Abstract
Regulatory T cells (Treg cells) have a pivotal role in the establishment and maintenance of immunological self-tolerance and homeostasis1,2. Transcriptional programming of regulatory mechanisms facilitates the functional activation of Treg cells in the prevention of diverse types of inflammatory responses3,4. It remains unclear how Treg cells orchestrate their homeostasis and interplay with environmental signals. Here we show that liver kinase B1 (LKB1) programs the metabolic and functional fitness of Treg cells in the control of immune tolerance and homeostasis. Mice with a Treg-specific deletion of LKB1 developed a fatal inflammatory disease characterized by excessive TH2-type-dominant responses. LKB1 deficiency disrupted Treg cell survival and mitochondrial fitness and metabolism, but also induced aberrant expression of immune regulatory molecules including the negative co-receptor PD-1 and the TNF receptor superfamily proteins GITR and OX40. Unexpectedly, LKB1 function in Treg cells was independent of conventional AMPK signalling or the mTORC1–HIF-1α axis, but contributed to the activation of β-catenin signalling for the control of PD-1 and TNF receptor proteins. Blockade of PD-1 activity reinvigorated the ability of LKB1-deficient Treg cells to suppress TH2 responses and the interplay with dendritic cells primed by thymic stromal lymphopoietin. Thus, Treg cells use LKB1 signalling to coordinate their metabolic and immunological homeostasis and to prevent apoptotic and functional exhaustion, thereby orchestrating the balance between immunity and tolerance.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Liston, A. & Gray, D. H. Homeostatic control of regulatory T cell diversity. Nat. Rev. Immunol. 14, 154–165 (2014)
Sakaguchi, S., Yamaguchi, T., Nomura, T. & Ono, M. Regulatory T cells and immune tolerance. Cell 133, 775–787 (2008)
van der Veeken, J., Arvey, A. & Rudensky, A. Transcriptional control of regulatory T-cell differentiation. Cold Spring Harb. Symp. Quant. Biol. 78, 215–222 (2013)
Panduro, M., Benoist, C. & Mathis, D. Tissue Tregs. Annu. Rev. Immunol. 34, 609–633 (2016)
Shackelford, D. B. & Shaw, R. J. The LKB1–AMPK pathway: metabolism and growth control in tumour suppression. Nat. Rev. Cancer 9, 563–575 (2009)
Rubtsov, Y. P. et al. Regulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity 28, 546–558 (2008)
Reber, L. L., Sibilano, R., Mukai, K. & Galli, S. J. Potential effector and immunoregulatory functions of mast cells in mucosal immunity. Mucosal Immunol. 8, 444–463 (2015)
Shimizu, J., Yamazaki, S., Takahashi, T., Ishida, Y. & Sakaguchi, S. Stimulation of CD25+CD4+ regulatory T cells through GITR breaks immunological self-tolerance. Nat. Immunol. 3, 135–142 (2002)
Vu, M. D. et al. OX40 costimulation turns off Foxp3+ Tregs. Blood 110, 2501–2510 (2007)
Sage, P. T., Francisco, L. M., Carman, C. V. & Sharpe, A. H. The receptor PD-1 controls follicular regulatory T cells in the lymph nodes and blood. Nat. Immunol. 14, 152–161 (2013)
Mahmud, S. A. et al. Costimulation via the tumor-necrosis factor receptor superfamily couples TCR signal strength to the thymic differentiation of regulatory T cells. Nat. Immunol. 15, 473–481 (2014)
Valzasina, B. et al. Triggering of OX40 (CD134) on CD4+CD25+ T cells blocks their inhibitory activity: a novel regulatory role for OX40 and its comparison with GITR. Blood 105, 2845–2851 (2005)
Franceschini, D. et al. PD-L1 negatively regulates CD4+CD25+Foxp3+ Tregs by limiting STAT-5 phosphorylation in patients chronically infected with HCV. J. Clin. Invest. 119, 551–564 (2009)
Rubtsov, Y. P. et al. Stability of the regulatory T cell lineage in vivo. Science 329, 1667–1671 (2010)
Levine, A. G., Arvey, A., Jin, W. & Rudensky, A. Y. Continuous requirement for the TCR in regulatory T cell function. Nat. Immunol. 15, 1070–1078 (2014)
