Abstract
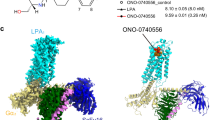
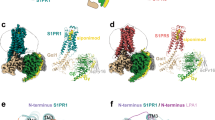
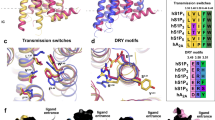
Lysophosphatidic acid (LPA) is a bioactive lipid composed of a phosphate group, a glycerol backbone, and a single acyl chain that varies in length and saturation. LPA activates six class A G-protein-coupled receptors to provoke various cellular reactions1. Because LPA signalling has been implicated in cancer2 and fibrosis3, the LPA receptors are regarded as promising drug targets. The six LPA receptors are subdivided into the endothelial differentiation gene (EDG) family (LPA1–LPA3)1 and the phylogenetically distant non-EDG family (LPA4–LPA6)4. The structure of LPA1 has enhanced our understanding of the EDG family of LPA receptors5. By contrast, the functional and pharmacological characteristics of the non-EDG family of LPA receptors have remained unknown, owing to the lack of structural information. Although the non-EDG LPA receptors share sequence similarity with the P2Y family of nucleotide receptors4, the LPA recognition mechanism cannot be deduced from the P2Y1 and P2Y12 structures6,7,8 because of the large differences in the chemical structures of their ligands. Here we determine the 3.2 Å crystal structure of LPA6, the gene deletion of which is responsible for congenital hair loss9,10, to clarify the ligand recognition mechanism of the non-EDG family of LPA receptors. Notably, the ligand-binding pocket of LPA6 is laterally open towards the membrane, and the acyl chain of the lipid used for the crystallization is bound within this pocket, indicating the binding mode of the LPA acyl chain. Docking and mutagenesis analyses also indicated that the conserved positively charged residues within the central cavity recognize the phosphate head group of LPA by inducing an inward shift of transmembrane helices 6 and 7, suggesting that the receptor activation is triggered by this conformational rearrangement.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Kihara, Y., Maceyka, M., Spiegel, S. & Chun, J. Lysophospholipid receptor nomenclature review: IUPHAR Review 8. Br. J. Pharmacol. 171, 3575–3594 (2014)
Mills, G. B. & Moolenaar, W. H. The emerging role of lysophosphatidic acid in cancer. Nat. Rev. Cancer 3, 582–591 (2003)
Tager, A. M. et al. The lysophosphatidic acid receptor LPA1 links pulmonary fibrosis to lung injury by mediating fibroblast recruitment and vascular leak. Nat. Med. 14, 45–54 (2008)
Yanagida, K., Kurikawa, Y., Shimizu, T. & Ishii, S. Current progress in non-Edg family LPA receptor research. Biochim. Biophys. Acta 1831, 33–41 (2013)
Chrencik, J. E. et al. Crystal structure of antagonist bound human lysophosphatidic acid receptor 1. Cell 161, 1633–1643 (2015)
Zhang, K. et al. Structure of the human P2Y12 receptor in complex with an antithrombotic drug. Nature 509, 115–118 (2014)
Zhang, J. et al. Agonist-bound structure of the human P2Y12 receptor. Nature 509, 119–122 (2014)
Zhang, D. et al. Two disparate ligand-binding sites in the human P2Y1 receptor. Nature 520, 317–321 (2015)
Pasternack, S. M. et al. G protein-coupled receptor P2Y5 and its ligand LPA are involved in maintenance of human hair growth. Nat. Genet. 40, 329–334 (2008)
Shimomura, Y. et al. Disruption of P2RY5, an orphan G protein-coupled receptor, underlies autosomal recessive woolly hair. Nat. Genet. 40, 335–339 (2008)
Khandoga, A. L., Pandey, D., Welsch, U., Brandl, R. & Siess, W. GPR92/LPA5 lysophosphatidate receptor mediates megakaryocytic cell shape change induced by human atherosclerotic plaques. Cardiovasc. Res. 90, 157–164 (2011)
Sumida, H. et al. LPA4 regulates blood and lymphatic vessel formation during mouse embryogenesis. Blood 116, 5060–5070 (2010)
Igarashi, H., Akahoshi, N., Ohto-Nakanishi, T., Yasuda, D. & Ishii, S. The lysophosphatidic acid receptor LPA4 regulates hematopoiesis-supporting activity of bone marrow stromal cells. Sci. Rep. 5, 11410 (2015)
