–1539 (2015)
Schultz, M. D. et al. Human body epigenome maps reveal noncanonical DNA methylation variation. Nature 523, 212–216 (2015)
Johnston, C. M. et al. Large-scale population study of human cell lines indicates that dosage compensation is virtually complete. PLoS Genet. 4, e9 (2008)
Tukiainen, T. et al. Chromosome X-wide association study identifies loci for fasting insulin and height and evidence for incomplete dosage compensation. PLoS Genet. 10, e1004127 (2014)
Deng, X., Berletch, J. B., Nguyen, D. K. & Disteche, C. M. X chromosome regulation: diverse patterns in development, tissues and disease. Nat. Rev. Genet. 15, 367–378 (2014)
Amos-Landgraf, J. M. et al. X chromosome-inactivation patterns of 1,005 phenotypically unaffected females. Am. J. Hum. Genet. 79, 493–499 (2006)
Lister, R. et al. Global epigenomic reconfiguration during mammalian brain development. Science 341, 1237905 (2013)
Zhang, Y. et al. Transcriptional profiling of human liver identifies sex-biased genes associated with polygenic dyslipidemia and coronary artery disease. PLoS ONE 6, e23506 (2011)
Berletch, J. B. et al. Escape from X inactivation varies in mouse tissues. PLoS Genet. 11, e1005079 (2015)
The GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015)
The GTEx Consortium. Genetic effects on gene expression across tissues. https://doi.org/10.1038/nature24277 (2017)
Melé, M. et al. The human transcriptome across tissues and individuals. Science 348, 660–665 (2015)
Balaton, B. P. & Brown, C. J. Escape artists of the X chromosome. Trends Genet. 32, 348–359 (2016)
Ross, M. T. et al. The DNA sequence of the human X chromosome. Nature 434, 325–337 (2005)
Pál, C., Papp, B. & Hurst, L. D. Highly expressed genes in yeast evolve slowly. Genetics 158, 927–931 (2001)
Winter, E. E., Goodstadt, L. & Ponting, C. P. Elevated rates of protein secretion, evolution, and disease among tissue-specific genes. Genome Res. 14, 54–61 (2004)
Stegle, O., Teichmann, S. A. & Marioni, J. C. Computational and analytical challenges in single-cell transcriptomics. Nat. Rev. Genet. 16, 133–145 (2015)
Chen, G. et al. Single-cell analyses of X chromosome inactivation dynamics and pluripotency during differentiation. Genome Res. 26, 1342–1354 (2016)
Deng, Q., Ramsköld, D., Reinius, B. & Sandberg, R. Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science 343, 193–196 (2014)
Reinius, B. et al. Analysis of allelic expression patterns in clonal somatic cells by single-cell RNA-seq. Nat. Genet. 48, 1430–1435 (2016)
Wang, M., Lin, F., Xing, K. & Liu, L. Random X-chromosome inactivation dynamics in vivo by single-cell RNA sequencing. BMC Genomics 18, 90 (2017)
Wainer-Katsir, K. & Linial, M. Single cell expression data reveal human genes that escape X-chromosome inactivation. Preprint at http://www.biorxiv.org/content/early/2016/10/09/079830 (2016)
Petropoulos, S. et al. Single-cell RNA-seq reveals lineage and X chromosome dynamics in human preimplantation embryos. Cell 165, 1012–1026 (2016)
Villani, A. C. et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science 356, eaah4573 (2017)
Borel, C. et al. Biased allelic expression in human primary fibroblast single cells. Am. J. Hum. Genet. 96, 70–80 (2015)
Kim, J. K., Kolodziejczyk, A. A., Ilicic, T., Teichmann, S. A. & Marioni, J. C. Characterizing noise structure in single-cell RNA-seq distinguishes genuine from technical stochastic allelic expression. Nat. Commun. 6, 8687 (2015)
Marinov, G. K. et al. From single-cell to cell-pool transcriptomes: stochasticity in gene expression and RNA splicing. Genome Res. 24, 496–510 (2014)
Bellott, D. W. et al. Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 508, 494–499 (2014)
Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015)
Xia, Z. et al. A 17q12 allele is associated with altered NK cell subsets and function. J. Immunol. 188, 3315–3322 (2012)
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010)
Lee, M. N. et al. Common genetic variants modulate pathogen-sensing responses in human dendritic cells. Science 343, 1246980 (2014)
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013)
Trombetta, J. J. et al. Preparation of single-cell RNA-seq libraries for next generation sequencing. Curr. Protoc. Mol. Biol. 107, 4.22.1–4.22.17 (2014)
DeLuca, D. S. et al. RNA-SeQC: RNA-seq metrics for quality control and process optimization. Bioinformatics 28, 1530–1532 (2012)
Rozowsky, J. et al. AlleleSeq: analysis of allele-specific expression and binding in a network framework. Mol. Syst. Biol. 7, 522 (2011)
Castel, S. E., Levy-Moonshine, A., Mohammadi, P., Banks, E. & Lappalainen, T. Tools and best practices for data processing in allelic expression analysis. Genome Biol. 16, 195 (2015)
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013)
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014)
