Abstract
Inflammatory bowel diseases are chronic gastrointestinal inflammatory disorders that affect millions of people worldwide. Genome-wide association studies have identified 200 inflammatory bowel disease-associated loci, but few have been conclusively resolved to specific functional variants. Here we report fine-mapping of 94 inflammatory bowel disease loci using high-density genotyping in 67,852 individuals. We pinpoint 18 associations to a single causal variant with greater than 95% certainty, and an additional 27 associations to a single variant with greater than 50% certainty. These 45 variants are significantly enriched for protein-coding changes (n = 13), direct disruption of transcription-factor binding sites (n = 3), and tissue-specific epigenetic marks (n = 10), with the last category showing enrichment in specific immune cells among associations stronger in Crohn’s disease and in gut mucosa among associations stronger in ulcerative colitis. The results of this study suggest that high-resolution fine-mapping in large samples can convert many discoveries from genome-wide association studies into statistically convincing causal variants, providing a powerful substrate for experimental elucidation of disease mechanisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
12 July 2017
The equation at the end Methods section ‘Establishing a P value threshold’ was corrected.
References
Kappelman, M. D. et al. Direct health care costs of Crohn’s disease and ulcerative colitis in US children and adults. Gastroenterology 135, 1907–1913 (2008)
Molodecky, N. A. et al. Increasing incidence and prevalence of the inflammatory bowel diseases with time, based on systematic review. Gastroenterology 142, 46–54.e42 (2012)
Jostins, L. et al. Host–microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012)
Liu, J. Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015)
van de Bunt, M., Cortes, A., Brown, M. A., Morris, A. P. & McCarthy, M. I. Evaluating the performance of fine-mapping strategies at common variant GWAS loci. PLoS Genet. 11, e1005535 (2015)
Maller, J. B. et al. Bayesian refinement of association signals for 14 loci in 3 common diseases. Nat. Genet. 44, 1294–1301 (2012)
Yang, J. et al. FTO genotype is associated with phenotypic variability of body mass index. Nature 490, 267–272 (2012)
Beecham, A. H. et al. Analysis of immune-related loci identifies 48 new susceptibility variants for multiple sclerosis. Nat. Genet. 45, 1353–1360 (2013)
Onengut-Gumuscu, S. et al. Fine mapping of type 1 diabetes susceptibility loci and evidence for colocalization of causal variants with lymphoid gene enhancers. Nat. Genet. 47, 381–386 (2015)
The 1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012)
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010)
Jostins, L. Using Next-Generation Genomic Datasets in Disease Association. PhD thesis, Univ. Cambridge (2012)
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009)
Howie, B., Marchini, J. & Stephens, M. Genotype imputation with thousands of genomes. G3 1, 457–470 (2011)
Goyette, P. et al. High-density mapping of the MHC identifies a shared role for HLA-DRB101:03 in inflammatory bowel diseases and heterozygous advantage in ulcerative colitis. Nat. Genet. 47, 172–179 (2015)
Rivas, M. A. et al. Deep resequencing of GWAS loci identifies independent rare variants associated with inflammatory bowel disease. Nat. Genet. 43, 1066–1073 (2011)
Yang, J. et al. Genome partitioning of genetic variation for complex traits using common SNPs. Nat. Genet. 43, 519–525 (2011)
Huang, H., Chanda, P., Alonso, A., Bader, J. S. & Arking, D. E. Gene-based tests of association. PLoS Genet. 7, e1002177 (2011)
Momozawa, Y. et al. Resequencing of positional candidates identifies low frequency IL23R coding variants protecting against inflammatory bowel disease. Nat. Genet. 43, 43–47 (2011)
Kheradpour, P. & Kellis, M. Systematic discovery and characterization of regulatory motifs in ENCODE TF binding experiments. Nucleic Acids Res. 42, 2976–2987 (2014)
Nechanitzky, R. et al. Transcription factor EBF1 is essential for the maintenance of B cell identity and prevention of alternative fates in committed cells. Nat. Immunol. 14, 867–875 (2013)
Trynka, G. et al. Chromatin marks identify critical cell types for fine mapping complex trait variants. Nat. Genet. 45, 124–130 (2013)
Farh, K. K.-H. et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature 518, 337–343 (2015)
Bernstein, B. E. et al. The NIH Roadmap Epigenomics Mapping Consortium. Nat. Biotechnol. 28, 1045–1048 (2010)
Nica, A. C. et al. Candidate causal regulatory effects by integration of expression QTLs with complex trait genetic associations. PLoS Genet. 6, e1000895 (2010)