Lin, M.-E., Rivera, R. R. & Chun, J. Targeted deletion of LPA5 identifies novel roles for lysophosphatidic acid signaling in development of neuropathic pain. J. Biol. Chem. 287, 17608–17617 (2012)
Yanagida, K. et al. Identification and characterization of a novel lysophosphatidic acid receptor, p2y5/LPA6 . J. Biol. Chem. 284, 17731–17741 (2009)
Lee, M. et al. P2Y5 is a Gαi, Gα12/13 G protein-coupled receptor activated by lysophosphatidic acid that reduces intestinal cell adhesion. Am. J. Physiol. Gastrointest. Liver Physiol. 297, G641–G654 (2009)
Inoue, A. et al. LPA-producing enzyme PA-PLA1α regulates hair follicle development by modulating EGFR signalling. EMBO J. 30, 4248–4260 (2011)
Kazantseva, A. et al. Human hair growth deficiency is linked to a genetic defect in the phospholipase gene LIPH. Science 314, 982–985 (2006)
Ketscher, A. et al. LSD1 controls metastasis of androgen-independent prostate cancer cells through PXN and LPAR6. Oncogenesis 3, e120 (2014)
Inoue, A. et al. TGFα shedding assay: an accurate and versatile method for detecting GPCR activation. Nat. Methods 9, 1021–1029 (2012)
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nat. Protocols 4, 706–731 (2009)
Zhang, C. et al. High-resolution crystal structure of human protease-activated receptor 1. Nature 492, 387–392 (2012)
Jiang, G., Inoue, A., Aoki, J. & Prestwich, G. D. Phosphorothioate analogs of sn-2 radyl lysophosphatidic acid (LPA): metabolically stabilized LPA receptor agonists. Bioorg. Med. Chem. Lett. 23, 1865–1869 (2013)
Kruse, A. C. et al. Activation and allosteric modulation of a muscarinic acetylcholine receptor. Nature 504, 101–106 (2013)
Shihoya, W. et al. Activation mechanism of endothelin ETB receptor by endothelin-1. Nature 537, 363–368 (2016)
Nishimasu, H. et al. Crystal structure of autotaxin and insight into GPCR activation by lipid mediators. Nat. Struct. Mol. Biol. 18, 205–212 (2011)
Sonoda, H. et al. A novel phosphatidic acid-selective phospholipase A1 that produces lysophosphatidic acid. J. Biol. Chem. 277, 34254–34263 (2002)
Dranoff, J. A. et al. A primitive ATP receptor from the little skate Raja erinacea. J. Biol. Chem. 275, 30701–30706 (2000)
Hanson, M. A. et al. Crystal structure of a lipid G protein-coupled receptor. Science 335, 851–855 (2012)
Cherezov, V. et al. High-resolution crystal structure of an engineered human β2-adrenergic G protein-coupled receptor. Science 318, 1258–1265 (2007)
Hirata, K. et al. Achievement of protein micro-crystallography at SPring-8 beamline BL32XU. J. Phys. Conf. Ser. 425, 12002 (2013)
Ueno, G . et al. Remote access and automation of SPring-8 MX beamlines. AIP Conf. Proc. 1741, 50021 (2016)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Vagin, A. A. et al. REFMAC5 dictionary: organization of prior chemical knowledge and guidelines for its use. Acta Crystallogr. D 60, 2184–2195 (2004)
Adams, P. D . et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Šali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993)
Sastry, G. M., Adzhigirey, M., Day, T., Annabhimoju, R. & Sherman, W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided Mol. Des. 27, 221–234 (2013)
Olsson, M. H. M., Søndergaard, C. R., Rostkowski, M. & Jensen, J. H. PROPKA3: consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory Comput. 7, 525–537 (2011)
Friesner, R. A . et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004)
Halgren, T. A . et al. Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J. Med. Chem. 47, 1750–1759 (2004)
Halgren, T. New method for fast and accurate binding-site identification and analysis. Chem. Biol. Drug Des. 69, 146–148 (2007)
Bowers, K. J. et al. Scalable algorithms for molecular dynamics simulations on commodity clusters. Proc. 2006 ACM/IEEE Conf. Supercomputing (SC06)http://dx.doi.org/10.1145/1188455.1188544 (Tampa, Florida, 11–17 November 2006)
