Abstract
The majority of genetic variants associated with common human diseases map to enhancers, non-coding elements that shape cell-type-specific transcriptional programs and responses to extracellular cues1,2,3. Systematic mapping of functional enhancers and their biological contexts is required to understand the mechanisms by which variation in non-coding genetic sequences contributes to disease. Functional enhancers can be mapped by genomic sequence disruption4,5,6, but this approach is limited to the subset of enhancers that are necessary in the particular cellular context being studied. We hypothesized that recruitment of a strong transcriptional activator to an enhancer would be sufficient to drive target gene expression, even if that enhancer was not currently active in the assayed cells. Here we describe a discovery platform that can identify stimulus-responsive enhancers for a target gene independent of stimulus exposure. We used tiled CRISPR activation (CRISPRa)7 to synthetically recruit a transcriptional activator to sites across large genomic regions (more than 100 kilobases) surrounding two key autoimmunity risk loci, CD69 and IL2RA. We identified several CRISPRa-responsive elements with chromatin features of stimulus-responsive enhancers, including an IL2RA enhancer that harbours an autoimmunity risk variant. Using engineered mouse models, we found that sequence perturbation of the disease-associated Il2ra enhancer did not entirely block Il2ra expression, but rather delayed the timing of gene activation in response to specific extracellular signals. Enhancer deletion skewed polarization of naive T cells towards a pro-inflammatory T helper (TH17) cell state and away from a regulatory T cell state. This integrated approach identifies functional enhancers and reveals how non-coding variation associated with human immune dysfunction alters context-specific gene programs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
13 June 2018
In this Letter, the data used to generate Extended Data Fig. 9d contained errors and were not correctly analysed. The corrected and reanalysed Extended Data Fig. 9d is shown in the Supplementary Information to the accompanying Amendment. The original Letter has not been corrected.
References
Farh, K. K.-H. et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature 518, 337–343 (2015)
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012)
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011)
Canver, M. C. et al. BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis. Nature 527, 192–197 (2015)
Korkmaz, G. et al. Functional genetic screens for enhancer elements in the human genome using CRISPR–Cas9. Nat. Biotechnol. 34, 192–198 (2016)
Rajagopal, N. et al. High-throughput mapping of regulatory DNA. Nat. Biotechnol. 34, 167–174 (2016)
Komor, A. C., Badran, A. H. & Liu, D. R. CRISPR-based technologies for the manipulation of eukaryotic genomes. Cell 168, 20–36 (2017)
Laguna, T. et al. New insights on the transcriptional regulation of CD69 gene through a potent enhancer located in the conserved non-coding sequence 2. Mol. Immunol. 66, 171–179 (2015)
Ziegler, S. F., Ramsdell, F. & Alderson, M. R. The activation antigen CD69. Stem Cells 12, 456–465 (1994)
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014)
Leonard, W. J., Krönke, M., Peffer, N. J., Depper, J. M. & Greene, W. C. Interleukin 2 receptor gene expression in normal human T lymphocytes. Proc. Natl Acad. Sci. USA 82, 6281–6285 (1985)
Kim, H. P., Imbert, J. & Leonard, W. J. Both integrated and differential regulation of components of the IL-2/IL-2 receptor system. Cytokine Growild-typeh Factor Rev. 17, 349–366 (2006)
Fontenot, J. D., Rasmussen, J. P., Gavin, M. A. & Rudensky, A. Y. A function for interleukin 2 in Foxp3-expressing regulatory T cells. Nat. Immunol. 6, 1142–1151 (2005)
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013)
Hnisz, D. et al. Convergence of developmental and oncogenic signaling pathways at transcriptional super-enhancers. Mol. Cell 58, 362–370 (2015)
Goudy, K. et al. Human IL-2Ra null mutation mediates immunodeficiency with lymphoproliferation and autoimmunity. Clin. Immunol. 146, 248–261 (2013)
