Abstract
Many retinal diseases lead to the loss of retinal neurons and cause visual impairment. The adult mammalian retina has little capacity for regeneration. By contrast, teleost fish functionally regenerate their retina following injury, and Müller glia (MG) are the source of regenerated neurons1,2,3,4,5,6. The proneural transcription factor Ascl1 is upregulated in MG after retinal damage1,7 in zebrafish and is necessary for regeneration8. Although Ascl1 is not expressed in mammalian MG after injury9, forced expression of Ascl1 in mouse MG induces a neurogenic state in vitro10 and in vivo after NMDA (N-methyl-d-aspartate) damage in young mice11. However, by postnatal day 16, mouse MG lose neurogenic capacity, despite Ascl1 overexpression11. Loss of neurogenic capacity in mature MG is accompanied by reduced chromatin accessibility, suggesting that epigenetic factors limit regeneration. Here we show that MG-specific overexpression of Ascl1, together with a histone deacetylase inhibitor, enables adult mice to generate neurons from MG after retinal injury. The MG-derived neurons express markers of inner retinal neurons, synapse with host retinal neurons, and respond to light. Using an assay for transposase-accessible chromatin with high-throughput sequencing (ATAC–seq), we show that the histone deacetylase inhibitor promotes accessibility at key gene loci in the MG, and allows more effective reprogramming. Our results thus provide a new approach for the treatment of blinding retinal diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Fischer, A. J. & Reh, T. A. Müller glia are a potential source of neural regeneration in the postnatal chicken retina. Nat. Neurosci. 4, 247–252 (2001)
Raymond, P. A., Barthel, L. K., Bernardos, R. L. & Perkowski, J. J. Molecular characterization of retinal stem cells and their niches in adult zebrafish. BMC Dev. Biol. 6, 36 (2006)
Fausett, B. V. & Goldman, D. A role for alpha1 tubulin-expressing Müller glia in regeneration of the injured zebrafish retina. J. Neurosci. 26, 6303–6313 (2006)
Wan, J. & Goldman, D. Retina regeneration in zebrafish. Curr. Opin. Genet. Dev. 40, 41–47 (2016)
Thummel, R., Kassen, S. C., Montgomery, J. E., Enright, J. M. & Hyde, D. R. Inhibition of Müller glial cell division blocks regeneration of the light-damaged zebrafish retina. Dev. Neurobiol. 68, 392–408 (2008)
Lenkowski, J. R. & Raymond, P. A. Müller glia: stem cells for generation and regeneration of retinal neurons in teleost fish. Prog. Retin. Eye Res. 40, 94–123 (2014)
Fausett, B. V., Gumerson, J. D. & Goldman, D. The proneural basic helix-loop-helix gene ascl1a is required for retina regeneration. J. Neurosci. 28, 1109–1117 (2008)
Ramachandran, R., Fausett, B. V. & Goldman, D. Ascl1a regulates Müller glia dedifferentiation and retinal regeneration through a Lin-28-dependent, let-7 microRNA signalling pathway. Nat. Cell Biol. 12, 1101–1107 (2010)
Karl, M. O. & Reh, T. A. Regenerative medicine for retinal diseases: activating endogenous repair mechanisms. Trends Mol. Med. 16, 193–202 (2010)
Pollak, J. et al. ASCL1 reprograms mouse Muller glia into neurogenic retinal progenitors. Development 140, 2619–2631 (2013)
Ueki, Y. et al. Transgenic expression of the proneural transcription factor Ascl1 in Müller glia stimulates retinal regeneration in young mice. Proc. Natl Acad. Sci. USA 112, 13717–13722 (2015)
Biedermann, B., Bringmann, A. & Reichenbach, A. High-affinity GABA uptake in retinal glial (Müller) cells of the guinea pig: electrophysiological characterization, immunohistochemical localization, and modeling of efficiency. Glia 39, 217–228 (2002)
Newman, E. A. Physiological properties and possible functions of Muller cells. Neurosci. Res. Suppl. 4, S209–S220 (1986)
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010)
