Abstract
Induced pluripotent stem cells (iPS cells) are a promising source for a cell-based therapy to treat Parkinson’s disease (PD), in which midbrain dopaminergic neurons progressively degenerate1,2. However, long-term analysis of human iPS cell-derived dopaminergic neurons in primate PD models has never been performed to our knowledge. Here we show that human iPS cell-derived dopaminergic progenitor cells survived and functioned as midbrain dopaminergic neurons in a primate model of PD (Macaca fascicularis) treated with the neurotoxin MPTP. Score-based and video-recording analyses revealed an increase in spontaneous movement of the monkeys after transplantation. Histological studies showed that the mature dopaminergic neurons extended dense neurites into the host striatum; this effect was consistent regardless of whether the cells were derived from patients with PD or from healthy individuals. Cells sorted by the floor plate marker CORIN did not form any tumours in the brains for at least two years. Finally, magnetic resonance imaging and positron emission tomography were used to monitor the survival, expansion and function of the grafted cells as well as the immune response in the host brain. Thus, this preclinical study using a primate model indicates that human iPS cell-derived dopaminergic progenitors are clinically applicable for the treatment of patients with PD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Kriks, S. et al. Dopamine neurons derived from human ES cells efficiently engraft in animal models of Parkinson’s disease. Nature 480, 547–551 (2011)
Doi, D. et al. Isolation of human induced pluripotent stem cell-derived dopaminergic progenitors by cell sorting for successful transplantation. Stem Cell Reports 2, 337–350 (2014)
Perrier, A. L. et al. Derivation of midbrain dopamine neurons from human embryonic stem cells. Proc. Natl Acad. Sci. USA 101, 12543–12548 (2004)
Chambers, S. M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat. Biotechnol. 27, 275–280 (2009)
Kirkeby, A. et al. Generation of regionally specified neural progenitors and functional neurons from human embryonic stem cells under defined conditions. Cell Reports 1, 703–714 (2012)
Doi, D. et al. Prolonged maturation culture favors a reduction in the tumorigenicity and the dopaminergic function of human ESC-derived neural cells in a primate model of Parkinson’s disease. Stem Cells 30, 935–945 (2012)
Hargus, G. et al. Differentiated Parkinson patient-derived induced pluripotent stem cells grow in the adult rodent brain and reduce motor asymmetry in Parkinsonian rats. Proc. Natl Acad. Sci. USA 107, 15921–15926 (2010)
Nguyen, H. N. et al. LRRK2 mutant iPSC-derived DA neurons demonstrate increased susceptibility to oxidative stress. Cell Stem Cell 8, 267–280 (2011)
Sánchez-Danés, A. et al. Disease-specific phenotypes in dopamine neurons from human iPS-based models of genetic and sporadic Parkinson’s disease. EMBO Mol. Med. 4, 380–395 (2012)
Kikuchi, T. et al. Idiopathic Parkinson’s disease patient-derived induced pluripotent stem cells function as midbrain dopaminergic neurons in rodent brains. J. Neurosci. Res. 95, 1829–1837 (2017)
Ono, Y. et al. Differences in neurogenic potential in floor plate cells along an anteroposterior location: midbrain dopaminergic neurons originate from mesencephalic floor plate cells. Development 134, 3213–3225 (2007)
Joksimovic, M. et al. Wnt antagonism of Shh facilitates midbrain floor plate neurogenesis. Nat. Neurosci. 12, 125–131 (2009)
Smidt, M. P. et al. A homeodomain gene Ptx3 has highly restricted brain expression in mesencephalic dopaminergic neurons. Proc. Natl Acad. Sci. USA 94, 13305–13310 (1997)
Katsukawa, M., Nakajima, Y., Fukumoto, A., Doi, D. & Takahashi, J. Fail-safe therapy by gamma-ray irradiation against tumor formation by human-induced pluripotent stem cell-derived neural progenitors. Stem Cells Dev. 25, 815–825 (2016)
Imbert, C., Bezard, E., Guitraud, S., Boraud, T. & Gross, C. E. Comparison of eight clinical rating scales used for the assessment of MPTP-induced parkinsonism in the Macaque monkey. J. Neurosci. Methods 96, 71–76 (2000)
Kikuchi, T. et al. Survival of human induced pluripotent stem cell-derived midbrain dopaminergic neurons in the brain of a primate model of Parkinson’s disease. J. Parkinsons Dis. 1, 395–412 (2011)
