Abstract
Mammalian organs vary widely in regenerative capacity. Poorly regenerative organs, such as the heart are particularly vulnerable to organ failure. Once established, heart failure commonly results in mortality1. The Hippo pathway, a kinase cascade that prevents adult cardiomyocyte proliferation and regeneration2, is upregulated in human heart failure. Here we show that deletion of the Hippo pathway component Salvador (Salv) in mouse hearts with established ischaemic heart failure after myocardial infarction induces a reparative genetic program with increased scar border vascularity, reduced fibrosis, and recovery of pumping function compared with controls. Using translating ribosomal affinity purification, we isolate cardiomyocyte-specific translating messenger RNA. Hippo-deficient cardiomyocytes have increased expression of proliferative genes and stress response genes, such as the mitochondrial quality control gene, Park2. Genetic studies indicate that Park2 is essential for heart repair, suggesting a requirement for mitochondrial quality control in regenerating myocardium. Gene therapy with a virus encoding Salv short hairpin RNA improves heart function when delivered at the time of infarct or after ischaemic heart failure following myocardial infarction was established. Our findings indicate that the failing heart has a previously unrecognized reparative capacity involving more than cardiomyocyte renewal.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Loehr, L. R., Rosamond, W. D., Chang, P. P., Folsom, A. R. & Chambless, L. E. Heart failure incidence and survival (from the Atherosclerosis Risk in Communities study). Am. J. Cardiol. 101, 1016–1022 (2008)
Halder, G. & Johnson, R. L. Hippo signaling: growth control and beyond. Development 138, 9–22 (2011)
Lopez, A. D., Mathers, C. D., Ezzati, M., Jamison, D. T. & Murray, C. J. Global and regional burden of disease and risk factors, 2001: systematic analysis of population health data. Lancet 367, 1747–1757 (2006)
Lenneman, A. J. & Birks, E. J. Treatment strategies for myocardial recovery in heart failure. Curr. Treat. Options Cardiovasc. Med. 16, 287 (2014)
Birks, E. J. Molecular changes after left ventricular assist device support for heart failure. Circ. Res. 113, 777–791 (2013)
Braunwald, E. Heart failure. JACC Heart Fail. 1, 1–20 (2013)
Halder, G., Dupont, S. & Piccolo, S. Transduction of mechanical and cytoskeletal cues by YAP and TAZ. Nat. Rev. Mol. Cell Biol. 13, 591–600 (2012)
Heallen, T. et al. Hippo signaling impedes adult heart regeneration. Development 140, 4683–4690 (2013)
Morikawa, Y. et al. Actin cytoskeletal remodeling with protrusion formation is essential for heart regeneration in Hippo-deficient mice. Sci. Signal. 8, ra41 (2015)
Tao, G. et al. Pitx2 promotes heart repair by activating the antioxidant response after cardiac injury. Nature 534, 119–123 (2016)
Matsuda, T. et al. NF2 activates Hippo signaling and promotes ischemia/reperfusion injury in the heart. Circ. Res. 119, 596–606 (2016)
Del Re, D. P. et al. Mst1 promotes cardiac myocyte apoptosis through phosphorylation and inhibition of Bcl-xL. Mol. Cell 54, 639–650 (2014)
Gao, X. M., Dart, A. M., Dewar, E., Jennings, G. & Du, X. J. Serial echocardiographic assessment of left ventricular dimensions and function after myocardial infarction in mice. Cardiovasc. Res. 45, 330–338 (2000)
Doupé, D. P. et al. A single progenitor population switches behavior to maintain and repair esophageal epithelium. Science 337, 1091–1093 (2012)
Leask, A. Getting to the heart of the matter: new insights into cardiac fibrosis. Circ. Res. 116, 1269–1276 (2015)
Pontén, A., Folestad, E. B., Pietras, K. & Eriksson, U. Platelet-derived growth factor D induces cardiac fibrosis and proliferation of vascular smooth muscle cells in heart-specific transgenic mice. Circ. Res. 97, 1036–1045 (2005)
Heineke, J. & Molkentin, J. D. Regulation of cardiac hypertrophy by intracellular signalling pathways. Nat. Rev. Mol. Cell Biol. 7, 589–600 (2006)
Meredith, A. J. et al. Circulating biomarker responses to medical management vs. mechanical circulatory support in severe inotrope-dependent acute heart failure. ESC Heart Fail. 3, 86–96 (2016)
Kostin, S., Hein, S., Arnon, E., Scholz, D. & Schaper, J. The cytoskeleton and related proteins in the human failing heart. Heart Fail. Rev. 5, 271–280 (2000)