Vahl, J. C. et al. Continuous T cell receptor signals maintain a functional regulatory T cell pool. Immunity 41, 722–736 (2014)
MacIver, N. J. et al. The liver kinase B1 is a central regulator of T cell development, activation, and metabolism. J. Immunol. 187, 4187–4198 (2011)
Faubert, B. et al. Loss of the tumor suppressor LKB1 promotes metabolic reprogramming of cancer cells via HIF-1α. Proc. Natl Acad. Sci. USA 111, 2554–2559 (2014)
Tamaka, K., Seike, M., Hagiwara, T., Sato, A. & Ohtsu, H. Histamine suppresses regulatory T cells mediated by TGF-β in murine chronic allergic contact dermatitis. Exp. Dermatol. 24, 280–284 (2015)
Gao, Y. et al. Control of T helper 2 responses by transcription factor IRF4-dependent dendritic cells. Immunity 39, 722–732 (2013)
Zhou, B. et al. Thymic stromal lymphopoietin as a key initiator of allergic airway inflammation in mice. Nat. Immunol. 6, 1047–1053 (2005)
Okazaki, T., Chikuma, S., Iwai, Y., Fagarasan, S. & Honjo, T. A rheostat for immune responses: the unique properties of PD-1 and their advantages for clinical application. Nat. Immunol. 14, 1212–1218 (2013)
Chae, W. J. et al. The Wnt antagonist Dickkopf-1 promotes pathological type 2 cell-mediated inflammation. Immunity 44, 246–258 (2016)
Ding, Y., Shen, S., Lino, A. C., Curotto de Lafaille, M. A. & Lafaille, J. J. β-catenin stabilization extends regulatory T cell survival and induces anergy in nonregulatory T cells. Nat. Med. 14, 162–169 (2008)
Ulges, A. et al. Protein kinase CK2 enables regulatory T cells to suppress excessive TH2 responses in vivo. Nat. Immunol. 16, 267–275 (2015)
Pearce, E. L. & Pearce, E. J. Metabolic pathways in immune cell activation and quiescence. Immunity 38, 633–643 (2013)
Wherry, E. J. et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity 27, 670–684 (2007)
Patsoukis, N. et al. PD-1 alters T-cell metabolic reprogramming by inhibiting glycolysis and promoting lipolysis and fatty acid oxidation. Nat. Commun. 6, 6692 (2015)
Bengsch, B. et al. Bioenergetic insufficiencies due to metabolic alterations regulated by the inhibitory receptor PD-1 are an early driver of CD8+ T cell exhaustion. Immunity 45, 358–373 (2016)
Chang, C. H. et al. Metabolic competition in the tumor microenvironment is a driver of cancer progression. Cell 162, 1229–1241 (2015)
Ito, T. et al. TSLP-activated dendritic cells induce an inflammatory T helper type 2 cell response through OX40 ligand. J. Exp. Med. 202, 1213–1223 (2005)
Rosenbluh, J. et al. β-catenin-driven cancers require a YAP1 transcriptional complex for survival and tumorigenesis. Cell 151, 1457–1473 (2012)
Liu, G., Yang, K., Burns, S., Shrestha, S. & Chi, H. The S1P1–mTOR axis directs the reciprocal differentiation of TH1 and Treg cells. Nat. Immunol. 11, 1047–1056 (2010)
Zeng, H. et al. mTORC1 couples immune signals and metabolic programming to establish Treg-cell function. Nature 499, 485–490 (2013)
Williams, A. L. et al. Sequence variants in SLC16A11 are a common risk factor for type 2 diabetes in Mexico. Nature 506, 97–101 (2014)
Shrestha, S. et al. Treg cells require the phosphatase PTEN to restrain TH1 and TFH cell responses. Nat. Immunol. 16, 178–187 (2015)
Schuijers, J., Mokry, M., Hatzis, P., Cuppen, E. & Clevers, H. Wnt-induced transcriptional activation is exclusively mediated by TCF/LEF. EMBO J. 33, 146–156 (2014)
Zheng, Y. ChIP-on-chip for FoxP3. Methods Mol. Biol. 707, 71–82 (2011)
Acknowledgements
The authors acknowledge A. Rudensky for Foxp3YFP-cre and Foxp3cre-ERT2 mice, Y. Wang for editing of the manuscript, Y. Dhungana and Y. Li for advice on bioinformatics analyses, and St Jude Immunology FACS core facility for cell sorting. This work was supported by NIH AI105887, AI101407, CA176624 and NS064599, and American Asthma Foundation (to H.C.).