Takagi, Y. et al. Dopaminergic neurons generated from monkey embryonic stem cells function in a Parkinson primate model. J. Clin. Invest. 115, 102–109 (2005)
Hallett, P. J. et al. Successful function of autologous iPSC-derived dopamine neurons following transplantation in a non-human primate model of Parkinson’s disease. Cell Stem Cell 16, 269–274 (2015)
Freed, C. R. et al. Transplantation of embryonic dopamine neurons for severe Parkinson’s disease. N. Engl. J. Med. 344, 710–719 (2001)
Olanow, C. W. et al. A double-blind controlled trial of bilateral fetal nigral transplantation in Parkinson’s disease. Ann. Neurol. 54, 403–414 (2003)
Kurowska, Z. et al. Signs of degeneration in 12–22-year old grafts of mesencephalic dopamine neurons in patients with Parkinson’s disease. J. Parkinsons Dis. 1, 83–92 (2011)
Li, W. et al. Extensive graft-derived dopaminergic innervation is maintained 24 years after transplantation in the degenerating parkinsonian brain. Proc. Natl Acad. Sci. USA 113, 6544–6549 (2016)
Yin, D. et al. Striatal volume differences between non-human and human primates. J. Neurosci. Methods 176, 200–205 (2009)
Redmond, D. E. Jr, Vinuela, A., Kordower, J. H. & Isacson, O. Influence of cell preparation and target location on the behavioral recovery after striatal transplantation of fetal dopaminergic neurons in a primate model of Parkinson’s disease. Neurobiol. Dis. 29, 103–116 (2008)
Turkheimer, F. E. et al. Reference and target region modeling of [11C]-(R)-PK11195 brain studies. J. Nucl. Med. 48, 158–167 (2007)
Shukuri, M. et al. In vivo expression of cyclooxygenase-1 in activated microglia and macrophages during neuroinflammation visualized by PET with 11C-ketoprofen methyl ester. J. Nucl. Med. 52, 1094–1101 (2011)
Kirkeby, A. et al. Predictive markers guide differentiation to improve graft outcome in clinical translation of hESC-based therapy for Parkinson’s disease. Cell Stem Cell 20, 135–148 (2017)
Liechti, R. et al. Characterization of fetal antigen 1/delta-like 1 homologue expressing cells in the rat nigrostriatal system: effects of a unilateral 6-hydroxydopamine lesion. PLoS ONE 10, e0116088 (2015)
Christophersen, N. S. et al. Midbrain expression of Delta-like 1 homologue is regulated by GDNF and is associated with dopaminergic differentiation. Exp. Neurol. 204, 791–801 (2007)
Bauer, G. et al. In vivo biosafety model to assess the risk of adverse events from retroviral and lentiviral vectors. Mol. Ther. 16, 1308–1315 (2008)
Okita, K. et al. An efficient nonviral method to generate integration-free human-induced pluripotent stem cells from cord blood and peripheral blood cells. Stem Cells 31, 458–466 (2013)
Miyazaki, T. et al. Laminin E8 fragments support efficient adhesion and expansion of dissociated human pluripotent stem cells. Nat. Commun. 3, 1236 (2012)
Nakagawa, M. et al. A novel efficient feeder-free culture system for the derivation of human induced pluripotent stem cells. Sci. Rep. 4, 3594 (2014)
Morizane, A., Doi, D., Kikuchi, T., Nishimura, K. & Takahashi, J. Small-molecule inhibitors of bone morphogenic protein and activin/nodal signals promote highly efficient neural induction from human pluripotent stem cells. J. Neurosci. Res. 89, 117–126 (2011)
Smith, S. M. et al. Advances in functional and structural MR image analysis and implementation as FSL. Neuroimage 23 (Suppl. 1), S208–S219 (2004)
Smith, S. M. Fast robust automated brain extraction. Hum. Brain Mapp. 17, 143–155 (2002)
Jenkinson, M. & Smith, S. A global optimisation method for robust affine registration of brain images. Med. Image Anal. 5, 143–156 (2001)
Jenkinson, M., Bannister, P., Brady, M. & Smith, S. Improved optimization for the robust and accurate linear registration and motion correction of brain images. Neuroimage 17, 825–841 (2002)
Zhang, Y., Brady, M. & Smith, S. Segmentation of brain MR images through a hidden Markov random field model and the expectation-maximization algorithm. IEEE Trans. Med. Imaging 20, 45–57 (2001)
Frey, S. et al. An MRI based average macaque monkey stereotaxic atlas and space (MNI monkey space). Neuroimage 55, 1435–1442 (2011)
Warschausky, S., Kay, J. B. & Kewman, D. G. Hierarchical linear modeling of FIM instrument growth curve characteristics after spinal cord injury. Arch. Phys. Med. Rehabil. 82, 329–334 (2001)
