Abstract
The ability to directly uncover the contributions of genes to a given phenotype is fundamental for biology research. However, ostensibly homogeneous cell populations exhibit large clonal variance1,2 that can confound analyses and undermine reproducibility3. Here we used genome-saturated mutagenesis to create a biobank of over 100,000 individual haploid mouse embryonic stem (mES) cell lines targeting 16,970 genes with genetically barcoded, conditional and reversible mutations. This Haplobank is, to our knowledge, the largest resource of hemi/homozygous mutant mES cells to date and is available to all researchers. Reversible mutagenesis overcomes clonal variance by permitting functional annotation of the genome directly in sister cells. We use the Haplobank in reverse genetic screens to investigate the temporal resolution of essential genes in mES cells, and to identify novel genes that control sprouting angiogenesis and lineage specification of blood vessels. Furthermore, a genome-wide forward screen with Haplobank identified PLA2G16 as a host factor that is required for cytotoxicity by rhinoviruses, which cause the common cold. Therefore, clones from the Haplobank combined with the use of reversible technologies enable high-throughput, reproducible, functional annotation of the genome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Cahan, P. & Daley, G. Q. Origins and implications of pluripotent stem cell variability and heterogeneity. Nat. Rev. Mol. Cell Biol. 14, 357–368 (2013)
Hou, Y. et al. Single-cell triple omics sequencing reveals genetic, epigenetic, and transcriptomic heterogeneity in hepatocellular carcinomas. Cell Res. 26, 304–319 (2016)
Begley, C. G. & Ellis, L. M. Drug development: raise standards for preclinical cancer research. Nature 483, 531–533 (2012)
Justice, M. J., Noveroske, J. K., Weber, J. S., Zheng, B. & Bradley, A. Mouse ENU mutagenesis. Hum. Mol. Genet. 8, 1955–1963 (1999)
Robertson, E., Bradley, A., Kuehn, M. & Evans, M. Germ-line transmission of genes introduced into cultured pluripotential cells by retroviral vector. Nature 323, 445–448 (1986)
Elbashir, S. M. et al. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411, 494–498 (2001)
Brummelkamp, T. R., Bernards, R. & Agami, R. A system for stable expression of short interfering RNAs in mammalian cells. Science 296, 550–553 (2002)
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012)
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR–Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826 (2013)
Morgens, D. W., Deans, R. M., Li, A. & Bassik, M. C. Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes. Nat. Biotechnol. 34, 634–636 (2016)
Evers, B. et al. CRISPR knockout screening outperforms shRNA and CRISPRi in identifying essential genes. Nat. Biotechnol. 34, 631–633 (2016)
Rouhani, F. et al. Genetic background drives transcriptional variation in human induced pluripotent stem cells. PLoS Genet. 10, e1004432 (2014)
Fedorov, L. M., Haegel-Kronenberger, H. & Hirchenhain, J. A comparison of the germline potential of differently aged ES cell lines and their transfected descendants. Transgenic Res. 6, 223–231 (1997)
Echeverri, C. J. et al. Minimizing the risk of reporting false positives in large-scale RNAi screens. Nat. Methods 3, 777–779 (2006)
Schnütgen, F. et al. Genomewide production of multipurpose alleles for the functional analysis of the mouse genome. Proc. Natl Acad. Sci. USA 102, 7221–7226 (2005)
Schnütgen, F. et al. Enhanced gene trapping in mouse embryonic stem cells. Nucleic Acids Res. 36, e133 (2008)
Mayasari, N. I. et al. Mixture of differentially tagged Tol2 transposons accelerates conditional disruption of a broad spectrum of genes in mouse embryonic stem cells. Nucleic Acids Res. 40, e97 (2012)
Ivics, Z., Hackett, P. B., Plasterk, R. H. & Izsvák, Z. Molecular reconstruction of sleeping beauty, a Tc1-like transposon from fish, and its transposition in human cells. Cell 91, 501–510 (1997)
Bellen, H. J. et al. The Drosophila gene disruption project: progress using transposons with distinctive site specificities. Genetics 188, 731–743 (2011)
Ivics, Z. et al. Transposon-mediated genome manipulation in vertebrates. Nat. Methods 6, 415–422 (2009)
Torres, M. et al. An α-E-catenin gene trap mutation defines its function in preimplantation development. Proc. Natl Acad. Sci. USA 94, 901–906 (1997)
