Abstract
Cpf1 is an RNA-guided endonuclease that is emerging as a powerful genome-editing tool1,2. Here we provide insight into its DNA-targeting mechanism by determining the structure of Francisella novicida Cpf1 with the triple-stranded R-loop generated after DNA cleavage. The structure reveals the machinery involved in DNA unwinding to form a CRISPR RNA (crRNA)–DNA hybrid and a displaced DNA strand. The protospacer adjacent motif (PAM) is recognized by the PAM-interacting domain. The loop-lysine helix–loop motif in this domain contains three conserved lysine residues that are inserted in a dentate manner into the double-stranded DNA. Unzipping of the double-stranded DNA occurs in a cleft arranged by acidic and hydrophobic residues facilitating the crRNA–DNA hybrid formation. The PAM single-stranded DNA is funnelled towards the nuclease site through a mixed hydrophobic and basic cavity. In this catalytic conformation, the PAM-interacting domain and the helix–loop–helix motif in the REC1 domain adopt a ‘rail’ shape and ‘flap-on’ conformations, respectively, channelling the PAM strand into the cavity. A steric barrier between the RuvC-II and REC1 domains forms the ‘septum’, separating the displaced PAM strand and the crRNA–DNA hybrid, avoiding DNA re-annealing. Mutations in key residues reveal a mechanism linking the PAM and DNA nuclease sites. Analysis of the Cpf1 structures3,4 proposes a singular working model of RNA-guided DNA cleavage, suggesting new avenues for redesign of Cpf1.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Vestergaard, G., Garrett, R. A. & Shah, S. A. CRISPR adaptive immune systems of Archaea. RNA Biol. 11, 156–167 (2014)
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR–Cas system. Cell 163, 759–771 (2015)
Dong, D. et al. The crystal structure of Cpf1 in complex with CRISPR RNA. Nature 532, 522–526 (2016)
Yamano, T. et al. Crystal structure of Cpf1 in complex with guide RNA and target DNA. Cell 165, 949–962 (2016)
Marraffini, L. A. CRISPR–Cas immunity in prokaryotes. Nature 526, 55–61 (2015)
Mohanraju, P. et al. Diverse evolutionary roots and mechanistic variations of the CRISPR–Cas systems. Science 353, aad5147 (2016)
Mojica, F. J., Díez-Villaseñor, C., García-Martínez, J. & Soria, E. Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J. Mol. Evol. 60, 174–182 (2005)
Makarova, K. S. et al. An updated evolutionary classification of CRISPR–Cas systems. Nat. Rev. Microbiol. 13, 722–736 (2015)
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016)
Horvath, P. & Barrangou, R. CRISPR/Cas, the immune system of bacteria and archaea. Science 327, 167–170 (2010)
Sorek, R., Lawrence, C. M. & Wiedenheft, B. CRISPR-mediated adaptive immune systems in bacteria and archaea. Annu. Rev. Biochem. 82, 237–266 (2013)
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013)
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012)
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013)
Makarova, K. S. & Koonin, E. V. Annotation and classification of CRISPR–Cas systems. Methods Mol. Biol. 1311, 47–75 (2015)
Deltcheva, E. et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471, 602–607 (2011)
Fonfara, I., Richter, H., Bratovicˇ, M., Le Rhun, A. & Charpentier, E. The CRISPR-associated DNA-cleaving enzyme Cpf1 also processes precursor CRISPR RNA. Nature 532, 517–521 (2016)
Fagerlund, R. D., Staals, R. H. & Fineran, P. C. The Cpf1 CRISPR–Cas protein expands genome-editing tools. Genome Biol. 16, 251 (2015)
Kim, Y. et al. Generation of knockout mice by Cpf1-mediated gene targeting. Nat. Biotechnol. 34, 808–810 (2016)
Kleinstiver, B. P. et al. Genome-wide specificities of CRISPR–Cas Cpf1 nucleases in human cells. Nat. Biotechnol. 34, 869–874 (2016)
Endo, A., Masafumi, M., Kaya, H. & Toki, S. Efficient targeted mutagenesis of rice and tobacco genomes using Cpf1 from Francisella novicida. Sci. Rep. 6, 38169 (2016)