Lappalainen, T. et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501, 506–511 (2013)
Wallace, C. et al. Statistical colocalization of monocyte gene expression and genetic risk variants for type 1 diabetes. Hum. Mol. Genet. 21, 2815–2824 (2012)
Dubois, P. C. A. et al. Multiple common variants for celiac disease influencing immune gene expression. Nat. Genet. 42, 295–302 (2010)
Wright, F. A. et al. Heritability and genomics of gene expression in peripheral blood. Nat. Genet. 46, 430–437 (2014)
Westra, H.-J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013)
Fairfax, B. P. et al. Innate immune activity conditions the effect of regulatory variants upon monocyte gene expression. Science 343, 1246949 (2014)
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014)
Franke, A. et al. Genome-wide meta-analysis increases to 71 the number of confirmed Crohn’s disease susceptibility loci. Nat. Genet. 42, 1118–1125 (2010)
Nejentsev, S., Walker, N., Riches, D., Egholm, M. & Todd, J. A. Rare variants of IFIH1, a gene implicated in antiviral responses, protect against type 1 diabetes. Science 324, 387–389 (2009)
Huang, J., Ellinghaus, D., Franke, A., Howie, B. & Li, Y. 1000 Genomes-based imputation identifies novel and refined associations for the Wellcome Trust Case Control Consortium phase 1 data. Eur. J. Hum. Genet. 20, 801–805 (2012)
Spain, S. L. & Barrett, J. C. Strategies for fine-mapping complex traits. Hum. Mol. Genet. 24 (R1), R111–R119 (2015)
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015)
The UK10K Consortium. The UK10K project identifies rare variants in health and disease. Nature 526, 82–90 (2015)
Roadmap Epigenomics Consortium et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015)
Shah, T. S. et al. optiCall: a robust genotype-calling algorithm for rare, low-frequency and common variants. 28, 1598–1603 (2012)
Price, A. L. et al. Long-range LD can confound genome scans in admixed populations. Am. J. Hum. Genet. 83, 132–135 (2008)
Anderson, E . et al. LAPACK Users’ Guide (Society for Industrial and Applied Mathematics, 1999)
Yang, J. et al. Genomic inflation factors under polygenic inheritance. Eur. J. Hum. Genet. 19, 807–812 (2011)
Bulik-Sullivan, B. K. et al. LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015)
de Lange, K. M. et al. Genome-wide association study implicates immune activation of multiple integrin genes in inflammatory bowel disease. Nat. Genet. 49, 256–261 (2017)
Delaneau, O., Marchini, J. & Zagury, J.-F. A linear complexity phasing method for thousands of genomes. Nat. Methods 9, 179–181 (2011)
Delaneau, O., Zagury, J.-F. & Marchini, J. Improved whole-chromosome phasing for disease and population genetic studies. Nat. Methods 10, 5–6 (2013)
Morris, J. A., Randall, J. C., Maller, J. B. & Barrett, J. C. Evoker: a visualization tool for genotype intensity data. Bioinformatics 26, 1786–1787 (2010)
Jostins, L. & McVean, G. Trinculo: Bayesian and frequentist multinomial logistic regression for genome-wide association studies of multi-category phenotypes. Bioinformatics 32, 1898–1900 (2016)
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011)
Madsen, P ., Su, G ., Labouriau, R . & Christensen, F. DMU-a package for analyzing multivariate mixed models. In Proc. 9th World Congress on Genetics Applied to Livestock Production 137 (Gesellschaft für Tierzuchtwissenschaften 2010), p. 137
Cox, D. R . & Snell, E. J. Analysis of Binary Data 2nd edn, Ch. 2 (CRC, 1989)
D’haeseleer, P. What are DNA sequence motifs? Nat. Biotechnol 24, 423–425 (2006)
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010)
The International HapMap Consortium. The International HapMap Project. Nature 426, 789–796 (2003)
Lin, S. M., Du, P., Huber, W. & Kibbe, W. A. Model-based variance-stabilizing transformation for Illumina microarray data. Nucleic Acids Res. 36, e11 (2008)
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003)
Du, P., Kibbe, W. A. & Lin, S. M. lumi: a pipeline for processing Illumina microarray. Bioinformatics 24, 1547–1548 (2008)
Storey, J. D. A direct approach to false discovery rates. J. Roy. Stat. Soc. B 64, 479–498 (2002)
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014)
Acknowledgements