Reithmayer, M., Reischl, A., Snyers, L. & Blaas, D. Species-specific receptor recognition by a minor-group human rhinovirus (HRV): HRV serotype 1A distinguishes between the murine and the human low-density lipoprotein receptor. J. Virol. 76, 6957–6965 (2002)
Hofer, F. et al. Members of the low density lipoprotein receptor family mediate cell entry of a minor-group common cold virus. Proc. Natl Acad. Sci. USA 91, 1839–1842 (1994)
Jaworski, K. et al. AdPLA ablation increases lipolysis and prevents obesity induced by high-fat feeding or leptin deficiency. Nat. Med. 15, 159–168 (2009)
Duncan, R. E., Sarkadi-Nagy, E., Jaworski, K., Ahmadian, M. & Sul, H. S. Identification and functional characterization of adipose-specific phospholipase A2 (AdPLA). J. Biol. Chem. 283, 25428–25436 (2008)
Tsutsumi, S. et al. Endoplasmic reticulum stress response is involved in nonsteroidal anti-inflammatory drug-induced apoptosis. Cell Death Differ. 11, 1009–1016 (2004)
Uyama, T. et al. Interaction of phospholipase A/acyltransferase-3 with Pex19p: a possible involvement in the down-regulation of peroxisomes. J. Biol. Chem. 290, 17520–17534 (2015)
Staring, J. et al. PLA2G16 represents a switch between entry and clearance of Picornaviridae. Nature 541, 412–416 (2017)
Potente, M., Gerhardt, H. & Carmeliet, P. Basic and therapeutic aspects of angiogenesis. Cell 146, 873–887 (2011)
del Toro, R. et al. Identification and functional analysis of endothelial tip cell-enriched genes. Blood 116, 4025–4033 (2010)
Strasser, G. A., Kaminker, J. S. & Tessier-Lavigne, M. Microarray analysis of retinal endothelial tip cells identifies CXCR4 as a mediator of tip cell morphology and branching. Blood 115, 5102–5110 (2010)
Jakobsson, L. et al. Endothelial cells dynamically compete for the tip cell position during angiogenic sprouting. Nat. Cell Biol. 12, 943–953 (2010)
Hellström, M. et al. Dll4 signalling through Notch1 regulates formation of tip cells during angiogenesis. Nature 445, 776–780 (2007)
Valdimarsson, G., De Sousa, P. A., Beyer, E. C., Paul, D. L. & Kidder, G. M. Zygotic expression of the connexin43 gene supplies subunits for gap junction assembly during mouse preimplantation development. Mol. Reprod. Dev. 30, 18–26 (1991)
Pitulescu, M. E., Schmidt, I., Benedito, R. & Adams, R. H. Inducible gene targeting in the neonatal vasculature and analysis of retinal angiogenesis in mice. Nat. Protoc. 5, 1518–1534 (2010)
Chaytor, A. T., Martin, P. E., Edwards, D. H. & Griffith, T. M. Gap junctional communication underpins EDHF-type relaxations evoked by ACh in the rat hepatic artery. Am. J. Physiol. Heart Circ. Physiol. 280, H2441–H2450 (2001)
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010)
Baker, M. 1,500 scientists lift the lid on reproducibility. Nature 533, 452–454 (2016)
Elling, U. et al. Haplobank methods collection. Protoc. Exch. http://dx.doi.org/10.1038/protex.2017.104 (2017)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010)
Jentsch, I., Adler, I. D., Carter, N. P. & Speicher, M. R. Karyotyping mouse chromosomes by multiplex-FISH (M-FISH). Chromosome Res. 9, 211–214 (2001)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009)
Roberts, A., Trapnell, C., Donaghey, J., Rinn, J. L. & Pachter, L. Improving RNA-seq expression estimates by correcting for fragment bias. Genome Biol. 12, R22 (2011)
Mohn, F. et al. Lineage-specific polycomb targets and de novo DNA methylation define restriction and potential of neuronal progenitors. Mol. Cell 30, 755–766 (2008)
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015)
Gombash Lampe, S. E., Kaspar, B. K. & Foust, K. D. Intravenous injections in neonatal mice. J. Vis. Exp. 93, e52037 (2014)
Golczak, M. et al. Structural basis for the acyltransferase activity of lecithin:retinol acyltransferase-like proteins. J. Biol. Chem. 287, 23790–23807 (2012)
Acknowledgements
We thank all members of our laboratories, IMBA/IMP and VBCF services for support and Life Science Editors for assistance; B. Knapp, I. Filipuzzi and T. Aust for clone picking, N. R. Movva and T. Bouwmeester (NIBR) for support, and K. Handler for the differentiation protocols. The Haplobank is funded by the Austrian National Bank (OeNB), an Advanced ERC grant and Era of Hope/National Coalition against Breast Cancer/DoD (to J.M.P.). U.E. is a Wittgenstein Prize fellow. D.B. is supported by FWF P23308-B13. A.S. is supported by an ERC Consolidator Grant, Boehringer Ingelheim and FFG.