Samuel, A., Housset, M., Fant, B. & Lamonerie, T. Otx2 ChIP–seq reveals unique and redundant functions in the mature mouse retina. PLoS One 9, e89110 (2014)
Brzezinski, J. A., IV, Kim, E. J., Johnson, J. E. & Reh, T. A. Ascl1 expression defines a subpopulation of lineage-restricted progenitors in the mammalian retina. Development 138, 3519–3531 (2011)
Guo, Z. et al. In vivo direct reprogramming of reactive glial cells into functional neurons after brain injury and in an Alzheimer’s disease model. Cell Stem Cell 14, 188–202 (2014)
Liu, Y. et al. Ascl1 converts dorsal midbrain astrocytes into functional neurons in vivo. J. Neurosci. 35, 9336–9355 (2015)
Niu, W. et al. SOX2 reprograms resident astrocytes into neural progenitors in the adult brain. Stem Cell Reports 4, 780–794 (2015)
Torper, O. et al. In vivo reprogramming of striatal NG2 glia into functional neurons that integrate into local host circuitry. Cell Reports 12, 474–481 (2015)
Srivastava, D. & DeWitt, N. In vivo cellular reprogramming: the next generation. Cell 166, 1386–1396 (2016)
Götz, M., Sirko, S., Beckers, J. & Irmler, M. Reactive astrocytes as neural stem or progenitor cells: in vivo lineage, in vitro potential, and genome-wide expression analysis. Glia 63, 1452–1468 (2015)
Heinrich, C., Spagnoli, F. M. & Berninger, B. In vivo reprogramming for tissue repair. Nat. Cell Biol. 17, 204–211 (2015)
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014)
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013)
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012)
Valensisi, C., Liao, J. L., Andrus, C., Battle, S. L. & Hawkins, R. D. cChIP–seq: a robust small-scale method for investigation of histone modifications. BMC Genomics 16, 1083 (2015)
Dunn, F. A., Doan, T., Sampath, A. P. & Rieke, F. Controlling the gain of rod-mediated signals in the mammalian retina. J. Neurosci. 26, 3959–3970 (2006)
Murphy, G. J. & Rieke, F. Network variability limits stimulus-evoked spike timing precision in retinal ganglion cells. Neuron 52, 511–524 (2006)
Bishop, D. et al. Near-infrared branding efficiently correlates light and electron microscopy. Nat. Methods 8, 568–570 (2011)
Della Santina, L. et al. Glutamatergic monopolar interneurons provide a novel pathway of excitation in the mouse retina. Curr. Biol. 26, 2070–2077 (2016)
Kerschensteiner, D., Morgan, J. L., Parker, E. D., Lewis, R. M. & Wong, R. O. Neurotransmission selectively regulates synapse formation in parallel circuits in vivo. Nature 460, 1016–1020 (2009)
Acknowledgements
The authors acknowledge the following funding sources for support of this work. NIH NEI 1R01EY021482 to T.A.R., EY14358 to R.O.W., Howard Hughes Medical Institute to F.R., Allen Distinguished Investigator award (Paul G. Allen Family Foundation) to T.A.R., F.R. and R.O.W., an NSF Fellowship to M.W. (DGE-0718124), a Cellular and Molecular Biology Training Grant (T32GM007270) to L.S.V., and the Vision Core Grant P30EY01730 for use of the imaging facilities. We thank members of the Reh and Bermingham-McDonogh laboratories for valuable discussion and technical advice. We thank O. Bermingham-McDonogh for constructive comments on the manuscript. We thank A. Wills and A. Chitsazan for their ATAC–seq protocol and data analytics suggestions. We thank J. Delrow, C. Bennett and A. Berger at the Fred Hutch Genomics Shared Resource and Flow Cytometry facilities for their contributions to our sequencing datasets. We thank the laboratory of R. D. Hawkins for the cChIP reagents and protocol (H3K27ac ChIP). We thank the laboratory of C. Trapnell, specifically D. Jackson and R. Chawla, for their help generating the 10× Genomics single-cell RNA-seq datasets. Lastly, the authors thank E. Levine (Vanderbilt University) for the Rlbp1-creER mouse line and M. Nakafuku (Cincinnati Children's) for the tetO-Ascl1-GFP mice.