Kim, S. Y., Morales, C. R., Gillette, T. G. & Hill, J. A. Epigenetic regulation in heart failure. Curr. Opin. Cardiol. 31, 255–265 (2016)
Jessup, M. & Brozena, S. Heart failure. N. Engl. J. Med. 348, 2007–2018 (2003)
Marín-García, J. & Akhmedov, A. T. Mitochondrial dynamics and cell death in heart failure. Heart Fail. Rev. 21, 123–136 (2016)
Dorn, G. W., II . Parkin-dependent mitophagy in the heart. J. Mol. Cell. Cardiol. 95, 42–49 (2016)
Kubli, D. A. et al. Parkin protein deficiency exacerbates cardiac injury and reduces survival following myocardial infarction. J. Biol. Chem. 288, 915–926 (2013)
Xin, M. et al. Hippo pathway effector Yap promotes cardiac regeneration. Proc. Natl Acad. Sci. USA 110, 13839–13844 (2013)
Itoh, N., Ohta, H., Nakayama, Y. & Konishi, M. Roles of FGF signals in heart development, health, and disease. Front. Cell Dev. Biol. 4, 110 (2016)
Korf-Klingebiel, M. et al. Conditional transgenic expression of fibroblast growth factor 9 in the adult mouse heart reduces heart failure mortality after myocardial infarction. Circulation 123, 504–514 (2011)
Singla, D. K., Singla, R. D., Abdelli, L. S. & Glass, C. Fibroblast growth factor-9 enhances M2 macrophage differentiation and attenuates adverse cardiac remodeling in the infarcted diabetic heart. PLoS ONE 10, e0120739 (2015)
Gong, G. et al. Parkin-mediated mitophagy directs perinatal cardiac metabolic maturation in mice. Science 350, aad2459 (2015)
Patterson, M. et al. Frequency of mononuclear diploid cardiomyocytes underlies natural variation in heart regeneration. Nat. Genet. 49, 1346–1353 (2017)
Bersell, K. et al. Moderate and high amounts of tamoxifen in αMHC-MerCreMer mice induce a DNA damage response, leading to heart failure and death. Dis. Model. Mech. 6, 1459–1469 (2013)
Koitabashi, N. et al. Avoidance of transient cardiomyopathy in cardiomyocyte-targeted tamoxifen-induced MerCreMer gene deletion models. Circ. Res. 105, 12–15 (2009)
Pugach, E. K., Richmond, P. A., Azofeifa, J. G., Dowell, R. D. & Leinwand, L. A. Prolonged Cre expression driven by the α-myosin heavy chain promoter can be cardiotoxic. J. Mol. Cell. Cardiol. 86, 54–61 (2015)
Sam, F. et al. Progressive left ventricular remodeling and apoptosis late after myocardial infarction in mouse heart. Am. J. Physiol. Heart Circ. Physiol. 279, H422–H428 (2000)
Wang, J. et al. A simple and fast experimental model of myocardial infarction in the mouse. Tex. Heart Inst. J. 33, 290–293 (2006)
Nascimento, D. S. et al. MIQuant – semi-automation of infarct size assessment in models of cardiac ischemic injury. PLoS ONE 6, e25045 (2011)
Bergmann, O. et al. Dynamics of cell generation and turnover in the human heart. Cell 161, 1566–1575 (2015)
Sanz, E. et al. Cell-type-specific isolation of ribosome-associated mRNA from complex tissues. Proc. Natl Acad. Sci. USA 106, 13939–13944 (2009)
Giudice, J. et al. Alternative splicing regulates vesicular trafficking genes in cardiomyocytes during postnatal heart development. Nat. Commun. 5, 3603 (2014)
Dow, L. E. et al. A pipeline for the generation of shRNA transgenic mice. Nat. Protocols 7, 374–393 (2012)
Morikawa, Y., Heallen, T., Leach, J., Xiao, Y. & Martin, J. F. Dystrophin–glycoprotein complex sequesters Yap to inhibit cardiomyocyte proliferation. Nature 547, 227–231 (2017)
Acknowledgements
This work was supported by grants from the National Institutes of Health (DE023177, HL127717, HL130804, HL118761 (J.F.M.); F31HL136065 (M.C.H.) and 5T32HL007676-23 (J.P.L.)), Vivian L. Smith Foundation (J.F.M.), State of Texas funding (J.F.M. and J.T.W.), LeDucq Foundation Transatlantic Networks of Excellence in Cardiovascular Research (14CVD01) ‘Defining the genomic topology of atrial fibrillation’ (J.F.M.). Supported by Intellectual and Developmental Disabilities Research Center grant number 1U54 HD083092 from the Eunice Kennedy Shriver National Institute of Child Health & Human Development and the Mouse Phenotyping Core at Baylor College of Medicine (U54 HG006348). T.H. was supported by American Heart Association Scientist Development Grant (16SDG26460001). This work was also supported in part by Neuroconnectivity core and Optical Imaging and Vital Microscopy core at Baylor College of Medicine. N. Stancel provided editorial assistance.