Author information
Authors and Affiliations
Contributions
U.E. generated the haploid library with technical support from A.Lei., C.H., J.L., M.H., A.-M.H., M.K., N.S., E.W., N.v.d.L., D.H., R.R. and D.E. U.E., R.A.W. and A.Leo. characterized cell lines. A.Lei., G.M., U.E., D.B. and T.D. performed rhinovirus work. A.S., T.B. and T.M. wrote the bioinformatics algorithms and set up the Haplobank website. S.Z. performed RACE experiments, F.Y. and B.F. performed karyotyping experiments and C.A.A. supported standardization. J.A.Z.M. and O.B. performed ATAC-sequencing. Z.I. advized on mutagenesis vectors. R.A.W., I.M.A., D.A., A.Leo. and H.G. performed blood vessel experiments. U.E. and J.M.P. coordinated the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks S. Narumiya and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Stem cell properties of the haploid subclone AN3-12.
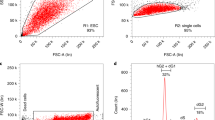
a, b, Various parthenogenic cell lines derived from independent embryos from an outcross of 129/Sv and C57BL/6 and thus containing different genomic backgrounds for different chromosomes were allowed to form embryoid bodies by placing 1,000 cells per hanging drop. We observed downregulation of pluripotency marker genes (a) and upregulation of markers from all three germ layers (b) in all cell lines assayed on day 0 (d0), day 5 (d5) and day 12 (d12). The HMSc2 subclone AN3-12 was chosen for further study based on its growth properties in serum/LIF and absence of feeders. Data are shown as individual data points of n = 2 technical replicates together the mean ± s.d. of one representative experiment. c, Growth curve of AN3-12 in the presence and absence of LIF. Data are shown as individual data points and mean values (lines) of three biological replicates. d, FACS analysis of chromosome content of AN3-12 cells (in LIF, same experiment as shown in c) shows the decrease in haploid (1n) cells from 35.5% to 24.9% during the seven-day culture period. e, AN3-12 cells, cultured as in c, maintain a robust haploid population when analysed on day 17 in ESCM despite rapid proliferation. f, Differentiation of AN3-12 cells into keratinocytes resulted in a near-complete loss of haploid cells among the keratin 14 (K14)-positive population; mES cells stained with anti-K14 are shown as a negative specificity control (grey curve in the K14 histogram). g, Immunostaining of AN3-12 cells cultured in ESCM as well as time course of removal of LIF with addition of 500 nM retinoic acid (analysed on the indicated days) shows downregulation of pluripotency markers Oct4, Nanog and Sox2. DAPI is shown as a nuclear counterstain. Scale bars, 50 μm. h, Histological examination of teratomas analysed 25 days after injection of 106 cells subcutaneously. All three germ layers were present in six analysed teratomas, representative H&E images are shown. Magnifications are indicated in each panel.