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015)
Smyth, G. K. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, 1–25 (2004)
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006)
Leek, J. T. & Storey, J. D. Capturing heterogeneity in gene expression studies by surrogate variable analysis. PLoS Genet. 3, 1724–1735 (2007)
Leek, J. T. & Storey, J. D. A general framework for multiple testing dependence. Proc. Natl Acad. Sci. USA 105, 18718–18723 (2008)
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012)
Yen, A. & Kellis, M. Systematic chromatin state comparison of epigenomes associated with diverse properties including sex and tissue type. Nat. Commun. 6, 7973 (2015)
Pirinen, M. et al. Assessing allele-specific expression across multiple tissues from RNA-seq read data. Bioinformatics 31, 2497–2504 (2015)
Rivas, M. A. et al. Effect of predicted protein-truncating genetic variants on the human transcriptome. Science 348, 666–669 (2015)
Acknowledgements
We thank J. Maller, F. Zhao and M. Lek for technical assistance and P. J. Siponen for assistance with figure design. T.T. was supported by the Academy of Finland (285725), Finnish Cultural Foundation, Orion-Farmos Research Foundation and Emil Aaltonen Foundation. K.J.K. is supported by a NIGMS Fellowship (F32GM115208). This work was supported by NIH grants U54DK105566, R01MH101820 and R01GM104371 to D.G.M. The Genotype-Tissue Expression (GTEx) project was supported by the Common Fund of the Office of the Director of the National Institutes of Health. Additional funds were provided by the NCI, NHGRI, NHLBI, NIDA, NIMH and NINDS. Donors were enrolled at Biospecimen Source Sites funded by NCI\SAIC-Frederick, Inc. (SAIC-F) subcontracts to the National Disease Research Interchange (10XS170), Roswell Park Cancer Institute (10XS171) and Science Care, Inc. (X10S172). The Laboratory, Data Analysis, and Coordinating Center (LDACC) was funded through a contract (HHSN268201000029C) to The Broad Institute; this grant also provided funding to D.G.M. and T.T. Biorepository operations were funded through an SAIC-F subcontract to the Van Andel Institute (10ST1035). Additional data repository and project management were provided by SAIC-F (HHSN261200800001E). The Brain Bank was supported by supplements to University of Miami grants DA006227 and DA033684 and to contract N01MH000028. Statistical Methods development grants were made to the University of Geneva (MH090941 and MH101814), the University of Chicago (MH090951, MH090937, MH101820 and MH101825), the University of North Carolina, Chapel Hill (MH090936 and MH101819), Harvard University (MH090948), Stanford University (MH101782), Washington University St. Louis (MH101810) and the University of Pennsylvania (MH101822).
Author information
Authors and Affiliations
Consortia
Contributions
T.T. and D.G.M. designed the study. A.-C.V. designed and conducted the scRNA-seq experiments. T.T., A.Y., M.A.R., M.A., L.G., M.F. and B.B.C. analysed the data. J.L.M., R.S., S.E.C., A.K., K.J.K., F.A., A.B., T.L., A.R., K.G.A., N.H. and D.G.M. provided tools and reagents. T.T. and D.G.M. wrote the manuscript with input from other authors.
Corresponding authors
Ethics declarations
Competing interests
D.G.M. is a founder with equity in Goldfinch Bio. The authors declare no other competing financial interests.
Additional information
Reviewer Information Nature thanks A. Clark and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Assessment of skew in XCI in GTEx female samples (v3 analysis release).
a, The estimated skew in XCI by tissue across individuals. b, The skew in XCI by individual across available tissue samples. The number in brackets after the tissue or sample name indicates the number of individuals or tissues, respectively, contributing to each box plot. Details of the analysis can be found in the Supplementary Note.
Extended Data Figure 2 Comparison of expression characteristics between reported genic XCI categories in the GTEx data.
a, The statistics for the comparison of the proportion of significantly biased (FDR <1%) genes by reported XCI status. Distributions are illustrated in Fig. 2b. n = 29 for all comparisons. b, The statistics for the comparison of the consistency in effect sizes across tissues. Distributions are illustrated in Fig. 2c. Only genes expressed in at least five of the 29 tissues are included. c, Number of tissues showing significant sex bias (FDR <1%) per gene by reported XCI status. d, Statistics for the comparison illustrated in c. e, Number of tissues in which genes are expressed by reported XCI status. f, Statistics for the comparison illustrated in e. All P values are from two-sided Wilcoxon rank-sum tests, except for a, where a paired, two-sided Wilcoxon rank-sum test was used. Only genes assessed for sex bias in at least one tissue are included, unless otherwise stated.