Author information
Authors and Affiliations
Contributions
K.Y. designed and performed experiments, and wrote the manuscript; B.D. and S.S. contributed to cellular experiments; G.N. performed microarray analysis; P.V. performed histological analysis; J.A., C.C. and C.W. contributed to metabolomics analysis; S.R., L.L. and A.K. contributed to molecular experiments; H.C. designed experiments, contributed to writing the manuscript, and provided overall direction.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks L. Turka and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Disrupted immune homeostasis in Foxp3creStk11fl/fl mice.
a, Expression of Stk11 mRNA (WT, n = 3; Foxp3creStk11fl/fl, n = 4) and LKB1 protein in CD4+YFP+ T cells (Treg cells) from wild-type and Foxp3creStk11fl/fl mice. b, Representative images of haematoxylin and eosin staining of colon (original magnification, ×20) and caecum (×10) from wild-type and Foxp3creStk11fl/fl mice. c, Representative images of MCPT1 (mast cell protease 1) staining of colon (original magnification, ×20) and caecum (×20) from wild-type and Foxp3creStk11fl/fl mice. MCPT1 (brown) labels the interepithelial mucosal mast cells (ieMMCs). d, Quantification of serum MCP-1 (WT, n = 10; Foxp3creStk11fl/fl, n = 12), TNF-α (WT, n = 11; Foxp3creStk11fl/fl, n = 12) and IFN-γ (WT, n = 9; Foxp3creStk11fl/fl, n = 8). e, Quantification of IgG2a/c, IgG2b, IgA and IgM in the serum from wild-type and Foxp3creStk11fl/fl mice (n = 6 each group). f, g, Flow cytometry of eosinophils (CD11b+Siglec-F+) in the spleen (f) and lung (g), and alveolar macrophages (CD11b–Siglec-F+) in the lung (g) from wild-type and Foxp3creStk11fl/fl mice. h, i, Flow cytometry of neutrophils (CD11b+Ly6G+, h) and macrophages (CD11b+F4/80+, i) in the spleen from wild-type and Foxp3creStk11fl/fl mice. j, k, Representative images of MBP (major basic protein, pink, which labels eosinophils) (j) and YM1 (brown, which labels M2 macrophages) (k) staining of skin (×20) from wild-type and Foxp3creStk11fl/fl mice. Data are representative of two (a–c, j, k), one (d, e) or at least three (f–i) independent experiments. Data are mean ± s.e.m. P values are determined by two-tailed Student’s t-test (a, d, e). NS, not significant; *P < 0.05, **P < 0.005, ***P < 0.0005. Numbers in gates indicate percentage of cells.
Extended Data Figure 2 Excessive T cell activation and TH2-dominant inflammatory responses in Foxp3creStk11fl/fl mice.
a, Expression of CD62L and CD44 on CD4+ and CD8+ T cells in the spleen from wild-type and Foxp3creStk11fl/fl mice. b, Flow cytometry and quantification of IL-4- (WT, n = 4; Foxp3creStk11fl/fl, n = 5) and IL-5- (n = 4 each group) producing CD4+ T cells in the spleen from wild-type and Foxp3creStk11fl/fl mice (approximately 4–5 weeks old) after in vitro stimulation for 4 h. c, Flow cytometry of IL-4- and IL-5-producing Treg cells in the spleen from wild-type and Foxp3creStk11fl/fl mice. d, e, Flow cytometry and quantification of IFN-γ- (n = 4 each group; d) and IL-17-producing (WT, n = 4; Foxp3creStk11fl/fl, n = 5; e) CD4+ T cells in the spleen from the mice in b. f, Flow cytometry of IL-4-producing CD4+ T cells in the lung (left panel) and colon (right panel) from wild-type and Foxp3creStk11fl/fl mice. g, Expression of IL-5 and IL-17 in CD4+ T cells in the spleen from young wild-type and Foxp3creStk11fl/fl mice (approximately 16 days old). Right, fold changes of IL-5- or IL-17-producing CD4+ T cells from Foxp3creStk11fl/fl versus wild-type counterparts (n = 5 each group). Data are representative of at least three (a–g) independent experiments. Data are mean ± s.e.m. P values are determined by Mann–Whitney test (b, d, e) or two-tailed Student’s t-test (g). *P < 0.05, **P < 0.005, ***P < 0.0005. Numbers in quadrants or gates indicate percentage of cells.