Extended Data Figure 2 Analysis of genome integrity.
a, M-FISH karyotypic analysis was performed on parental mouse haploid cells (AN3-12) to evaluate genomic stability. Randomly selected metaphases were karyotyped and examined by M-FISH and DAPI banding. Approximately, 200 metaphases from AN3-12 were counted for the diploid versus haploid frequency and 10 well-spread metaphases were fully karyotyped by M-FISH and DAPI-banding pattern. Images of normal female diploid and haploid karyotypes (19, X) are shown. Images were captured on a Zeiss AxioImager D1 fluorescent microscope equipped with narrow band-pass filters for DAPI, DEAC, FITC, CY3, Texas red and CY5. b, CNV analysis of haploid AN3-13 cells by genome sequencing using Illumina HiSeq2500. Mapped reads were analysed relative to male genomes of parental mouse strains C57BL/6J and 129/Sv, respectively, quotient to closer parental strain is shown. As expected, the X chromosome is overrepresented whereas the Y chromosome is absent. Regions of detected variation are highlighted with red boxes and are shown in c. Chromosome numbers are indicated. c, In AN3-12 haploid mES cells three very small deletions (on chromosomes 2, 10 and 12) and 1 duplication (on chromosome 13) were detectable as highlighted. d, Chromosomal distribution of SNP densities for in-house 129/Sv and AN3-12 mES cells relative to in-house C57BL/6 are shown. Numbers of SNPs were calculated for all non-overlapping 100-kb windows across the mm10 C57BL/6J mouse reference genome. SNP density in AN3-12 shows regions of high and low number of SNPs relative to the C57BL/6J genome, as expected for a haploid cell line derived from an F1 female between 129/Sv and C57BL/6.
Extended Data Figure 3 Molecular characterization of mutagenesis vectors.
a, Schematic illustration of the universal NGS strategy. Optimized primer-binding sites compatible with Illumina sequencing and two restriction enzymes with four base-pair recognition sites were placed adjacent to the terminal elements (LTR, TR). An internal barcode of 32 bases with alternating weak and strong bases was inserted in a parallel cloning step. b, For mapping of integration sites, genomic DNA was amplified by iPCR to introduce adaptor sequences and the experimental index for NGS. Paired-end sequencing maps the genomic integration in the first read using a custom primer, the experimental index as well as the internal barcode using standard Illumina primers binding to the integrated complementary sequence. Barcode (BC) PCR was performed on genomic DNA. c, Meta-analysis of mutagen integrations around transcriptional start sites (TSS) (excluding the precise TSS site). In particular Tol2 and retrovirus show a preference to integrate in proximity to the TSS. Retroviruses also frequently integrate into the promotor regions, whereas lentiviral integrations are typically located within the entire gene body. IPKM, insertions per kilobase per million. The vectors used are described in the legend of Fig. 1. d, Distribution of integration sites. Binning the number of integrations in genic and 2-kb upstream regions per 10-kb windows illustrates pronounced cold spots of mutagenesis using retroviral mutagenesis, where one can observe bins devoid of integrations. e, Genomic region surrounding the Gapdh locus exemplifying the distributions of integrations. While retroviral integrations strongly cluster, Tol2 displays a more uniform distribution of integration sites. Tracks are + strand (top) and − strand (bottom) integration sites. Bar lengths indicate NGS read numbers, subsequent to iPCR. f, Heat map illustrating overlap of epigenetic histone marks with integrations of the indicated mutagens, normalized to peak size. Only retrovirus and Tol2 integrations strongly correlate with DNA accessibility determined by ATAC-seq and active marks such as H3K4me3 and H3K27ac. In silico mutagenesis is shown as a control.
Extended Data Figure 4 Insertional preferences and generation of the mutant mES cell library.