Jucaite, A., Fernell, E., Halldin, C., Forssberg, H. & Farde, L. Reduced midbrain dopamine transporter binding in male adolescents with attention-deficit/hyperactivity disorder: association between striatal dopamine markers and motor hyperactivity. Biol. Psychiatry 57, 229–238 (2005)
Leroy, C. et al. Assessment of 11C-PE2I binding to the neuronal dopamine transporter in humans with the high-spatial-resolution PET scanner HRRT. J. Nucl. Med. 48, 538–546 (2007)
Logan, J. et al. Distribution volume ratios without blood sampling from graphical analysis of PET data. J. Cereb. Blood Flow Metab. 16, 834–840 (1996)
Patlak, C. S. & Blasberg, R. G. Graphical evaluation of blood-to-brain transfer constants from multiple-time uptake data. Generalizations. J. Cereb. Blood Flow Metab. 5, 584–590 (1985)
Sossi, V., Holden, J. E., de la Fuente-Fernandez, R., Ruth, T. J. & Stoessl, A. J. Effect of dopamine loss and the metabolite 3-O-methyl-[18F]fluoro-dopa on the relation between the 18F-fluorodopa tissue input uptake rate constant Kocc and the [18F]fluorodopa plasma input uptake rate constantK i . J. Cereb. Blood Flow Metab. 23, 301–309 (2003)
Acknowledgements
We thank K. Sekiguchi, J. Toga and E. Yagi for providing recombinant LM511-E8, Y. Ono for anti-CORIN and anti-NURR1 antibodies, H. Doi, A. Mawatari, M. Tsuji, K. Takahashi, M. Goto, Y. Wada, A. Yamazaki, T. Kawasaki, C. Takeda, N. Shibata, S. Kurai, A. Igesaka, T. Mori, R. Zochi, E. Hayashinaka, M. Yamano, T. Ose, M. Ohno and K. Onoe for supporting the PET study, H. Ohmori for an electrophysiological study, S. Nolbrant for discussions about gene expression by the donor cells, and Astellas Pharma Inc. for FK506. We also thank P. Karagiannis for reading of the manuscript, K. Kubota, Y. Ishii, Y. Morita and Y. Katano for technical assistance, K. Nishimura, M. Motono, Y. Ioroi, B. Samata, Y. Koshiba, Y. Nakajima and Y. Miyawaki for taking care of the animals, and S. Tsuji, J. Mitsui and S. Morishita for whole-exome analysis of patients with PD. This study was supported by grants from the Highway Project for Realization of Regenerative Medicine from the Ministry of Education, Culture, Sports, Science and Technology (MEXT), the Network Program for Realization of Regenerative Medicine from the Japan Agency for Medical Research and Development (AMED) and the Program for Intractable Diseases Research using disease-specific iPS cells from AMED (to H.I.). M.P. is a New York Stem Cell Foundation - Robertson Investigator.
Author information
Authors and Affiliations
Contributions
T.K. designed the study, performed the culture, transplantation, MRI study, data analysis and interpretation, and wrote the manuscript. A.M. and D.D. assisted with cell culture, cell sorting, transplantation and the MRI study. H.Ma. generated the PD model monkeys and performed the behavioural analysis. H.O., T.H., H.Mi. and S.T. performed the PET imaging and corresponding analysis and interpretation. R.T., H.I., K.O. and M.N. generated the iPS cells. S.M. and M.Y. performed statistical analyses. M.P. provided fetal mesencephalic samples and discussed the gene expression analyses. J.T. conceived and designed the study, assembled the data, carried out the data analysis and interpretation, wrote the manuscript, and gave final approval of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks R. Barker, A. Björklund and F. Gage for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 In vitro analysis of dopaminergic neuron progenitors.
a, The protocol for the induction of dopamine neuron progenitors. Y, Y-27632; BDNF, brain-derived neurotrophic factor; GDNF, glial cell line-derived neurotrophic factor; AA, ascorbic acid; dbcAMP, dibutyryladenosine cyclic monophosphate. b, Percentages of CORIN+ cells at day 12. Values are mean ± s.d. (n = 4 each for PD group and healthy group). c, Representative images of immunostaining for FOXA2, NURR1, TUJ1, PAX6, and SOX1 at day 26. Scale bar, 50 μm. d, e, Quantification of immunostaining for FOXA2 (d) and NURR1 (e) at day 26. Values are mean ± s.d. (n = 4 each for PD group and healthy group). f, A representative current-clamp recording of the action potentials induced by brief current pulses at day 70 (1231A3). g, A representative chromatogram from HPLC analysis at day 42 (1231A3). DOPAC, 3,4-dihydroxyphenylacetic acid. t-tests were performed in b, d, and e. There was no significant difference between healthy and PD groups.
Extended Data Figure 2 Behavioural analysis of monkeys.