Extended Data Figure 3 LKB1 regulates Treg cell homeostasis and function.
a, Flow cytometry of Treg cells in the lung (left panel) and colon (right panel) from wild-type and Foxp3creStk11fl/fl mice. b, BrdU incorporation in splenic Treg cells from wild-type and Foxp3creStk11fl/fl mice at 16 h after injection of BrdU. c, d, Expression of active caspase-3 (c) and Bim (d) in CD45.2+ Treg cells from CD45.1+:wild-type and CD45.1+:Foxp3creStk11fl/fl mixed bone marrow chimaeras. e, Flow cytometry of splenic Treg cells from young wild-type and Foxp3creStk11fl/fl mice (approximately 16 days old). Right, proportions of Treg cells (WT, n = 7; Foxp3creStk11fl/fl, n = 5). f, g, Expression of Bim (f), and CD62L and CD44 (g) in splenic CD4+ T cells from young wild-type and Foxp3creStk11fl/fl mice (approximately 16 days old). h, i, Representative images of haematoxylin and eosin staining of the lung (h) and measurement of IgE in the serum (i) from young (approximately 16 days old) wild-type (n = 7) and Foxp3creStk11fl/fl mice (n = 4). j, Flow cytometry of splenic Treg cells from wild-type, Foxp3creStk11fl/fl and Foxp3creStk11fl/fl Bcl2l11fl/fl mice. k, Expression of CD62L and CD44 on CD4+ T cells from wild-type, Foxp3creStk11fl/fl and Foxp3creStk11fl/flBcl2l11fl/fl mice. l, Flow cytometry of splenic Treg cells from wild-type and Foxp3creBcl2l11fl/fl mice. m, Expression of IL-4 and IFN-γ in CD4+ T cells from wild-type and Foxp3creBcl2l11fl/fl mice after in vitro stimulation for 4 h. Data are representative of at least three (a–d) or two (e–m) independent experiments. Data are mean ± s.e.m. P values are determined by Mann–Whitney test (e) or two-tailed Student’s t-test (i). NS, not significant; *P < 0.01. Numbers above graphs indicate the mean fluorescence intensity; numbers in quadrants or gates indicate percentage of cells.
Extended Data Figure 4 LKB1 regulates the expression of distinct Treg signature molecules.
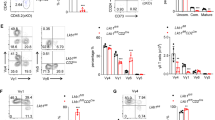
a, Comparison of PD-1, GITR and OX40 expression on Treg cells from wild-type and Foxp3creStk11fl/fl mice. b, Flow cytometry of resting (CD62LhiCD44low) and activated (CD62LlowCD44hi) Treg cells in the spleen from wild-type and Foxp3creStk11fl/fl mice. c, Comparison of GATA3, IRF4, T-bet and RORγt in Treg cells from wild-type and Foxp3creStk11fl/fl mice. d, Comparison of Foxp3, ICOS, CTLA-4 and CD25 expression in Treg cells from wild-type and Foxp3creStk11fl/fl mice. e, Comparison of Foxp3, ICOS, CTLA-4 and CD25 expression in CD45.2+ wild-type and CD45.2+ Foxp3creStk11fl/fl Treg cells from mixed bone marrow chimaeras. f, Expression of Stk11 mRNA in CD4+Foxp3–GFP+YFP+ T cells from Foxp3cre-ERT2Stk11fl/+ and Foxp3cre-ERT2Stk11fl/fl mice (n = 7 each group). g, Flow cytometry of Foxp3–GFP+YFP+ Treg cells from Foxp3cre-ERT2Stk11fl/+ and Foxp3cre-ERT2Stk11fl/fl mice at day 20 after the last tamoxifen administration. h, Proportion of DAPI+ Foxp3–GFP+YFP+ Treg cells in g. i, Flow cytometry of Foxp3–GFP+YFP+ Treg cells from Foxp3cre-ERT2Stk11fl/+ and Foxp3cre-ERT2Stk11fl/fl mice at day 6 after the last tamoxifen administration. j, k, Expression of IL-4 versus IL-13 (j) and IFN-γ versus IL-17 (k) in CD4+ T cells from Foxp3cre-ERT2Stk11fl/+ and Foxp3cre-ERT2Stk11fl/fl mice after in vitro stimulation for 4 h. l, Comparison of Foxp3–GFP, ICOS and CD25 expression on Foxp3–GFP+YFP+ Treg cells from Foxp3cre-ERT2Stk11fl/+ and Foxp3cre-ERT2Stk11fl/fl mice. m, Flow cytometry of resting (CD62LhiCD44lo) and activated (CD62LloCD44hi) Treg cells in the spleen from Foxp3cre-ERT2Stk11fl/+ and Foxp3cre-ERT2Stk11fl/fl mice. Data are representative of at least three (a–i, l, m) or two (j, k) independent experiments. Data are mean ± s.e.m. P values are determined by two-tailed Student’s t-test (f). ***P < 0.0001. Numbers above graphs indicate the mean fluorescence intensity; numbers in quadrants or gates indicate percentage of cells.