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016)
Huang, H. et al. Fine-mapping inflammatory bowel disease loci to single-variant resolution. Nature 547, 173–178 (2017)
Huang, J., Ellinghaus, D., Franke, A., Howie, B. & Li, Y. 1000 Genomes-based imputation identifies novel and refined associations for the Wellcome Trust Case Control Consortium phase 1 Data. Eur. J. Hum. Genet. 20, 801–805 (2012)
Onengut-Gumuscu, S. et al. Fine mapping of type 1 diabetes susceptibility loci and evidence for colocalization of causal variants with lymphoid gene enhancers. Nat. Genet. 47, 381–386 (2015)
Ye, C. J. et al. Intersection of population variation and autoimmunity genetics in human T cell activation. Science 345, 1254665 (2014)
Laurence, A. et al. Interleukin-2 signaling via STAT5 constrains T helper 17 cell generation. Immunity 26, 371–381 (2007)
Fujino, S. et al. Increased expression of interleukin 17 in inflammatory bowel disease. Gut 52, 65–70 (2003)
Furtado, G. C., Curotto de Lafaille, M. A., Kutchukhidze, N. & Lafaille, J. J . Interleukin 2 signaling is required for CD4+ regulatory T cell function. J. Exp. Med. 196, 851–857 (2002)
Chatenoud, L. & Bluestone, J. A. CD3-specific antibodies: a portal to the treatment of autoimmunity. Nat. Rev. Immunol. 7, 622–632 (2007)
Kuhn, C. & Weiner, H. L. Therapeutic anti-CD3 monoclonal antibodies: from bench to bedside. Immunotherapy 8, 889–906 (2016)
Klatzmann, D. & Abbas, A. K. The promise of low-dose interleukin-2 therapy for autoimmune and inflammatory diseases. Nat. Rev. Immunol. 15, 283–294 (2015)
Horlbeck, M. A. et al. Nucleosomes impede Cas9 access to DNA in vivo and in vitro. eLife 5, 2767 (2016)
Kampmann, M., Bassik, M. C. & Weissman, J. S. Functional genomics platform for pooled screening and generation of mammalian genetic interaction maps. Nat. Protocols 9, 1825–1847 (2014)
Marcel, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011)
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012)
Sinha, R. et al. Index switching causes ‘spreading-of-signal’ among multiplexed samples in Illumina HiSeq 4000 DNA Sequencing. Preprint at https://doi.org/10.1101/125724 (2017)
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Gen. Biol. 16, 259–270 (2015)
Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015)
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016)
Pimentel, H. J., Bray, N., Puente, S., Melsted, P. & Pachter, L. Differential analysis of RNA-Seq incorporating quantification uncertainty. Preprint at https://doi.org/10.1101/058164 (2016)
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015)
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010)
Acknowledgements
We thank members of Marson and Corn laboratories, as well as A. Abbas, S. Qi, L. Gilbert, J. Hiatt, M. Lee, V. Nguyen, J. Weissman, J. Roose, M. Gavin and W. Leonard for suggestions and technical assistance. This research was supported by NIH grants DP3DK111914-01 (A.M., M.S.A., C.Y.), R01HG0081410-01 (A.M, W.G.), R01HL109102 (K.M.A., A.M.), P50-HG007735 (H.Y.C., W.J.G.), Scleroderma Research Foundation (H.Y.C.), the UCSF Sandler Fellowship (A.M.), a gift from Jake Aronov (A.M.), a National Multiple Sclerosis Society grant (A.M.; CA 1074-A-21), and the Marcus Program in Precision Medicine Innovation (A.M.). A.M. holds a Career Award for Medical Scientists from the Burroughs Wellcome Fund and is a Chan Zuckerberg Biohub Investigator. J.E.C. is supported by the Li Ka Shing Foundation. B.G.G. is supported by the IGI-AstraZeneca Postdoctoral Fellowship. K.M.A. is a Leukemia & Lymphoma Society Scholar. J.D.G. is a National Science Foundation Predoctoral Fellow. K.S. is supported by a DFG Postdoctoral Fellowship. We thank Jackson Laboratories for generating the SNP and EDEL mice and Agilent for generating oligo pools for cloning of the CRISPRa gRNA library. We thank UC Berkeley High Throughput Screening Facility and Flow Cytometry Facility. This work used the Vincent J. Coates Genomics Sequencing Laboratory at UC Berkeley, supported by NIH S10 Instrumentation Grants S10RR029668 and S10RR027303. We also relied on the Flow Cytometry Core at UCSF, supported by the Diabetes Research Center grant NIH P30 DK063720.