Author information
Authors and Affiliations
Contributions
N.L.J. and T.A.R. designed and conceived the experiments. N.L.J. performed all injections, immunohistochemistry, imaging, western blots and cell reconstructions. T.Y. and R.O.W. prepared samples for SBFSEM and provided guidance for cell reconstructions and synaptic staining. W.N.G. and F.R. performed all electrophysiological recordings. S.G.W. performed FACS on retinas for ChIP and single-cell mRNA-seq. M.S.W. designed and conceived the experiments and performed Otx2 ChIP–seq and analysis. ATAC–seq and H3K27ac ChIP data was generated by L.S.V. and analysed by L.S.V. and T.A.R. All other figures and text were prepared by N.L.J. and T.A.R. with feedback and contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks Z. He, H. Song and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Animal age and numbers used in study.
a, Histogram showing ages at which all Glast-creER and Rlbp1-creER mice used for the study received their first tamoxifen injection. There were 108 Glast-creER and Rlbp1-creER mice used for the study (122.7 ± 64.1 days, mean postnatal age ± s.d.). b, Histogram showing ages at which all ANT-treated mice used for the study received their first tamoxifen injection. There were 54 ANT-treated mice used for the study (136.4 ± 57.8 days, mean postnatal age ± s.d.). ANT-treated mice are classified according to experimental assay (H3K27ac ChIP-PCR; RNA-seq (10x Genomics)/H3K27ac ChIP-PCR, cells from these mice were split for both assays; RNA-seq (Fluidigm), single-cell RNA-seq; Otx2 ChIP–seq; IHC, immunohistochemistry; SBFSEM, serial block-face scanning electron microscopy; Ephys, whole-cell electrophysiology; ATAC–seq).
Extended Data Figure 2 TSA increases histone acetylation.
a, Western blot for H3K27ac and H3 24 h after TSA or DMSO intravitreal injection. b, Graph shows TSA significantly increases H3K27ac relative to H3 by unpaired t-test at **P = 0.0033, n = 5 per group. c, Graph showing the percentage of GFP+ cells that express Otx2 in various conditions in Rlbp1-creER animals. Numbers were similar to Glast-creER animals, showing no Otx2 in control mice lacking Ascl1 and approximately 36% of GFP+ cells expressing Otx2 when all three factors were present. Data were analysed by one-way ANOVA with Tukey’s post hoc test. d, Graph showing the percentage of GFP+ cells that express Sox9 in various conditions in Glast-creER and Rlbp1-creER animals. A significant reduction in the number of cells expressing Sox9 was seen by one-way ANOVA with Tukey’s post hoc test at *P < 0.05, **P < 0.01, ***P < 0.001 when animals expressing Ascl1 were treated with NMDA and TSA. Data in b–d are shown as mean ± standard error. e, Graph showing the percentage of ANT-treated GFP+ cells that express Otx2 is not significantly different with age by Student’s t-test (n = 4 and 11 mice for 7 weeks and 15–21 weeks, respectively). Data in e is shown as mean ± s.d.
Extended Data Figure 3 Neurogenesis and proliferation from Ascl1-expressing MG.
a, Panel of images showing morphologies between MG and neurons. Images are from retinas that were collected 2–5 weeks post-TSA. Examples of ‘hybrid’ morphologies were observed at all post-TSA time points analysed. First image of panel has normal glial morphology for comparison. b, Example of EdU-labelled GFP+Otx2+ MG in ONL; cells stained with all three markers were very rare: <1% of GFP+ MG (approximately 1 cell per retinal section from 3 mice). c, Diagram showing experimental treatment paradigm for proliferation analysis. d, Ascl1-overexpressing mice treated with mouse epidermal growth factor (mEGF) and NMDA showed examples of Sox9+EdU+ MG. e, Majority of MG did not label with EdU when Ascl1-overexpressing mice only received intravitreal EdU injections (No Treatment). f, Graph showing the percentage of GFP+ cells expressing Sox9 and that were labelled with EdU. A significant increase in Sox9+EdU+ MG was seen when treated with NMDA and mEGF by Student’s t-test at **P = 0.0033, n = 5 per group. g, Ascl1-overexpressing mice treated with mEGF and NMDA showed examples of Otx2+EdU+ MG. h, Ascl1-overexpressing mice only receiving intravitreal EdU injections (No Treatment) had no MG that expressed Otx2 and labelled with EdU. i, Graph showing the percentage of GFP+ cells that express Otx2 and were labelled with EdU. A significant increase in Otx2+EdU+ MG was seen when treated with NMDA and mEGF by Student’s t test at *P = 0.0105, n = 5 per group. All data are shown as mean ± standard error, all scale bars are 10 μm.