Author information
Authors and Affiliations
Contributions
J.P.L., J.T.W., and J.F.M. conceived and designed experiments and interpreted data. J.P.L., T.H., M.Z., M.C.H., Y.M., and M.R. performed experiments. J.P.L. and J.F.M. analysed data and compiled figures. A.S. provided human tissue samples J.P.L., J.T.W., and J.F.M. wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks M. Schneider and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Activated Hippo signalling in human heart failure.
a–c, Western blots of human heart samples. Ctrl: non-failing non-transplantable, n = 6 (a–c). HF: non-ischaemic idiopathic cardiomyopathy in end-stage heart failure, n = 6 (a, b). iHF: ischaemic heart in end-stage heart failure, n = 6 (c). Quantification presented in Fig. 1.
Extended Data Figure 2 Mouse model of systolic heart failure.
a–d, Systolic diameter (a), diastolic diameter (b), systolic volume (c), and diastolic volume (d); n values indicated in Fig. 1a; ANOVA, Tukey’s pairwise post-hoc test. e, f, Haematoxylin and eosin oedema liquid (pink transudate fluid) in lung tissue 3 weeks after myocardial infarction, sham (n = 3) (e) and myocardial infarction (n = 5) (f); scale bar, 50 μm. g, h, Prussian blue haemosiderin (blue) in lung tissue 3 weeks after myocardial infarction, sham (n = 3) (g) and myocardial infarction (n = 5) (h); scale bar, 50 μm. i, Natriuretic peptide B (BNP) in blood serum 3 weeks after myocardial infarction, Mann–Whitney U-test (i). j, Weight gain 3 weeks after myocardial infarction (j), t-test. k, l, Longitudinal echocardiography beginning 3 weeks after myocardial infarction; data are a subset of Fig. 2c. Control sham and control myocardial infarction samples were split by Cre genotype or by injection type, indicated in parenthesis (k). No significant effect of Cre or tamoxifen (Tam) was observed; ANOVA, Tukey’s pairwise post-hoc test (k). Data are mean ± s.e.m., P > 0.05 non-significant (NS), *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 3 Histological analysis at 3 weeks after myocardial infarction.
Masson’s trichrome of serial sagittal sections 3 weeks after myocardial infarction, no tamoxifen was delivered; genotype is indicated, n = 3 per group, scar boundaries (open arrows), cardiomyocytes in the ischaemic region (solid arrows); scale bar, 2 mm.
Extended Data Figure 4 Histological analysis at 4 weeks after myocardial infarction.
Masson’s trichrome of serial sagittal sections 4 weeks after myocardial infarction, 1 week after tamoxifen; control (αMHC-mcm; ROSAmT/mG) and SalvCKO (αMHC-mcm; ROSAmT/mG; Salvfl/fl), n = 3 per group, scar boundaries (open arrows), cardiomyocytes in the ischaemic region (solid arrows); scale bar, 2 mm.
Extended Data Figure 5 Histological analysis at 6 weeks after myocardial infarction.
Masson’s trichrome of serial sagittal sections 6 weeks after myocardial infarction, 3 weeks after tamoxifen; control (αMHC-mcm; ROSAmT/mG) and SalvCKO (αMHC-mcm; ROSAmT/mG; Salvfl/fl), n = 3 per group, scar boundaries (open arrows), cardiomyocytes in the ischaemic region (solid arrows); scale bar, 2 mm.
Extended Data Figure 6 Vessel growth in the border zone of Hippo-deficient mouse hearts.
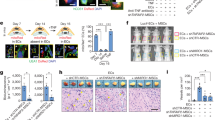
a, qPCR of known markers of heart failure at 6 weeks after myocardial infarction; myosin heavy chain 6 (Myh6), myosin heavy chain 7 (Myh7), natriuretic peptide A (Nppa), natriuretic peptide B (Nppb), n = 3 per group; ANOVA, Bonferroni’s post-hoc test. b, Masson’s trichrome (scale bar, 100 μm) and immunofluorescence staining for isolectin B4 (scale bar, 25 μm) and CD31 (scale bar, 25 μm), control (n = 3) SalvCKO (n = 5). c–e, Quantification 9 weeks after myocardial infarction in the border zone for capillary density (c), isolectin+ vessels (d), and CD31+ cells (e), control (n = 3) SalvCKO (n = 5), Mann–Whitney U-test. f, qPCR of angiogenic growth factors, cardiomyocyte-specific TRAP RNA, 6 weeks after myocardial infarction, angiopoietin 1 (Angpt1) and 2 (Angpt2), fibroblast growth factor 14 (Fgf14) and 18 (Fgf18), vascular endothelial growth factor B (Vegfb) and C (Vegfc), n = 3 per group; ANOVA, Bonferroni’s post-hoc test. Data are mean ± s.e.m., P > 0.05 non-significant (NS), *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 7 TRAP RNA sequencing reproducibility.