a, Correlation between integration probabilities (IPKM, insertions per kilobase per million) and expression level (mean log2(FPKM)). Strongest correlation is seen for lentiviral constructs as well as Retro-GT without osteopontin-enhancer elements. All mutagenesis vectors are described in Fig. 1 and Methods. b, 5′ RACE on a set of pooled clones with confirmed antisense integration sites revealed multiple spurious transcription initiation sites in the intronic part of the gene-trap vector around the lox site, but we failed to detect spliced transcripts. Transcriptional initiation within the lox5171 site is highlighted. Red-labelled sequence is marking polyGs used for 5′ tailing. c, Intersection of integration sites of the indicated mutagenesis vectors (see Fig. 1) with genomic features. Coding sequences (CDS), 5′ and 3′ untranslated regions (5′ UTR and 3′ UTR), 1st intron, all other introns, excluding the first intron (intron), non-coding exons (ncExon), upstream regions (defined as 2-kb upstream of TSS) and intergenic regions are indicated. Mutagenesis by piggyback transposons as well as in silico random mutagenesis and ATAC-seq results are shown for comparison. d, Schematic work flow for generation of the mutant haploid mES cell library. Single-cell-derived clones were manually picked 10 or 11 days after seeding, expanded in 96-well plates, and either frozen in quadruplicates or further processed for mapping of the integration sites. e, Schematic illustration of the first step of 4D pooling. Each plate was pooled into the respective slice tray as well as a master plate, uniting identical well coordinates of all plates. f, Schematic illustration of the second step of 4D pooling. Each master-plate was pooled into a master tower pool, a plate with lamella uniting columns, and a plate with lamella uniting rows, thereby generating pools for rows and columns over all samples. g, 4-Dimensional pooling of 9,600 clones in 8 rows, 12 columns, 10 slices and 10 towers resulting in 40 pools. After iPCR to introduce experimental indices, pools were combined and deep sequenced. Amplification of internal barcodes confirmed clonal identity and mapping in 4 dimensions. All mapped clones were deposited in the Haplobank (https://www.haplobank.at/).
Extended Data Figure 5 Numbers of independent gene trap clones and intragenic distribution.
a, Numbers of independent available cell lines, carrying a single integration per cell, per gene. For about 37% (RefSeq) to 38% (Ensembl) of genes targeted, there is one gene-trap clone available (5′ UTR, intron or coding sequence), whereas about 18% of genes are targeted in two independent clones, and for around 43% of genes three or more independent clones are available. b, 24.8% (RefSeq) to 26.8% (Ensembl) of genes are represented by a single cell line if one takes all clones into account and about 40% of genes are hit in three or more clones. c, Separation of all gene traps combined into biotypes in single-integration clones of the Haplobank. Antisense and intergenic insertions are observed in all systems, in particular for enhanced gene-trap vectors. d, To map the integration sites of our clones from the Haplobank to the open reading frames (ORFs) of the respective genes dissected ORFs into 5% intervals and annotated integration sites in introns and exons relative to the position within the ORF. All mutagenesis systems (see Fig. 1) show a strong bias towards transcript truncation proximal to the 5′ end of the ORFs and are thus predicted to result in loss-of-function alleles. We defined integrations in the first 50% of the coding sequence (green bars) as optimal for a gene-trap allele; these clones are highlighted by a yellow star on the Haplobank homepage.
Extended Data Figure 6 Interaction of Pla2g16 with Cox inhibitors in mES cells.
a, Titration series of the indicated Cox inhibitors in the presence and absence of rhinovirus (RV-A1a) in mES cells. No protective effect of inhibition of prostaglandin biosynthesis was detected at non-toxic concentrations. Because mES cells do not generate infectious RV-A1a efficiently, conditioned supernatant containing RV-A1a was added daily. Data are shown as individual data points and mean values. b, mES cell clones from the Haplobank that had mutations in Ldlr and Pla2g16, respectively, were mixed as sister cells in sense (red) and antisense (green) orientation labelled by GFP and mCherry. Subsequently, cells were cultured in the presence and absence of rhinovirus RV-A1a for four days and ratios of red and green cells were then quantified using FACS. Selection pressure for loss of Ldlr and Pla2g16 was not affected by inhibition of Cox. Data are shown as mean ± s.d. of three biological replicates.