Extended Data Figure 5 LKB1 regulates Treg cell function independently of mTOR–HIF-1α axis or AMPK signalling.
a, b, Phosphorylation of S6 (a), and phosphorylation of AKT S473 and FOXO1 (b) in wild-type and Foxp3creStk11fl/fl Treg cells stimulated with or without anti-CD3/CD28 antibodies for 48 h. c, Flow cytometry of splenic Treg cells from wild-type, Foxp3creStk11fl/fl, Foxp3creHif1afl/fl and Foxp3creStk11fl/flHif1afl/fl mice. d, Expression of CD62L and CD44 on CD4+ T cells from the mice in c. e, Expression of IL-4 and IL-17 in CD4+ T cells from the mice in c after in vitro stimulation for 4 h. f, Phosphorylation of AMPK in wild-type and LKB1-deficient Treg cells stimulated with anti-CD3/CD28 antibodies for 48 h. g, Expression of CD62L and CD44 on CD4+ (upper) and CD8+ T cells (lower) from wild-type and Foxp3crePrkaa1fl/flPrkaa2fl/fl mice. h, Expression of IFN-γ, IL-4 and IL-17 in CD4+ (upper) and CD8+ T cells (lower) from wild-type and Foxp3crePrkaa1fl/flPrkaa2fl/fl mice after in vitro stimulation for 4 h. Data are representative of two (a–f) or three (g, h) independent experiments. Numbers in quadrants or gates indicate percentage of cells.
Extended Data Figure 6 LKB1-dependent gene expression and metabolic programs in Treg cells.
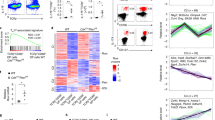
a, Gene set enrichment analysis (GSEA) of transcriptional profiles in wild-type and Foxp3creStk11fl/fl Treg cells from mixed bone marrow chimaeras (n = 5 each group). The list of top 10 gene sets downregulated in Foxp3creStk11fl/fl Treg cells were shown. NES, normalized enrichment score. b, c, The gene sets of TCA cycle and respiratory electron transport (b) and mitochondrial protein import (c) were enriched among the top 10 downregulated pathways in Foxp3creStk11fl/fl Treg cells. d, Heat maps showing relative abundance of metabolites differentially expressed and unsupervised hierarchical clustering. e, f, Relative abundance of metabolites implicated in TCA cycle (e) or glycolytic pathways (f) in Foxp3creStk11fl/+ and Foxp3creStk11fl/fl Treg cells (Foxp3creStk11fl/+, n = 3; Foxp3creStk11fl/fl, n = 4). DHAP, glyceraldehyde 3P; PEP, phosphoenolpyruvic acid. g, Comparison of mitochondrial mass, TMRM (indicative of mitochondrial membrane potential) and ROS production in wild-type and LKB1-deficient Treg cells. h, Relative amount of intracellular ATP in wild-type and LKB1-deficient Treg cells (n = 4 each group). i, j, Oxygen consumption rate (OCR) (n = 8 each group, i) and extracellular acidification rate (ECAR) (n = 4 each group, j) of activated Foxp3–GFP+YFP+ Treg cells from Foxp3cre-ERT2Stk11fl/+ and Foxp3cre-ERT2Stk11fl/fl mice. k, Comparison of lipid droplets in wild-type and LKB1-deficient Treg cells. l, Oxygen consumption rate of activated Foxp3–GFP+YFP+ Treg cells responding to the treatment of inhibitors oligomycin (Oligo), FCCP, etomoxir (Eto) and Rotenone/antimycin A (Rot/AA). m, De novo lipid biosynthesis of Foxp3–GFP+YFP+ Treg cells stimulated with anti-CD3/CD28 antibodies for 48 h (WT, n = 4; Foxp3creStk11fl/fl, n = 3). Data are representative of one (a–c; n = 5 mice each group; d–f; Foxp3creStk11fl/+, n = 3; Foxp3creStk11fl/fl, n = 4) or two (g–m) independent experiments. Data are mean ± s.e.m. P values are determined by two-tailed Student’s t-test (e, f, h–j, m) or two-way ANOVA (l). NS, not significant; *P < 0.05, **P < 0.005. Numbers above graphs indicate the mean fluorescence intensity.