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577 (1995)
Okudaira, M. et al. Separation and quantification of 2-acyl-1-lysophospholipids and 1-acyl-2-lysophospholipids in biological samples by LC-MS/MS. J. Lipid Res. 55, 2178–2192 (2014)
Hayashi, R., Inoue, A., Suga, Y., Aoki, J. & Shimomura, Y. Analysis of unique mutations in the LPAR6 gene identified in a Japanese family with autosomal recessive woolly hair/hypotrichosis: establishment of a useful assay system for LPA6. J. Dermatol. Sci. 78, 197–205 (2015)
Devost, D. et al. Conformational profiling of the AT1 angiotensin II receptor reflects biased agonism, G protein coupling, and cellular context. J. Biol. Chem. 292, 5443–5456 (2017)
Ehlert, F. J ., Griffin, M. T ., Sawyer, G. W . & Bailon, R. A simple method for estimation of agonist activity at receptor subtypes: comparison of native and cloned M3 muscarinic receptors in guinea pig ileum and transfected cells. J. Pharmacol. Exp. Ther. 289, 981–992 (1999)
Hattori, M., Hibbs, R. E. & Gouaux, E. A fluorescence-detection size-exclusion chromatography-based thermostability assay for membrane protein precrystallization screening. Structure 20, 1293–1299 (2012)
Acknowledgements
We thank H. Nishimasu for critical comments on the manuscript; T. Nakane for assistance in the diffraction data analyses; K. Ohgomori for technical assistance; A. Inoue for flow cytometry analyses; F. M. N. Kadji for the TGFα shedding assay; Y. Nakamura for the LPA mass spectrometry analysis; and the beamline staff at BL32XU of SPring-8 (Hyogo, Japan) for assistance with data collection. The diffraction experiments were performed at SPring-8 BL32XU (proposals 2015A1024, 2015A1057, 2015B2024, 2015B2057 and 2016A2527), with the approval of RIKEN. This work was supported by grants from the AMED-CREST, by the Platform for Drug Discovery, Informatics and Structural Life Science from AMED, and by a MEXT Grant-in-Aid for Specially Promoted Research (grant 16H06294) to O.N. This work was also supported by JSPS KAKENHI (grant 16J07583 to R.T.; 15H06862 to K.Y.; 17H05000 to T.N.; 16H06574 to R.I.). A.I. was funded by JST, PRESTO (grant JPMJPR1331), and the PRIME from AMED. H.K. and T.N. were funded by JST, PRESTO (grants JPMJPR14L9 and JPMJPR14L8, respectively). J.A. received funding from the AMED-CREST, AMED, and a MEXT Grant-in-Aid for Scientific Research on Innovative Areas (grant 15H05897).
Author information
Authors and Affiliations
Contributions
R.T. purified and crystallized LPA6, determined the structure, planned the mutational analyses, and performed the ligand-binding assays and thermostability measurements. A.I. and A.U. performed the TGFα shedding assays, flow cytometry experiments, and analysed the data. M.S. performed the ligand-docking simulations under the supervision of Y.O. and T.O. K.Y. and K.H. assisted with the diffraction data collection and the data analyses. M.Y. performed the ligand synthesis under the supervision of T.D. Y.T. performed the preliminary expression check of LPA6. H.E.K. and Y.N.-N. assisted with the structure determination. T.N. assisted with the data analyses. R.T., A.I., M.S., T.N., J.A. and O.N. wrote the manuscript, with feedback from all of the authors. T.O., R.I., J.A. and O.N. supervised the research.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks T. Shimizu, I. von Kügelgen and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Sequence alignment of LPA6
The amino acid sequence alignment of zebrafish LPA6 (DrLPA6), human LPA6 (HsLPA6), mouse LPA6 (MmLPA6) and chicken LPA6 (GgLPA6). Fully conserved residues are highlighted in blue, and well-conserved residues are highlighted in grey. The secondary structure of LPA6 is shown above the alignment. Dashed lines indicate the disordered regions in the crystal structure. The residue numbers are based on DrLPA6.