Extended Data Figure 4 MG-derived neurons form synaptic specializations in IPL.
a, Confocal images of MG-derived bipolar-like cells with processes in IPL (scale bars, 10 μm). b, Super-resolution Zeiss Airyscan raw images of white boxes from a showing IPL processes stained with presynaptic ribbon marker CtBP2 (Ribeye, white) and postsynaptic marker PSD95 (magenta). c, Images from b masked in Amira to show presynaptic CtBP2 within GFP processes. d, Images of masked presynaptic CtBP2 staining apposed to postsynaptic PSD95 staining. e, Images of masks that were traced in Amira around GFP processes in IPL (scale bars, 10 μm). f, Images of masked CtBP2 staining that is within GFP processes. g, Images of CtBP2 superposed with PSD95 staining from c (white arrows) rotated in xz plane (scale bars, 1 μm). Images are from retinas that were collected 11 days post-TSA.
Extended Data Figure 5 MG-derived GFP+ cells exhibit diverse morphologies.
Panel of images showing 2D projections of GFP+ cells imaged post-recording (scale bars, 20 μm).
Extended Data Figure 6 FACS purification of GFP+ cells.
a, b, Live imaging of Ascl1-overexpressing retinas either untreated (a) or ANT-treated (b) before dissociation and FACS purification. c, Representative graph shows gating (P1 gate = viable cells) for viable cells from non-viable cells and debris. d, Gating P1 fraction for GFP fluorescence (P3 gate = GFP+ fraction). e, f, Post-sort on P3 fraction shows majority of GFP+ cells (88.7%) are viable after sorting.
Extended Data Figure 7 ANT treatment results in MG-derived bipolar/amacrine-like cell cluster.
a, Principal component analyses of 10x Genomics single-cell RNA-seq from aggregate wild-type and ANT-treated cells showing k-means clustering. b, Clusters identifying wild-type (brown, 701 cells) and ANT-treated (green, 825 cells) cells. c, Plots showing expression levels of glial genes Glul and Rlbp1. Expression of glial genes is reduced in the ANT-treated cells and further reduced in the MG-derived bipolar/amacrine-like cells. d, Plots showing expression levels of progenitor genes Ascl1 and Dll1. Ascl1 expression was only seen in ANT-treated cells. Dll1 appears to be increased in the large ANT-treated cluster but decreased in the bipolar/amacrine-like cluster. e, Plots showing expression levels of bipolar genes Otx2 and Cabp5. Both genes are enriched in the bipolar/amacrine-like cluster. f, Plots showing expression levels for amacrine genes Prox1 and Neurod4. Both genes are enriched in the bipolar/amacrine-like cluster. g, Plots showing expression levels for rod genes Rho and Nrl. Rod contamination shows enrichment for both genes in small cluster. h, Heat map of top 20 upregulated and differentially expressed genes for each cluster (scale, log2 fold change). Single-cell data are from retinas that were collected from ANT-treated mice 19 days post-TSA.