Author information
Authors and Affiliations
Contributions
D.R.S., B.G.G, J.E.C. and A.M. designed the study and wrote the manuscript. B.G.G., M.B., N.L.B., T.M., G.J.R. and G.L.C. performed and analysed CRISPRa screens. B.G.G., D.R.S., N.N., J.S.C. and H.M. performed luciferase reporter cloning and experiments. D.R.S., T.L.R., J.D.G., Y.L., A.C., M.L.N., Z.L., J.M.W, E.B., F.V.G, V.R.T, R.E.G., M.S. and K.S. contributed to functional experiments on CaRE4 and rs61839660. M.R.M., A.T.S., W.J.G. and H.Y.C. generated and analysed HiChIP data. D.S.L. and C.Y. performed ImmVar QTL analysis. H.H., R.L., K.K.F. and M.J.D. contributed to fine-mapping analysis of rs61839660 disease association. D.R.S, J.D.G. and K.M.A. contributed to T cell differentiation. M.S. and J.A.B. advised on functional studies in murine models. D.R.S. and B.G.G. are joint first authors. J.E.C. and A.M. are co-corresponding and co-senior authors.
Corresponding authors
Ethics declarations
Competing interests
H.Y.C. and W.J.G. are co-founders of Epinomics. A.M. and J.E.C. are co-founders of Spotlight Therapeutics. J.E.C. serves as an advisor to Mission Therapeutics and the Corn laboratory has received sponsored research support from AstraZeneca and Pfizer. A.M. serves as an advisor to Juno Therapeutics and PACT Therapeutics and the Marson laboratory has received sponsored research support from Juno Therapeutics and Epinomics.
Additional information
Reviewer Information Nature thanks F. Urnov and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Upregulation of target gene expression on gRNA-expressing cells.
a, Distribution of CD69 expression on Jurkat-dCas9-VP64 cells transduced with the CD69 tiling gRNA library. b, c, Representative flow cytometry plots of CD69 expression on Jurkat (b) or HuT78 cells (c) transduced with dCas9-VP64 and individual gRNAs. For each target region or control, solid black lines represent gRNA 1 and dashed black lines represent gRNA 2. Shaded grey histograms represent isotype control staining. Cells stimulated for 48 h with plate-bound anti-CD3/CD28 antibodies are shown for comparison. d, e, Isotype-subtracted geometric MFI of data in b and c. f, g, Representative flow cytometry plots of IL-2Ra expression on Jurkat (f) and HuT78 cells (g) as in d. h, i, Isotype-subtracted geometric MFI of data in f and g. Statistical tests were performed on log-transformed MFI values. PBS and anti-CD3/CD28-treated samples were compared using an unpaired two-tailed Student’s t-test. TSS and CaRE gRNA samples were compared to each non-targeting (NT) gRNA sample using one-way ANOVA followed by Sidak’s multiple comparisons test. Data are presented as mean ± s.d., n = 3 biological replicates. Data in b–i are representative of at least 2 independent experiments. **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001. j, Jurkat dCas9-VP64 cells were transduced with individual gRNAs from the IL-2Ra library, and surface IL-2Ra expression was measured by flow cytometry. The isotype-subtracted geometric MFI of the transduced cells is plotted against gRNA enrichment in the indicated bin in the IL-2Ra screen.
Extended Data Figure 2 Correlation of results across CRISPRa screen replicates.
a, b, Normalized read counts for gRNAs in the indicated cell populations are compared between biological replicates of the CD69 screen (a) and IL-2Ra screen (b).
Extended Data Figure 3 Activation of CaREs by CRISPRa specifically upregulates IL2RA.
a, Transcriptome comparison of HuT78 cells expressing dCas9-VP64 transduced with individual gRNAs targeting the IL2RA TSS, CaRE3 or CaRE4 versus a non-targeting sgRNA. Cells stimulated for 48 h with plate-bound CD3 and CD28 antibodies were also analysed. Scatter plots show gene-level abundance estimates averaged over two replicates for each condition. Genes called as differentially expressed for each targeting guide, as described in Methods, are highlighted in red in their respective plot. For visualization purposes, transcripts per million values have been scaled by the transformation x→x1/10. b, RNA-seq read coverage for IL2RA non-targeting, TSS, CaRE3 and CaRE4 gRNA samples.