Extended Data Figure 3 Change in the proportion of discovered sex-biased genes by XCI category with varying q value cut-offs.
a, The proportion of sex-biased genes across tissues. Here a gene is classified as sex-biased if the q value for association falls below the given threshold in at least one tissue. b–f, Examples of the change in the proportion of sex-biased expression in individual tissues. The dashed black line indicates the FDR <1% cut-off applied in the analyses to determine sex-biased expression. ADPSBQ, adipose, subcutaneous; WHLBLD, whole blood; BRNCTXA, brain, cortex; SKINNS, skin not sun exposed (suprapubic).
Extended Data Figure 4 Heat map representation of male–female expression differences in all assessed X-chromosomal genes (n = 681) across 29 GTEx tissues.
The colour scale displays the direction of sex bias, with red colour indicating higher female expression. Genes that were too weakly expressed to be assessed in a given tissue type in the sex bias analysis are coloured grey. Dots mark the observations where sex bias was significant at FDR <1%.
Extended Data Figure 5 Comparison of expression characteristics between Xp and Xq, the evolutionary newer and older regions of chrX, respectively, by XCI status and for the whole chromosome.
a, b, The level of median expression across GTEx tissues in log2 RPKM units. c, d, The breadth of expression measured as the number of tissues (max = 29) in which genes are expressed (median expression across samples >0.1 RPKM and expressed in more than 10 individuals at >1 counts per million). P values are calculated using the Wilcoxon rank-sum test. All genes expressed in at least one tissue are included in the comparisons.
Extended Data Figure 6 X-chromosomal RNA-seq and WGS data in the GTEx donor with fully skewed XCI (GTEX-UPIC).
a, Allelic expression in chrX in 16 RNA-sequenced tissue samples available from the donor. Dashed red lines indicate PAR1 and PAR2 boundaries. b, Allele balance and allele depth across chrX in WGS for GTEX-UPIC and two female and one male GTEx WGS samples that were randomly chosen.
Extended Data Figure 7 Expressed alleles at biallelically expressed ASE sites in scRNA-seq.
a, X-chromosomal genes repeatedly biallelic in scRNA-seq (see Methods for details). b, Illustration of the relative expression from the two alleles at all X-chromosomal ASE sites that were repeatedly biallelically expressed across cells in either of the two scRNA-seq samples that showed random XCI (Y035 and 24A). Narrow white lines separate observations from individual cells.
Extended Data Figure 8 Assessment of the level of Xi expression at escape genes and in different regions of the X chromosome.
a, The ratio of Xi-to-Xa expression in the single-cell samples (left; each circle represents a sample), in the skewed XCI donor from GTEx (middle; each circle represents a tissue), and the female-to-male ratio in expression (right; each circle represents a tissue) at reported escape genes. Genes are ordered according to their location in the X chromosome with genes in the pseudoautosomal region residing in the top part. A dark border around a circle indicates that there was significant evidence for Xi expression greater than the baseline in the given sample or tissue (left and middle) or significant sex-bias in the given tissue (right). Given some outliers, for example, XIST, the Xi-to-Xa ratio is capped at 1.75 and female-to-male ratio at 2.25. b, The relative expression arising from the X and Y chromosome at PAR1 genes in skeletal muscle in eight males. The allelic expression at these genes was assigned to the two chromosomes using parental genotypes available for these samples (see Methods for details). The dashed line at 0.5 indicates the point where expression from X and Y chromosomes is equal. The error bars give the 95% confidence intervals for the observed read ratio. c, Heat map representation of the change in pattern of sex-bias at 13 X–Y homologous gene pairs (see Methods for details) in nonPAR from only including the X-chromosomal expression (heat map on the left) to accounting for the Y-chromosomal expression (heat map on the right). The colour scale displays the direction of sex-bias with red colour indicating higher female expression. Genes that were too lowly expressed in the given tissue type to be assessed in the sex-bias analysis are coloured grey. Dots mark the observations where sex-bias was significant at FDR <1%. The grey bars on top of the heat maps indicate the location of the gene in the X chromosome: dark grey indicating Xp and lighter grey Xq.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, a Supplementary Discussion, the Supplementary Table guide and a Supplementary Note describing the analysis of skew in XCI in GTEx female samples. (PDF 664 kb)
Supplementary Tables
This file contains Supplementary Tables 1-14. (ZIP 1653 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International (CC BY 4.0) licence. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tukiainen, T., Villani, AC., Yen, A. et al. Landscape of X chromosome inactivation across human tissues. Nature 550, 244–248 (2017). https://doi.org/10.1038/nature24265
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24265
This article is cited by
-
Testosterone deficiency promotes arterial stiffening independent of sex chromosome complement
Biology of Sex Differences (2024)
-
Overcoming genetic and cellular complexity to study the pathophysiology of X-linked intellectual disabilities
Journal of Neurodevelopmental Disorders (2024)
-
X-chromosome inactivation in human iPSCs provides insight into X-regulated gene expression in autosomes
Genome Biology (2024)
-
Insights into estrogen impact in oral health & microbiome in COVID-19
BMC Microbiology (2024)
-
Identification of skewed X chromosome inactivation using exome and transcriptome sequencing in patients with suspected rare genetic disease
BMC Genomics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.