Extended Data Figure 7 Deletion of IL-4 partially rescues dysregulated type 2 immune responses, but not Treg cell function or overall immune homeostasis.
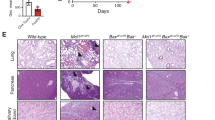
a, b, Expression of IL-4 and IL-13 (a) and IL-5 (b) in splenic CD4+ T cells from wild-type, Foxp3creStk11fl/fl, Il4–/– and Foxp3creStk11fl/flIl4–/– mice after in vitro stimulation for 4 h. c, d, Flow cytometry of eosinophils (CD11b+Siglec-F+) (c) and CD11b+PD-L2+ DCs (gated on CD11c+MHC-II+) (d) in the spleen from the mice in (a). e, Quantification of IgE and IgG1 in the serum from the mice in a (n = 3 each group). f, Flow cytometry of splenic Treg cells from the mice in a. g, Mean fluorescence intensity (MFI) of PD-1, GITR and OX40 expression on Treg cells from the mice in a (WT, n = 4; Foxp3creStk11fl/fl, n = 3; Il4–/–, n = 3; Foxp3creStk11fl/flIl4–/–, n = 3). h, Expression of CD62L and CD44 on splenic CD4+ T cells from the mice in a. i, Expression of IFN-γ and IL-4 in splenic CD4+ T cells from the mice in a after in vitro stimulation for 4 h. j, Representative images of haematoxylin and eosin staining of the lung (original magnification, ×10), liver (×10) and skin (×20) from the mice in a. Data are representative of at least three (a–d, f, h, i) or two (e, g, j) independent experiments. Data are mean ± s.e.m. P values are determined by one-way ANOVA (e, g). NS, not significant; *P < 0.05, **P < 0.005. Numbers in quadrants or gates indicate percentage of cells.
Extended Data Figure 8 Defects of LKB1-deficient Treg cells in suppressing DC maturation and TH2 cell differentiation.
a, Production of IL-25 and IL-33 in the homogenate of the lung from wild-type and Foxp3creStk11fl/fl mice, normalized by the weight of the lung (WT, n = 7; Foxp3creStk11fl/fl, n = 5). b, Fold change of CD80 and CD86 expression on splenic DCs from C57BL/6 mice stimulated with TSLP for 18 h (n = 3 each group). c, Mean fluorescence intensity (MFI) of CD80 and CD86 expression on DCs (with or without TSLP) cultured alone, or together with wild-type or Foxp3creStk11fl/fl Treg cells for 48 h (n = 4 each group). d, Naive CD4+ T cells were co-cultured with TSLP-DCs alone, or together with Treg cells from Foxp3cre-ERT2Stk11fl/+ or Foxp3cre-ERT2Stk11fl/fl mice (after tamoxifen treatment) for 5 days, and expression of Il4, Il5 and Il13 mRNA in co-cultured CD4+ T cells was measured after restimulation with anti-CD3 for 5 h. e, f, Naive CD4+ T cells were co-cultured with LPS-primed DCs alone, or together with Treg cells from Foxp3cre-ERT2Stk11fl/+ or Foxp3cre-ERT2Stk11fl/fl mice (after tamoxifen treatment) for 5 days, followed by analyses of IFN-γ expression after in vitro stimulation for 4 h (e), and Ifng mRNA expression after restimulation with anti-CD3 for 5 h (f). Data are representative of two (a, d–f) or three (b, c) independent experiments. Data are mean ± s.e.m. P values are determined by two-tailed Student’s t-test (a, b) or one-way ANOVA (c). NS, not significant; *P < 0.05, **P < 0.005. Numbers in gates indicate percentage of cells.