Extended Data Figure 8 Epigenetic changes in MG-derived neurons.
a, Violin plots of ATAC–seq tag density in the region 1 kb upstream of the promoters of genes that either decrease by twofold or increase by twofold between the wild-type and ANT-reprogrammed MG to show that the changes in gene expression are accompanied by changes in DNA accessibility during reprogramming with Ascl1, NMDA and TSA. The changes in gene expression were taken from the average of all Ascl1+ cells in the single-cell RNA-seq data from the Fluidigm data and compared with the wild-type MG from the same dataset. Notably, many progenitor genes, like Hes6 and Ascl1, show chromatin accessibility at putative cis-regulatory regions in all treatment conditions (including wild type) even though these genes are not normally expressed in the mature retina. b, H3K27ac ChIP was carried out on FACS-purified ANT-reprogrammed MG and wild-type MG. The data are shown as a per cent of control for three neural genes that show increases in expression and a control gene (Hoxb2) that does not change with ANT treatment. There was a substantial increase in H3K27ac for each of the genes tested. ChIP-PCR data are from ANT-treated retinas that were collected 19 days post-TSA.
Extended Data Figure 9 FACS-purified ANT-treated cells show increased neuronal gene expression and Otx2 binding.
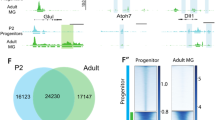
a, Principal component analysis from Fluidigm single-cell mRNA-seq showing 48 wild-type MG (black), 9 ANT-treated cells (red), and 1 cell that was probably a contaminating rod photoreceptor (asterisk). The ANT-treated cells formed a separate cluster from the wild-type MG. b, Heat map showing ANT-treated MG-derived cells and wild-type MG. Each column is an individual cell, with example neural and glial genes as rows. Reprogramming with ANT leads to downregulation of glial genes and an increase in neural genes. Scale is log2(CPM). ANT-treated retinas were collected 46 days post-TSA. c, IGV browser tracks at bipolar genes Crx, Cplx4 and Cabp5 showing: Otx2 ChIP–seq from whole adult mouse retina15 (grey); Otx2 ChIP–seq from FACS-purified ANT-treated and Ascl1-only cells (blue); and DNA accessibility from ATAC–seq of FACS-purified ANT-treated cells (red). Scale for all tracks shown to the right as reads per million. Black arrows mark open chromatin in promoter regions that are bound by Otx2 in ANT-treated but not control Ascl1 cells, and are appropriate regions of binding based on a whole-retina Otx2 ChIP–seq dataset. d, e, Otx2 ChIP–seq peaks from ANT-treated cells show central enrichment at Otx2 ChIP–seq sites from whole adult retina15, but this is not present in the Otx2 ChIP–seq from MG that expressed Ascl1 but received none of the other treatments (TSA, NMDA). f, Top-scoring transcription factor binding motif enrichment (HOMER software suite) from ANT-treated Otx2 ChIP–seq peak calls along with the closest matching transcription factor motif (TF match) and P value for de novo motif enrichment. g, Gene Ontology enrichment for category ‘Mouse Phenotype’ of peaks from d, showing P value for term enrichment, the number of associated genes (blue bars), and example genes from these enriched terms (expanded at right). ChIP–seq was performed on 490,430 and 692,271 pooled cells from 17 ANT-treated Rlbp1-Ascl1 and 15 untreated Rlbp1-Ascl1 mice, respectively. Retinas were collected 9–18 days post-TSA.
Supplementary information
Rights and permissions
About this article
Cite this article
Jorstad, N., Wilken, M., Grimes, W. et al. Stimulation of functional neuronal regeneration from Müller glia in adult mice. Nature 548, 103–107 (2017). https://doi.org/10.1038/nature23283
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature23283
This article is cited by
-
Identifying Hmga2 preserving visual function by promoting a shift of Müller glia cell fate in mice with acute retinal injury
Stem Cell Research & Therapy (2024)
-
Exosomes derived from induced cardiopulmonary progenitor cells alleviate acute lung injury in mice
Acta Pharmacologica Sinica (2024)
-
Epithelial–mesenchymal transition in tissue repair and degeneration
Nature Reviews Molecular Cell Biology (2024)
-
Regulation of chromatin organization during animal regeneration
Cell Regeneration (2023)
-
Common and divergent gene regulatory networks control injury-induced and developmental neurogenesis in zebrafish retina
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.