Extended Data Figure 2 Characterization and comparison of zebrafish and human LPA6 activities with various LPA species.
a, Chemical structures of LPA6 ligands used for the functional analyses and the docking simulation. The structures of 1-oleoyl (18:1)-LPA, 1-linoleoyl (18:2)-LPA, 2-linoleoyl (18:2)-LPA, and 2-alkyl-OMPT-(R) are shown. As compared with 2-oleoyl (18:1)-LPA (not shown), 2-alkyl-OMPT-(R) has three modifications: a thiophosphate polar head group (open arrowhead), which makes the molecule resistant to degradation by a lipid phosphate phosphatase; a methoxy group at the sn-1 position (filled arrowhead), which blocks intramolecular acyl chain migration; and an ester linkage (alkyl group) at the sn-2 position (arrow). b, G12/13-dependent TGFα shedding response of LPA6. Receptor activities of N-terminally epitope-tagged and C-terminally truncated zebrafish LPA6 (DrLPA6) and full-length human LPA6 (HsLPA6) were measured by the TGFα shedding assay using G12/13-deficient cells, Gq/11-deficient cells and their parental HEK293A cells. Receptor-specific signals (grey symbols) indicate the percentage difference in AP–TGFα release between LPA6-expressing cells (filled symbols) and mock-transfected cells (open symbols). Data are mean and s.e.m. of four independent experiments, each performed with two or three biological replicates. Values at the top of each panel indicate mean ± s.e.m. (n = 4) of pEC50 values, calculated by fitting a sigmoidal concentration–response curve to the receptor-specific signals. Note that the origin of the parental cells is different from the HEK293FT cells, which have a low background LPA response and were used in the rest of the experiments. c, Activities of 1-acyl LPA species (with acyl chains differing in length and/or saturation) and 2-alkyl-OMPT-(R) towards LPA6. Data are mean and s.e.m. of four independent experiments, each performed with three biological replicates. Values on the right indicate mean ± s.e.m. (n = 4) of pEC50 values, calculated by fitting a sigmoidal concentration–response curve to the receptor-specific signals. Values in parentheses indicate mean ± s.e.m. (n = 4) of log10 relative intrinsic activity (RAi) values, in which LPA (18:1) was set as the reference compound (by definition, the relative intrinsic activity values for LPA (18:1) and its logarithm are equal to 1 and 0, respectively). Although the purchased LPA species were synthesized as 1-acyl forms, 1-acyl LPA and 2-acyl LPA co-exist in an approximately 9:1 ratio in an aqueous solution, owing to intramolecular migration of the acyl chain. The measured receptor activity reflects the responses towards both 1-acyl and 2-acyl LPA species.
Extended Data Figure 3 Electron density maps of LPA6
a, Overall structure of the LPA6 crystallization construct, including T4L fused between TM5 and TM6, and the C-terminal linker (A(+1)SSEDLYFQ(+9)) derived from the TEV protease cleavage sequence. The receptor is coloured and viewed as in Fig. 1a. T4L is shown as grey cylinders, and the C-terminal linker is coloured purple, with the side chains shown as stick models. b, Close-up view of the C-terminal linker between the N and C lobes of T4L. The 2Fo − Fc electron density map of the C-terminal loops (I309–S312 of LPA6 and the linker region), contoured at 1.0σ, is shown as a green mesh. The disordered side chain is labelled with an asterisk. c, Stereo view of the 2Fo − Fc electron density map (contoured at 1.0σ) around the conserved motifs (D/ERY, NPXXY), viewed from the same orientation as in b. The map is shown as a green mesh. TM1, TM2, TM4 and MO3 are omitted for clarity. d, Stereo view of the 2Fo − Fc electron density map (contoured at 1.0σ) around the vertical cleft, viewed from the same orientation as in a. TM1, TM2, TM6 and TM7 are omitted for clarity.