Extended Data Figure 9 Appropriate control of PD-1 expression is important for Treg cells in suppressing TH2 immune responses.
a, b, Fold change of PD-1, GITR and OX40 expression on lung Treg cells (a), or splenic Treg cells (b) from OVA-sensitized C57BL/6 mice with or without OVA inhalation challenge (n = 3 each group). c, Expression of IL-4 in CD4+ T cells co-cultured with TSLP-DCs, or together with Treg cells transduced with control retrovirus (RV) or PD-1-expressing retrovirus (Pdcd1-RV). d, Expression of IL-4 in CD4+ T cells co-cultured with TSLP-DCs, or together with wild-type or PD-1-deficient Treg cells. e, Flow cytometry of eosinophils (CD11b+Siglec-F+) in the lung and spleen from Rag1–/– mice reconstituted with Scurfy:wild-type or Scurfy:Pdcd1–/– bone marrow cells. f, Concentration of IgG1 and IgG2a/c in the serum from the mice in (e) (n = 4 each group). Data are representative of two (a–f) independent experiments. Data are mean ± s.e.m. P values are determined by two-tailed Student’s t-test (a, b, f). NS, not significant; *P < 0.05, **P < 0.005. Numbers in gates indicate percentage of cells.
Extended Data Figure 10 Treg cells require the LKB1–β-catenin signalling axis to enforce their functional fitness in maintaining immune homeostasis.
a, GSEA reveals the significant enrichment of the Wnt signalling gene set among the downregulated pathways in LKB1-deficient Treg cells. b, Relative expression of Ctnnb1 mRNA in activated wild-type and LKB1-deficient Treg cells. c, Expression of total and phosphorylated β-catenin in activated wild-type and LKB1-deficient Treg cells. Right, relative phosphorylation of β-catenin normalized by total β-catenin. d, ChIP and real-time PCR analysis of β-catenin-bound DNA of the Pdcd1 locus from activated Foxp3cre-ERT2Stk11fl/+ and Foxp3cre-ERT2Stk11fl/fl Treg cells (following in vivo tamoxifen treatment) (n = 4 each group). e, Mean fluorescence intensity (MFI) of GITR and CD25 expression on wild-type and Foxp3creStk11fl/fl Treg cells transduced with control retrovirus (RV) or mutant β-catenin-expressing retrovirus (β-cat-RV) (n = 3 each group). f, MFI of PD-1, GITR and CD25 expression on Foxp3cre-ERT2Stk11fl/+ (n = 5) and Foxp3cre-ERT2Stk11fl/fl (n = 4) Treg cells (following in vivo tamoxifen treatment) transduced with control RV or β-cat-RV. g, Expression of PD-L2, CD80 and CD86 on DCs co-cultured with wild-type or LKB1-deficient Treg cells transduced with RV or β-cat-RV. Numbers above graphs indicate the MFI. Data are representative of one (a) or two (b–g) independent experiments. Data are mean ± s.e.m. P values are determined by two-tailed Student’s t-test (d) or two-way ANOVA (e, f). NS, not significant; **P < 0.005, ***P < 0.0005. h, Schematics of LKB1 signalling in the regulation of Treg cell function and immune homeostasis. LKB1 signalling in Treg cells establishes metabolic and homeostatic fitness required for preventing undesired immune responses through selectively controlling the expression of inhibitory regulators, including PD-1, GITR and OX40. Consequently, uncontrolled expression of PD-1 and possible other receptors impairs the capability of Treg cells in suppressing TH2 immune responses triggered by TSLP-induced PD-L2+ DCs. Although not depicted here, IL-4 contributes to the induction of PD-L2 on DCs and the amplification of TH2-mediated immunopathology.
Supplementary information
Supplementary information
This file contains Supplementary Figure 1, the uncropped gels with size marker indications.
Source data
Rights and permissions
About this article
Cite this article
Yang, K., Blanco, D., Neale, G. et al. Homeostatic control of metabolic and functional fitness of Treg cells by LKB1 signalling. Nature 548, 602–606 (2017). https://doi.org/10.1038/nature23665
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23665
This article is cited by
-
Lipid metabolism in tumor-infiltrating regulatory T cells: perspective to precision immunotherapy
Biomarker Research (2024)
-
Lkb1 orchestrates γδ T-cell metabolic and functional fitness to control IL-17-mediated autoimmune hepatitis
Cellular & Molecular Immunology (2024)
-
The regulation and differentiation of regulatory T cells and their dysfunction in autoimmune diseases
Nature Reviews Immunology (2024)
-
WNT/β-catenin regulatory roles on PD-(L)1 and immunotherapy responses
Clinical and Experimental Medicine (2024)
-
Peach extract induces systemic and local immune responses in an experimental food allergy model
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.