We thank M. Khan and B. Wong for their assistance in designing illustrations, and K. de Lange for comments on the Supplementary Methods. We received support from the following grants. M.J.D. and R.J.X.: P30DK43351, U01DK062432, R01DK64869, Helmsley grant 2015PG-IBD001 and Crohn’s & Colitis Foundation of America. G.T., C.A.A. and J.C.B: Wellcome Trust grant 098051. M.G.: Fonds de la Recherche Scientifique-FNRS for the FRFS-WELBIO under grant no. WELBIO-CR-2012A-06 (CAUSIBD), BELSPO-IUAP-P7/43-BeMGI, Fédération Wallonie-Bruxelles (ARC IBD@Ulg), and Région Wallonne (CIBLES, FEDER). H.H.: ASHG/Charles J. Epstein Trainee Award. J.L.: Wellcome Trust 098759/Z/12/Z. D.M.: Olle Engkvist Foundation and Swedish Research Council (grants 2010-2976 and 2013-3862). R.K.W.: VIDI grant (016.136.308) from the Netherlands Organization for Scientific Research. J.D.R.: Canada Research Chair, National Institute of Diabetes and Digestive and Kidney Diseases grants DK064869 and DK062432, CIHR GPG-102170 from the Canadian Institutes of Health Research, GPH-129341 from Genome Canada and Génome Québec, and Crohn’s Colitis Canada. J.H.C.: DK062429, DK062422, DK092235, DK106593, and the Sanford J. Grossman Charitable Trust. R.H.D.: Inflammatory Bowel Disease Genetic Research Chair at the University of Pittsburgh, U01DK062420 and R01CA141743. E.D.: Marie-Curie Fellowship. A.-S.G: Fonds de la Recherche Scientifique-FNRS (F.R.S.-FNRS) and Fonds Léon Fredericq fellowships. J.H.: Örebro University Hospital Research Foundation and the Swedish Research Council (grant number 521 2011 2764). C.G.M. and M.P.: National Institute for Health Research (NIHR) Biomedical Research Centre awards to Guy’s & St Thomas’ NHS Trust/King’s College London and to Addenbrooke’s Hospital/University of Cambridge School of Clinical Medicine. D.E.: German Federal Ministry of Education and Research (SysInflame grant 01ZX1306A), DFG Excellence Cluster number 306 ‘Inflammation at Interfaces’. A.F.: Professor of Foundation for Experimental Medicine (Zurich, Switzerland). D.P.B.M.: DK062413, AI067068, U54DE023789-01, 305479 from the European Union, and The Leona M. and Harry B. Helmsley Charitable Trust. Additional acknowledgements for the original data are in the Supplementary Information.
Author information
Authors and Affiliations
Consortia
Contributions
Overall project supervision and management: M.J.D. J.C.B., M.G. Fine-mapping algorithms: H.H., M.F., L.J. TFBS analyses: H.H., K.F. Epigenetic analyses: M.U.M., G.T. eQTL dataset generation: E.L., E.T., J.D., E.D., M.E., R.M., M.M., Y.M., V.D., A.G. eQTL analyses: M.F., J.D., L.J., A.C. Variance component analysis: T.M., M.F. Contribution to overall statistical analyses: G.B. Primary drafting of the manuscript: M.J.D., J.C.B, M.G., H.H., L.J. Major contribution to drafting of the manuscript: M.F., M.U.M., J.H.C., D.P.B.M., J.D.R., C.G.M., R.H.D., R.K.W. The remaining authors contributed to the study conception, design, genotyping quality control, and/or writing of the manuscript. All authors saw, had the opportunity to comment on, and approved the final draft.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks A. Morris, C. Polychronakos, L. Scott, and T. Vyse for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Lists of participants and their affiliations appear in the Supplementary Information.
Extended data figures and tables
Extended Data Figure 1 Power of the fine-mapping analysis.
Power (y axis) to identify the causal variant in a correlated pair (strength of correlation shown by colour) increases with the significance of the association (x axis), and therefore with sample size and effect size. The vertical dashed line shows the genome-wide significance level. To estimate the relationship between the strength of association and our ability to fine-map it, we assume that the association has only two causal variant candidates, and we define the signal as successfully fine-mapped if the ratio of Bayes factors between the true causal variant and the non-causal variant was greater than 10 (a 91% posterior, assuming equal priors for the two candidate variants). Using equation (8) in Supplementary Methods, we have  in which θ is the maximum likelihood estimate of the parameter values. The log-likelihood ratio follows a χ2 distribution:
in which θ is the maximum likelihood estimate of the parameter values. The log-likelihood ratio follows a χ2 distribution: in which λ is the χ2 statistic of the lead variant and r is the correlation coefficient between the two variants. Because of the additive property of the χ2 distribution, logBF follows a non-central χ2 distribution with 1 degree of freedom and non-centrality parameter λ(1 − r2)/2. Therefore, the power can calculated as the probability that logBF > log(10), given by the cumulative distribution function of the non-central χ2 distribution.
in which λ is the χ2 statistic of the lead variant and r is the correlation coefficient between the two variants. Because of the additive property of the χ2 distribution, logBF follows a non-central χ2 distribution with 1 degree of freedom and non-centrality parameter λ(1 − r2)/2. Therefore, the power can calculated as the probability that logBF > log(10), given by the cumulative distribution function of the non-central χ2 distribution.