Extended Data Figure 4 Structural comparison of LPA6 with class A GPCRs.
a, Superimposition of the LPA6 structure with the P2Y1–antagonist complex structure (PDB code 4XNV), viewed from the extracellular side, the membrane plane, and the cytoplasmic side. b, Comparison of the structures around the D/ERY and the NPXXY motifs. The structures of LPA6, the P2Y1–antagonist complex (PDB code 4XNV), and the P2Y12–antagonist complex (PDB code 4NTJ) are shown, with the inactive and active structures of the β2 adrenergic receptor (β2AR; PDB codes 2RH1 and 3SN6, respectively), the M2 muscarinic acetylcholine receptor (M2R; PDB codes 3UON and 4MQS, respectively), and the μ-opioid receptor (μOR; PDB codes 4DKL and 5C1M, respectively). In each image, TM6 is omitted for clarity. In the active structures of β2AR, M2R and μOR, the relative motions of R3.50 and Y7.53, as compared to the inactive structures, are depicted as red arrows. The D7.49 residues of P2Y1 and P2Y12 were mutated to N in the determined structures, and thus these residues are labelled with asterisks. The orientations of the R3.50 and Y7.53 side chains of LPA6 indicate that the LPA6 structure represents the inactive state. c–e, Structural comparison of LPA6 with LPA1. Overall structures of the monoolein-bound LPA6 (c), the LPA-docked model of LPA6 (binding model 2) (d), and the LPA1–antagonist complex (PDB code 4Z34) (e) are shown as both cylinder models and surface representations. In each panel, the two images on the left are viewed from the same orientation as in Fig. 1a. The bottom right images are the cut-away surface representations of the structures, showing the shapes of the ligand-binding cavities. In c, MO2 and MO3 are omitted for clarity.
Extended Data Figure 5 Mutagenesis analyses using the TGFα shedding assay.
a, Receptor activity of wild-type zebrafish LPA6 towards 1-oleoyl (18:1)-LPA, 1-linoleoyl (18:2)-LPA and 2-alkyl-OMPT-(R). Cells transfected with N-terminally epitope-tagged and C-terminally truncated zebrafish LPA6 (wild type) or an empty vector (mock transfection) were subjected to the TGFα shedding assay. Raw AP–TGFα release signals in cells transfected with wild-type LPA6 are shown as coloured symbols (1-oleoyl (18:1)-LPA, red; 1-linoleoyl (18:2)-LPA, yellowish green; 2-alkyl-OMPT-(R), light blue), and those in mock-transfected cells are denoted as grey symbols. Receptor-specific AP–TGFα release signals (percentage difference in AP–TGFα release between LPA6-transfected cells and mock-transfected cells; legends are on the right) are shown in each panel and overlaid in the right panel. b, Receptor activities of 32 LPA6 point mutants examined in this study. As in a, each mutant LPA6 was expressed in HEK293 cells and its receptor-specific responses towards the three ligands were examined using the TGFα shedding assay. Data are mean and s.e.m. of 5–14 independent experiments, each performed with two or three biological replicates. See Extended Data Table 2 for parameters obtained from the concentration–response curves.