Extended Data Figure 7 Interactions of PLA2G16 with Cox inhibitors in human HEK293T cells and domain mapping.
a, RV-A1a exposure causes cell death in HEK293T cells in a dose-dependent manner. Cell viability was quantified three days after infection using Alamar blue. Data are shown as individual data points of five biological replicates and mean values. b, Titration series for ibuprofen and indomethacin treatment in the presence and absence of rhinovirus (RV-A1a) in HEK293T cells. Protective effects of ibuprofen and indomethacin were detected at a high drug concentration. Cell viability was quantified 2.5 days after infection using Alamar blue. Data are shown as individual data points of 4 biological replicates and mean values. c, Competitive growth assays in HEK293T cells. Cells containing sgRNAs targeting PLA2G16 did not show a growth difference in the absence of RV-A1a or when treated with indomethacin at 100 μM, but were significantly enriched when challenged with RV-A1a, indicating preferential survival. By contrast, control-guide-treated cells did not show growth advantages at any experimental condition. Data are shown as individual data points and mean ± s.d. of biological triplicates analysed on days 0, 3, 6 and 10 after RV-A1a exposure. d, Scheme of Pla2g16 domains. The enzymatic centre of Pla2g16 is located in the cytoplasm (green); an α helix in the transmembrane domain (yellow) connects the core to a short vesicular domain (blue), located in endosomes49. e, Design of CRISPR sgRNAs targeting the mRNA regions encoding the vesicular domain of seven amino acids (sgRNA1) and the 3′ UTR (sgRNA2) in haploid mES cells to test essentiality of these domains in RV-A1a infections. f, Cells carrying sgRNA2 showed editing in only 1 out of 12 cases, but upon selection with RV-A1a were enriched for deletions within the vesicular domain. For sgRNA2, all mapped deletions in control cells only affected the 3′ UTR, where the expected Cas9 cuts occur; upon RV-A1a exposure, the majority of observed deletions affected the transmembrane domain, the vesicular domain and, in some cases, even extended into the cytoplasmatic region. Colour codes: grey, deletion; red, alternative reading frame and insertions.
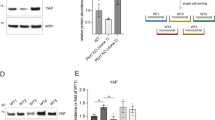
Extended Data Figure 8 Blood-vessel sprouting in Notch1 mutant mES cells and candidate tip cell genes.
a, Assessment of four independent Notch1-targeted clones from Haplobank. The locations of the integrations are shown: two antisense (AS) clones marked by green triangles, one sense (S) clone marked by the red triangle, and one clone with an upstream (AS-up) integration (blue triangle). Flipping of the gene traps upon Cre infection is shown by PCR in the left panel. Loss of Notch1 protein (intracellular domain, ICD) expression (clones A4, H7), and re-expression (clone D5) upon Cre recombination are shown by western blot (right panel). β-Actin is shown as a loading control. WT, parental clone without any gene trap integration. Uncropped blots are shown in Supplementary Fig. 1. b, Notch1 inactivation leads to a hypersprouting phenotype. Note the advanced progression and increased density of the vascular networks upon Notch1 deletion (sense clone) compared to antisense sister cells (top, bright field images; bottom, IB4 immunostaining to mark endothelial cells). Scale bars, 500 μm. c, Angiogenic sprouting is not affected when the gene trap is located 1,500 bp upstream of the Notch1 gene (A2 clone). GFP+ and Cre-reverted mCherry+ sister cells were analysed in 3D blood-vessel organoid cultures. Bright field images are shown. Scale bars, 500 μm. d, Differentially expressed genes in endothelial tip cells versus stalk cells from two published datasets30,31 in the mouse retina were filtered for genes that have also been associated (ingenuity pathway analysis) with candidate genes/pathways for vascular diseases in humans. Scatterplot showing the frequency of independent associations of tip cell genes with various human vascular diseases. Genes available in the Haplobank at the beginning of the project were chosen for functional analysis in the 3D organoids. For most of the listed candidate genes, there were no functional vascular data available. e, Quantification of IB4-positive vascular structures from the indicated sister clones carrying sense and repaired antisense integrations. Clones were classified according to their sprouting capacity from low (hyposprouting) to high (hypersprouting). f, Different clones with independent integrations in the same gene showed reproducible phenotypes in sprouting angiogenesis. Vascular outgrowths were stained for endothelial cell-specific IB4 expression, number of vessels counted and normalized to the respective antisense sister clones. e, f, Data are shown as individual data points from a minimum of n = 3 independent experiments for each sense/antisense sister clone combination together with the mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001; two-tailed Student’s t-test.
Extended Data Figure 9 Sprouting angiogenesis in reversible sister clones.