Extended Data Figure 2 Procedures in the fine-mapping analysis.
Details for each stage are described in Methods. The dashed line means the imputation was performed only once after the manual inspection (not iteratively).
Extended Data Figure 3 Variance explained.
Variance explained by secondary, tertiary, … variants as a fraction of the primary signal at each locus.
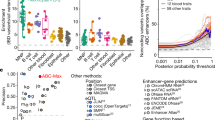
Extended Data Figure 4 Functional annotations.
a, Functional annotation for 45 variants having posterior probability > 50%. b, Functional annotation for 116 association signals fine-mapped to ≤50 variants. Annotations are defined in Methods. We additionally grouped eQTLs into ‘Immune/Blood’ (CD4+, CD8+, CD19+, CD14+ CD15+, platelets) and ‘Gut’ (ileum, transverse colon, and rectum). The eQTLs were generated from the ULg dataset using the ‘Frequentist co-localization using conditional P values’ approach (Methods).
Extended Data Figure 5 Size of credible sets.
Comparison of credible set sizes for primary signals using each of our fine-mapping methods (methods 1, 2, and 3), the combined approach (as adopted in final results) and the approach described in ref. 6 (y axis) and the r2 > 0.6 cut-off (x axis). Fine-mapping maps most signals to smaller numbers of variants. The trend line (blue) and the confidence interval (grey) were calculated using the geom_smooth function in the R ggplot2 package using the linear model.
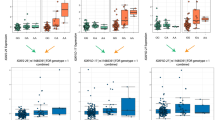
Extended Data Figure 6 Distributions of the allele frequency and the imputation quality.
a–c, Distribution of the risk allele frequency for 45 variants having >50% posterior probability plotted against (a) posterior probability, (b) significance of the association as –log10(P), and (c) odds ratio of the association. Variants are colour coded according to their functions. Odds ratio for IBD associations was the larger of odds ratios for Crohn’s disease and ulcerative colitis. d–f, Distribution of imputation quality (INFO measure from the IMPUTE2 program) for variants having MAF ≥ 5% (d), between 5% and 1% (e), and < 1% (f).
Extended Data Figure 7 Merging and adjudicating signals across methods.
The number of signals for each method is shown in the brackets, and for each method a black bar indicates a signal with P < 1.35 × 10−6, and a grey bar a signal that does not reach that threshold. The coloured bar shows the final status of each signal after merging and model selection (Methods). Label ‘low INFO’ corresponds to INFO < 0.8 (the threshold used for signals reported by one or two methods), and ‘rare and imputed’ to MAF < 0.01 and no genotyped variants in the credible set, regardless of INFO (Methods).
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Notes, a Supplementary Box, the full list of members of the International Inflammatory Bowel Disease Genetics Consortium and acknowledgements for the original data. (PDF 1246 kb)
Supplementary Table 1
This table contains a list of all fine-mapped signals, a list of all variants in fine-mapped signals and Functional annotation for all fine-mapped signals. (XLSX 974 kb)
Supplementary Table 2
This table shows enrichment for histone marks in various cell lines. (XLSX 77 kb)
Supplementary Table 3
This table shows test of heterogeneity between the balanced and imbalanced cohorts. (XLSX 75 kb)
Rights and permissions
About this article
Cite this article
Huang, H., Fang, M., Jostins, L. et al. Fine-mapping inflammatory bowel disease loci to single-variant resolution. Nature 547, 173–178 (2017). https://doi.org/10.1038/nature22969
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22969
This article is cited by
-
Ensemble learning for integrative prediction of genetic values with genomic variants
BMC Bioinformatics (2024)
-
Transcriptome-wide association studies associated with Crohn’s disease: challenges and perspectives
Cell & Bioscience (2024)
-
Neutrophils: from IBD to the gut microbiota
Nature Reviews Gastroenterology & Hepatology (2024)
-
Pleiotropy, epistasis and the genetic architecture of quantitative traits
Nature Reviews Genetics (2024)
-
Phenotypic and genetic effect of carotid intima-media thickness on the risk of stroke
Human Genetics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.