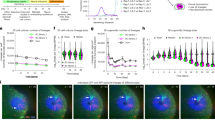
a, Rating scale of PD model monkeys. b, PD scores of each monkey. c–f, The video analysis system. Schematic view of the video recording system (c), simplified illustration of the special cage for the video recording (d), a photo of the cage with the LED backlight switched on (e), and a representative capture from the video recording (f). g–j, Representative captures of the video analysis when the monkey’s number of movements was quantified as 5,316 (g) and 15,212 (i), and the moving time of each monkey analysed by video recording when the threshold was set to 5,000 (h) or 10,000 (j) pixels per 0.033 seconds. In h and j, values are shown relative to each pre-operative value, which was set to 1. k, l, Improvement of monkey PD scores (k) or fold change in spontaneous movement analysed by video recording (l) after administration of one-shot l-DOPA or transplantation. Horizontal bars designate the mean value. Two-tailed Wilcoxon matched-pairs signed rank-test was performed. *P < 0.05.
Extended Data Figure 3 Estimation of graft growth.
a, Estimated maximum volume of the grafts within 95% confidence upper limit analysed by a linear mixed effect model. b, Correlation between the graft volumes calculated from MRI and measured by histological analysis (n = 16). Data were compared using a two-tailed Pearson’s correlation analysis, and r and P values and linear regression lines are shown.
Extended Data Figure 4 Tyrosine hydroxylase histology of monkeys.
a, Representative tyrosine hydroxylase staining of each monkey. Scale bars, 5 mm. b, Representative magnified view of TH+ cells in each graft. Scale bars, 100 μm (left) and 50 μm (right).
Extended Data Figure 5 [18F]DOPA- and [11C]PE2I-PET of monkeys.
a, Binding potential (BPnd) values of [11C]PE2I-PET. Lines show mean values (n = 3 for vehicle and PD groups, 4 for healthy group). b, BPnd values of [11C]PE2I-PET in each monkey. c, [18F]DOPA- and [11C]PE2I-PET of each monkey. Dotted white lines designate the putamen.
Extended Data Figure 6 Correlation between surviving TH+ cells and functional recovery.
a, b, Correlation between the number of surviving TH+ cells and score improvement (a) and moving time analysed from the video recording (b). c, d, Correlation between tyrosine hydroxylase-innervated area and score improvement (c) and moving time (d). Two-tailed Pearson’s correlation analysis was performed, and r and P values are shown. Data for the healthy group are shown in blue, the PD group in red, and the vehicle group in black.
Extended Data Figure 7 Inflammation in the brains of cell-transplanted monkeys.
a, Uptake ratios of [11C]PK11195-PET and S-[11C]KTP-Me-PET in representative monkeys. Dotted white lines designate the putamen. b, c, Ratio of standardized uptake values of [11C]PK11195-PET (b) and S-[11C] KTP-Me-PET (c). d, Representative images of MHC class II, CD45, and monkey IgG staining of monkey number 9. Dotted lines designate the grafted area. Scale bar, 5 mm for the left side of MHC class II, CD45, and monkey IgG, and 50 μm for the right side of MHC class II.
Extended Data Figure 9 Gene expression analysis of the transplanted cells.
a, Principal component analysis of the transplanted cells (healthy group in blue, PD group in red), two iPS cells (836B3 and 1231A3, black), fetal ventral midbrain tissue (fVM, green), fetal dorsal midbrain tissue (fDM, green), adult whole brain tissue (WB, navy), and adult substantia nigra tissue (SN, navy). b, Gene list obtained from the microarray analysis. c, Quantitative PCR analysis of the transplanted cells. Values are expressed as relative quantity (RQ).
Supplementary information
Source data
Rights and permissions
About this article
Cite this article
Kikuchi, T., Morizane, A., Doi, D. et al. Human iPS cell-derived dopaminergic neurons function in a primate Parkinson’s disease model. Nature 548, 592–596 (2017). https://doi.org/10.1038/nature23664
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23664
This article is cited by
-
Continuous immunosuppression is required for suppressing immune responses to xenografts in non-human primate brains
Cell Regeneration (2024)
-
CRISPR/sgRNA-directed synergistic activation mediator (SAM) as a therapeutic tool for Parkinson´s disease
Gene Therapy (2024)
-
Graft-derived neurons and bystander effects are maintained for six months after human iPSC-derived NESC transplantation in mice’s cerebella
Scientific Reports (2024)
-
Evaluation of the Efficacy of Stem Cells Therapy in the Periodontal Regeneration: A Meta-Analysis and Mendelian Randomization Study
Stem Cell Reviews and Reports (2024)
-
Unravelling the Parkinson’s puzzle, from medications and surgery to stem cells and genes: a comprehensive review of current and future management strategies
Experimental Brain Research (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.