a, Reproducibility correlation matrices of the RNA-seq read count, linear regression, n = 3 per group. b, c, Plot of the per-gene s.d. across samples, against the rank (mean) and read count, variance stabilizing transformation, total RNA-seq (b), and TRAP RNA-seq (c). d, log2(fold change) between TRAP and total RNA-seq for control myocardial infarction and SalvCKO myocardial infarction was highly correlated.
Extended Data Figure 8 A reparative molecular response to heart failure in Hippo-deficient hearts.
a, Gene lists of the top 10 genes with the highest fold change in each GO category for total RNA: SalvCKO myocardial infarction versus control myocardial infarction. b, Boxplot of the normalized read count for Tnnt2, Cdh5, and Malat1. The bars indicate minimum and maximum values. c, Volcano plot TRAP-seq: control myocardial infarction versus control sham. d, e, GO upregulated (d) and downregulated (e) genes. f, g, Gene lists of the top ten genes with the highest fold change in each GO category for TRAP RNA: control myocardial infarction versus control sham (f); TRAP RNA: SalvCKO myocardial infarction versus control myocardial infarction (g).
Extended Data Figure 9 Requirement of Park2 in the regenerating mouse heart.
a–c, Human heart western blots; quantification presented in Fig. 4, tubulin# blot is repeated from Extended Data Fig. 1. d, Scar size 21 days after myocardial infarction in P1 Park2 wild-type (+/+) and null (−/−) mice; Mann–Whitney U-test. e, f, Echocardiography: ejection fraction (EF) (e), and fractional shortening (FS) (f); ANOVA, Bonferroni’s post-hoc test. g, Cardiomyocyte cell size measured by cross-sectional area, mean (red dashed line), t-test. h, Masson’s trichrome, 21 days after myocardial infarction in P1 Park2 wild-type (+/+) (n = 8) and null (−/−) (n = 4) mice; scale bar, 2 mm. i, Summary of results indicating Park2 is necessary for cardiac regeneration. Data are mean ± s.e.m., P > 0.05 non-significant (NS), *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Figure 10 Requirement of Park2 in the P8 Hippo-deficient regenerating mouse heart.
a, Mitochondrial DNA content 4 days after myocardial infarction in P8 control and SalvCKO (n = 3 per group). b, Park2 protein levels in border zone (BZ) and distal zone (DZ) myocardium at 4 days after myocardial infarction in P8 control and SalvCKO (n = 3 per group). c, Scar size 21 days after myocardial infarction in P8 Park2 wild-type (+/+) and mutant (mut; −/− or +/−) mice, in combination with SalvCKO; ANOVA, Bonferroni’s post-hoc test. d, e, Echocardiography: ejection fraction (EF) (d) and fractional shortening (FS) (e); ANOVA, Bonferroni’s post-hoc test. f, Masson’s trichrome, 21 days after myocardial infarction in P8 Park2 mutant (n = 20) and SalvCKO; Park2 double-mutant mice (n = 15); scale bar, 2 mm. g, Summary of results indicating Park2 is necessary for regeneration. Data are mean ± s.e.m., P > 0.05 non-significant (NS), *P < 0.05, **P < 0.01, ***P < 0.001.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1 and Supplementary Table 1. Supplementary Figure 1 shows the uncropped scans with size marker indications and Supplementary Table 1 shows a list of qPCR primers. (PDF 4977 kb)
Rights and permissions
About this article
Cite this article
Leach, J., Heallen, T., Zhang, M. et al. Hippo pathway deficiency reverses systolic heart failure after infarction. Nature 550, 260–264 (2017). https://doi.org/10.1038/nature24045
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24045
This article is cited by
-
Animal models to study cardiac regeneration
Nature Reviews Cardiology (2024)
-
Toward drug-induced heart regeneration
Nature Cardiovascular Research (2024)
-
Regeneration of the heart: from molecular mechanisms to clinical therapeutics
Military Medical Research (2023)
-
Recent advances in regulating the proliferation or maturation of human-induced pluripotent stem cell-derived cardiomyocytes
Stem Cell Research & Therapy (2023)
-
Optogenetic control of YAP can enhance the rate of wound healing
Cellular & Molecular Biology Letters (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.