Representative images of the indicated sense (S) and antisense (AS) sister clones. IB4 was used to mark endothelial cells. GFP or mCherry expression indicates the respective flipped gene traps. Note that some sense clones are GFP+ whereas others are mCherry+; this is owing to the original orientation of the integration in sense or antisense, which was then reverted by the mCherry-Cre-expressing virus. Scale bar, 500 μm. For quantification of data see Fig. 3e.
Extended Data Figure 10 Generation of a chimaeric vasculature in vivo and gap junction α-1 protein localization to tip cells.
a, Representative fluorescence image of a haploid mES-cell-derived teratoma stained for endothelial cell-specific IB4. Endothelial cells arising from haploid mES cells are positive for mCherry and IB4 (yellow), whereas host endothelial cells are only positive for IB4 and appear green. Scale bars, 50 μm. b, Representative FACS analysis of teratomas following injection of chimaeric embryoid bodies into immunocompromised mice. Myst3 antisense (mCherry+) and sense (GFP+) sister clones were mixed at a 1:1 ratio. VE-cadherin-negative non-endothelial cells were also determined within the teratomas. c, Parental haploid mES cells stably expressing GFP or mCherry-Cre were assessed for their ability to generate IB4+ vascular structures in the presence of VEGFA. The number and ratios of IB4+ vessels per organoid were not apparently different between GFP-expressing and mCherry-Cre-expressing cells. Scale bars, 500 μm. Data are shown as mean ± s.e.m. and individual data points from n = 4 independent experiments. P = 0.207; two-tailed Student’s t-test. d, GFP-expressing and mCherry-Cre-expressing parental haploid mES cells contribute equally to tip cells (49.2% GFP+; 50.8% mCherry-Cre+) in 1:1 mixed mosaic cultures. Data are shown as mean ± s.e.m. and individual data points from of n = 4 independent experiments. P = 0.823; two-tailed Student’s t-test. e, Localization of gap junction α-1 protein in the mouse retina at postnatal day 6 (P6). Endothelial cells are marked by IB4 staining. At the angiogenic front, gap junction α-1 protein expression is found in endothelial cells, primarily localized at tip cells (arrows). Scale bars, 50 μm. f, Retinas were stained for gap junction α-1 protein expression and the endothelial marker IB4 to visualize the vascular networks on P6. Note the punctate pattern of gap junction α-1 protein adjacent to the IB4+ vessels, suggestive of gap junction α-1 protein expression in perivascular cells. Scale bar, 50 μm (left) and 20 μm (right). g, Gap junction α-1 protein predominantly localizes to the tip cells (arrows) in the 3D blood vessels. Vessels are marked by CD31 immunostaining and counterstained by DAPI. Bar graph indicates percentages of vessels with the highest gap junction α-1 protein expression in the tip cell. Data are shown as individual data points of eight independent embryoid bodies and mean ± s.d. of vessels. Scale bars, 20 μm (right) and 10 μm (left).
Supplementary information
Supplementary Figures
This file contains full uncropped scans of DNA gels and Western blots used in Extended Data Figure 8. (PDF 2098 kb)
Supplementary Table
This table shows the numbers of clones available with respect to different mutagens, orientation of the inserted gene trap to gene transcription, as well as the number of different genes hit. A gene is defined as the genomic region between the transcriptional start and stop sites. www.haplobank.at (PDF 490 kb)
Rights and permissions
About this article
Cite this article
Elling, U., Wimmer, R., Leibbrandt, A. et al. A reversible haploid mouse embryonic stem cell biobank resource for functional genomics. Nature 550, 114–118 (2017). https://doi.org/10.1038/nature24027
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24027
This article is cited by
-
Crimean–Congo haemorrhagic fever virus uses LDLR to bind and enter host cells
Nature Microbiology (2024)
-
Loss of cohesin regulator PDS5A reveals repressive role of Polycomb loops
Nature Communications (2023)
-
Identification of CCZ1 as an essential lysosomal trafficking regulator in Marburg and Ebola virus infections
Nature Communications (2023)
-
SCON—a Short Conditional intrON for conditional knockout with one-step zygote injection
Experimental & Molecular Medicine (2022)
-
Organelle degradation in the lens by PLAAT phospholipases
Nature (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.