Extended Data Figure 4 Chromatin features and enhancer activity of CD69 CaREs.
a, Results of the CD69 CRISPRa screen are overlapped with DNase I hypersensitivity, H3K27ac, and H3K4me1 datasets from various primary human haematopoietic cell types. Data are shown for the indicated reference epigenomes from the Roadmap Epigenomics Project. Jurkat DNase HS data are from ENCODE. b, Jurkat cells were nucleofected with luciferase reporter constructs containing sequences from CD69 CaREs upstream of a generic minimal promoter. 18 h after nucleofection, cells were split between a stimulation plate coated with anti-CD3/CD28 antibodies or a PBS control plate. Cells were lysed after 24 h of stimulation, followed by measurement of luciferase activity. Data are presented as mean ± s.d., n = 4 biological replicates. Data are representative of two independent experiments. The dotted line represents the threshold of relevant luciferase activity defined as two times the value from a sequence-scrambled IL-2Ra CaRE4 control construct. ****P ≤ 0.0001 by one-way ANOVA followed by Dunnet’s multiple comparisons test.
Extended Data Figure 5 Chromatin features and enhancer activity of IL2RA CaREs.
a, Results of the IL2RA CRISPRa screen are overlapped with with DNase I hypersensitivity, H3K27ac, and H3K4me1 datasets from primary human haematopoietic cell types. Data from the Roadmap Epigenomics Project. Jurkat DNase HS data are from ENCODE. b, Jurkat cells were nucleofected with luciferase reporter constructs containing IL2RA CaRE sequences upstream of a generic minimal promoter. 18 h after nucleofection, cells were split to a stimulation plate coated with anti-CD3/CD28 or a PBS control plate. Cells were lysed 24 h later and luciferase activity was measured. Data are presented as mean ± s.d., n = 4 biological replicates, representative of two independent experiments. Dotted line represents relevant luciferase activity defined as two times activity of sequence-scrambled IL2RA CaRE4 control. ****P ≤ 0.0001 by one-way ANOVA followed by Dunnet’s multiple comparisons test.
Extended Data Figure 6 IL2RA CaRE4 harbours a risk variant that is linked to Crohn’s disease and reduced IL2RA expression in stimulated CD4+ T cells.
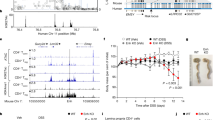
a, HiChIP looping data anchored at IL2RA CaRE4 reveals that in addition to interacting with the IL2RA promoter, CaRE4 physically associates with other sites in the IL2RA locus as well as the promoters of IL15RA and RBM17. b, IL2RA regional association plot. P values of variants associated to Crohn’s disease were taken from the inflammatory bowel diseases fine-mapping study18, including all SNPs and indels in the 1000 Genomes phase 1 project. New SNPs and INDELs from the 1000 Genomes phase 3 and the UK10K projects were not included in this figure, but none of these has high LD with rs61839660 that could explain the SNP association. Genes within 150 kbp of IL2RA (from UCSC Genome Browser human GRCh37 assembly) were plotted. Figure generated using Locuszoom (http://locuszoom.org). c, Reduced IL2RA levels in stimulated primary human T cells with the natural rs61839660 variant. The minor ‘T’ allele of rs61839660 is associated with reduced IL2RA levels in stimulated primary human T cells without conditioning. d, rs61839660 is the mostly highly associated SNP with IL2RA levels in stimulated primary human T cells at 48 h after conditioning on rs2476491.
Extended Data Figure 7 Il2ra enhancer-edited mice show no steady-state immune dysfunction.
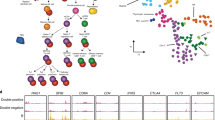
Enhancer-edited mice and littermate controls were immunophenotyped at 2–4 months of age. a, Spleens from wild-type, SNP and 12DEL mice. b, Total number of cells in spleen, peripheral lymph nodes (peri-LNs) and thymus. c, Percentage of naive (CD4+CD62L+CD44−) and memory (CD4+CD62L−CD44+) CD4+ T cells isolated from spleen and peri-LNs. d, Percentage of thymocytes in T cell developmental stages from the thymus. Data shown for CD4/CD8 single-positive, double-positive, and double-negative populations. e, f, Quantification of percentage IL-2Ra+ double-negative thymocytes and IL-2Ra MFI on IL-2Ra+ double-negative thymocytes. g, Percentage of Treg cells of CD4+ T cells in tissues of enhancer edited mice and littermate controls at 2–4 months of age. h, Quantification of IL-2Ra surface staining (geometric MFI) on FOXP3+ cells. All data are presented as mean ± s.d. and are representative of at least two independent experiments. Data are biological replicates of wild-type (n = 7), SNP (n = 4), and 12DEL (n = 5) mice (a–f) and of wild-type (n = 6), SNP (n = 4), and 12DEL (n = 5) mice (g, h). A non-parametric one-way ANOVA (*P ≤ 0.05) followed by Dunn’s multiple comparison test was used to compare enhancer-edited mice to wild-type controls.