Extended Data Figure 6 Biochemical analyses of the mutant LPA6 constructs and the LPA-producing enzyme PA-PLA1α.
a, b, The total binding and the non-specific binding of [3H]1-oleoyl (18:1)-LPA towards the mutants of positively charged residues (n = 9) (a) and the mutants of residues within the hydrophobic cleft (n = 8) (b). Data are mean ± s.e.m. c, Representative fluorescence-detection size-exclusion chromatography (FSEC) profiles of LPA6–eGFP, heated at the respective temperatures and detected by eGFP fluorescence. d, Melting curve of LPA6–eGFP derived from eight repetitive FSEC-TS measurements. Note that the error bars are smaller than the symbols. e, List of the melting temperatures (Tm values), determined by fitting the melting curves to a sigmoidal dose–response equation. f, LPA production by PA-PLA1α accumulated in the cellular fraction, but not in conditioned media. HEK293 cells were transfected with a plasmid encoding wild-type PA-PLA1α or a catalytically inactive mutant (S154A). After a 1-day incubation, conditioned media (CM) and cells were prepared, and the amounts of LPA species were quantified by LC–MS/MS. Among the LPA species measured by LC–MS/MS, 1-oleoyl (18:1)-LPA and palmitoyl (16:0)-LPA were detectable. ND, not detectable. g, LPA6 activation by PA-PLA1α. HEK293 cells were transfected with a mixture of a plasmid encoding wild-type or S154A PA-PLA1α, a human LPA6-encoding plasmid or an empty vector (Mock), and the AP–TGFα reporter-encoding plasmid. After a 1-day incubation, the amount of AP–TGFα released into the conditioned media was quantified. Data are expressed by subtracting the values obtained from a mock transfection (an empty plasmid together with AP–TGFα-encoding plasmid and human LPA6-encoding plasmid) from those of the PA-PLA1α plasmid-transfected condition. Data are representative of two independent experiments, each performed with biological triplicates.
Extended Data Figure 7 Preference of the LPA6 mutants for LPA species.
Receptor activities of zebrafish LPA6 mutants (L1153.41F, L1534.52F, and V2015.45F) towards LPA species (with acyl chains differing in length and/or saturation) and 2-alkyl-OMPT-(R). All of the data and the values were collected, analysed and represented in the same manner as in Extended Data Fig. 2c, but using point mutants of zebrafish LPA6 constructs. Data are mean and s.e.m. of three to five independent experiments each performed with three biological replicates. Values indicate mean ± s.e.m. (n = 3–5) of pEC50 values, with log10 relative intrinsic activity (RAi) values in parentheses.
Extended Data Figure 8 Sequence alignment of LPA6 and phylogenetically related receptors.
The amino acid sequence alignment of zebrafish LPA6 (DrLPA6), human non-EDG family LPA receptors (HsLPA4, HsLPA5, HsLPA6), human LPS receptors (HsLPS1 (also known as GPR34), HsLPS2 (P2Y10) HsLPS3 (GPR174)), human P2Y1 receptor (HsP2Y1), and human P2Y12 receptor (HsP2Y12). Strictly conserved residues are highlighted with coloured letters in the blue boxes, and highly conserved residues are highlighted with coloured letters in a grey background. The residue types are coloured as follows: hydrophobic residues (A, V, L, I and M), magenta; aromatic residues (F, Y and W), blue; polar residues (S, T, N and Q), brown; acidic residues (D and E), red; basic residues (H, K and R), cyan; glycine and proline (G and P), orange; cysteine (C), green. The positions of the LPA6 transmembrane helices are shown above the alignment. The residue numbers are based on DrLPA6.
Supplementary information
Supplementary Information
This file contains Supplementary Methods.
Rights and permissions
About this article
Cite this article
Taniguchi, R., Inoue, A., Sayama, M. et al. Structural insights into ligand recognition by the lysophosphatidic acid receptor LPA6. Nature 548, 356–360 (2017). https://doi.org/10.1038/nature23448
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23448
This article is cited by
-
Structural basis for lysophosphatidylserine recognition by GPR34
Nature Communications (2024)
-
Bitter taste receptor activation by cholesterol and an intracellular tastant
Nature (2024)
-
Structural basis of hydroxycarboxylic acid receptor signaling mechanisms through ligand binding
Nature Communications (2023)
-
Structural basis of lysophosphatidylserine receptor GPR174 ligand recognition and activation
Nature Communications (2023)
-
LPAR2 correlated with different prognosis and immune cell infiltration in head and neck squamous cell carcinoma and kidney renal clear cell carcinoma
Hereditas (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.