Extended Data Figure 8 IL-2Ra induction in stimulated SNP and 12DEL T cells.
a, Wild-type, SNP and 12DEL cells that were stimulated only (‘stim’ (anti-CD3/CD28)) or stim + 50 U ml−1 IL-2 over 3 days. b, Percentage of CD69+ cells by surface levels on wild-type and enhancer edited cells 1 day after stimulation. c, Statistical analysis using Fisher’s LSD at each day of stimulation time course comparing wild-type and SNP naive T cells, with or without IL-2. d, IL-2Ra MFI on IL-2Ra+ T cells with stim or stim + 50 U ml−1 IL-2 over 3 days. Table shows the Fisher’s LSD statistical analysis at each day of T cell stimulation time course. e, IL-2Ra MFI on 12DEL naive T cells with stim, with stim + IL-2 or 10 μg ml−1 anti-IL-2 blocking antibody. Data displayed in c, d and e are representative of two independent experiments. Data in d are from wild-type (n = 3) and SNP (n = 3) gender matched littermate controls. All data are normalized to IL-2Ra MFI on wild-type stim only cells at day 3. A two-way ANOVA with multiple comparisons testing followed by Fisher’s LSD test was used for statistical analysis. Data in e are from wild-type (n = 2) and 12DEL (n = 2) littermate controls. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001.
Extended Data Figure 9 Characterization of Il2ra enhancer deletion (EDEL) on the NOD background.
a, Representative spleens from wild-type and EDEL mice. b, Naive (CD62L+CD44−) and memory (CD44+ CD62L−) compositions of CD4+ T cells in wild-type and EDEL. c, Representative lymph node staining showing Treg (CD4+FOXP3+) and Teff (CD4+FOXP3−) compartments. d, Quantification of Treg cell abundance across multiple different tissues. e, As we did not uncover defects in steady state T cells, we isolated naive T cells and activated them in vitro with anti-CD3/CD28 antibodies. qPCR on naive T cells from wild-type or EDEL mice 8 h after stimulation. Relative transcript levels for Il2ra, CD69 (control), Il15ra (adjacent gene), and Rbm17 (adjacent gene) are shown. The average Ct value for each transcript on wild-type cells is shown. f, CD69 protein surface expression on wild-type and EDEL naive T cells 1 day after stimulation with anti-CD3/CD28. g, Representative flow plot of 3 day time course with naive T cells stim only (anti-CD3/CD28), stim + 50 U ml−1 IL-2 or stim + 10 μg ml−1 anti-IL-2. (h) Quantification of percentage of IL-2Ra− cells in the time course. i, Quantification of IL-2Ra MFI on IL-2Ra+ cells. Data were generated from two independent experiments with wild-type (n = 6) and EDEL (n = 6) mice. EDEL and wild-type mice were treated with 50 μg anti-CD3 to assess the in vivo T cell response to stimulation. Mice were killed one day after treatment and IL-2Ra surface levels were checked by flow cytometry on T cells from spleen, peripheral lymph nodes (pLN), mesenteric lymph nodes (mLN) and colon. j, Representative IL-2Ra MFI histograms on CD4+ and CD8+ T cells from various tissues. k–m, Quantification of IL-2Ra MFI on CD4+FOXP3− Teff, CD4+FOXP3+ Treg and CD8+ T cells from different tissues. n, Abundance of regulatory T cells in tissues following acute stimulation with anti-CD3 antibody. Data are representative of two experiments. EDEL (n = 8) and wild-type (n = 3) littermate mice were used for experiments. A two-way ANOVA with Holm–Sidak multiple comparisons test was used for statistical analysis. *P ≤ 0.05, **P ≤ 0.01, ****P ≤ 0.0001.
Extended Data Figure 10 Il2ra enhancer deletion promotes TH17 and inhibits iTreg CD4+ T cell differentiation in IL-2-limiting conditions.
Naive CD4+ T cells were activated with anti-CD3/anti-CD28 and differentiated in the presence of TGFβ, anti-IL-4, anti-IFNγ with high (20 ng ml−1), medium (2 ng ml−1) or no IL-6. The IL-2 activity was varied within each IL-6 concentration by adding IL-2-blocking antibody (10 ng ml−1, 1 ng ml−1 or 0.01 ng ml−1), no IL-2 or 50 U ml−1 IL-2. a–e, Five days after initial activation flow cytometry was used to assess IL-17A for TH17 differentiation (a), FOXP3 for iTreg differentiation (b), viability (c), and IL-2Ra induction (d, e). Experiments were carried out with wild-type (n = 3) and EDEL (n = 3) age matched and sex matched littermate controls. A two-way ANOVA with Holm–Sidak multiple comparisons test was used for statistical analysis. **P ≤ 0.01, ***P ≤ 0.001.
Supplementary information
Supplementary Figure
Flow cytometry gating strategies. (a) Gating strategy for mouse immunophenotyping of spleen. Data for peripheral and mesenteric lymph nodes, as well as large intestine were acquired and analyzed in a similar manner. (b) Thymus flow cytometry data. (c) Sort sort strategy for naive CD4+ T cells from spleen and peripheral lymph node negatively enriched for CD4+ T cells. (d) Example of naive CD4+ T cell activation staining. (e) FACS from anti-CD3 treated mice. (f) NOD EDEL naive CD4+ T cell differentiation flow cytometry gating. (PDF 1215 kb)
Supplementary Table 1
CRISPRa screen raw data and analysis. This table contains the following information for each sgRNA in the CD69 and IL2RA libraries: raw read counts for each replicate, normalized read counts for each replicate, log2 fold-change over background for each replicate, and the mean log2 fold-change across screen replicates. (XLSX 13536 kb)
Supplementary Table 2
Differentially expressed genes from CRISPRa of IL2RA TSS, CaRE3 and CaRE4. This table lists the genes called as differentially expressed, along with q-value and log2 fold-change. (TXT 1 kb)
Supplementary Table 3
Full RNA sequencing data from CRISPRa of IL2RA TSS, CaRE3 and CaRE4. This table contains the q-value and log2 fold-change for all genes analyzed in the RNA-Seq experiment. (TXT 3105 kb)
Supplementary Table 4
A list of antibodies used in the study. (XLSX 11 kb)
Supplementary Table 5
A list of primer sequences used in the study. (XLSX 49 kb)
Supplementary Table 6
A guide to RNA sequences used in the study. (XLSX 42 kb)
Supplementary Table 7
HiChIP interaction matrices for IL2RA TSS and CaREs. (XLSX 629 kb)
Supplementary Table 8
Constructs for luciferase assays. This table contains details about each luciferase reporter construct used. (XLSX 43 kb)
Supplementary Table 9
Analysis matrix for CRISPRa RNA sequencing in Extended Data Figure 3. This table contains the analysis matrix used by sleuth to determine differential gene expression. (TXT 0 kb)
Supplementary Table 10
Epigenetic track documentation. This table contains information on each track of epigenetic data shown in Extended Data Figures 4 and 5. (XLSX 43 kb)
Rights and permissions
About this article
Cite this article
Simeonov, D., Gowen, B., Boontanrart, M. et al. Discovery of stimulation-responsive immune enhancers with CRISPR activation. Nature 549, 111–115 (2017). https://doi.org/10.1038/nature23875
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23875
This article is cited by
-
Mapping the functional impact of non-coding regulatory elements in primary T cells through single-cell CRISPR screens
Genome Biology (2024)
-
CRISPR technologies for genome, epigenome and transcriptome editing
Nature Reviews Molecular Cell Biology (2024)
-
The regulation and differentiation of regulatory T cells and their dysfunction in autoimmune diseases
Nature Reviews Immunology (2024)
-
Multicenter integrated analysis of noncoding CRISPRi screens
Nature Methods (2024)
-
GCLiPP: global crosslinking and protein purification method for constructing high-resolution occupancy maps for RNA binding proteins
Genome Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.