Jiang, F. et al. Structures of a CRISPR–Cas9 R-loop complex primed for DNA cleavage. Science 351, 867–871 (2016)
Gao, P., Yang, H., Rajashankar, K. R., Huang, Z. & Patel, D. J. Type V CRISPR–Cas Cpf1 endonuclease employs a unique mechanism for crRNA-mediated target DNA recognition. Cell Res. 26, 901–913 (2016)
Anders, C., Niewoehner, O., Duerst, A. & Jinek, M. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 513, 569–573 (2014)
Jiang, F., Zhou, K., Ma, L., Gressel, S. & Doudna, J. A. Structural biology. A Cas9-guide RNA complex preorganized for target DNA recognition. Science 348, 1477–1481 (2015)
Szczelkun, M. D. et al. Direct observation of R-loop formation by single RNA-guided Cas9 and Cascade effector complexes. Proc. Natl Acad. Sci. USA 111, 9798–9803 (2014)
Yang, H., Gao, P., Rajashankar, K. R. & Patel, D. J. PAM-dependent target DNA recognition and cleavage by C2c1 CRISPR–Cas endonuclease. Cell 167, 1814–1828 (2016)
Kim, D. et al. Genome-wide analysis reveals specificities of Cpf1 endonucleases in human cells. Nat. Biotechnol. 34, 863–868 (2016)
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016)
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015)
Sternberg, S. H., Redding, S., Jinek, M., Greene, E. C. & Doudna, J. A. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature 507, 62–67 (2014)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D 69, 1204–1214 (2013)
Vonrhein, C. et al. Data processing and analysis with the autoPROC toolbox. Acta Crystallogr. D 67, 293–302 (2011)
Sheldrick, G. M. A short history of SHELX. Acta Crystallogr. A 64, 112–122 (2008)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Adams, P. D. et al. The Phenix software for automated determination of macromolecular structures. Methods 55, 94–106 (2011)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Kleywegt, G. J. & Jones, T. A. Model building and refinement practice. Methods Enzymol. 277, 208–230 (1997)
Bricogne G. et al. BUSTER version 2.10.3 (2016)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Diederichs, K. & Karplus, P. A. Better models by discarding data? Acta Crystallogr. D 69, 1215–1222 (2013)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Acknowledgements
The Novo Nordisk Foundation Center for Protein Research is supported financially by the Novo Nordisk Foundation (Grant NNF14CC0001). This work was also supported by a grant from the Danish Cancer Society to G.M. We thank the beamline staff of X06SA at the Swiss Light Source (PSI, Switzerland) and the EMBL beamline P14 at DESY (Germany) for support during data collection. Data processing has been performed in the Computerome, the Danish National Computer for Life Sciences. We also thank H. Koc and M. Melynis from the Protein Production Facility Platform at CPR for excellent technical assistance.
Author information
Authors and Affiliations
Contributions
S.S. expressed and purified the Cpf1–R-loop complex with the help of P.A. S.S. and P.A. prepared the crystals and P.A. performed the cleavage and saturation experiments designed by S.S. All authors collected diffraction data. G.M. performed data processing and crystallographic analysis. All authors were involved in model building and refinement. All authors discussed the data, and helped G.M. to write the manuscript and prepare the figures.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks J. Doudna, D. Liu and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Sequence alignment of representative members of the Cpf1 family.
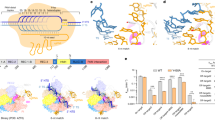
The amino acid sequences of Cpf1 from Francisella novicida (FnCpf1), Acidaminococcus sp. (AsCpf1), Lachnospiraceae bacterium (LbCpf1), Francisella tularensis (FtCpf1) and Porphyromonas crevioricans (PcCpf1) were aligned by Clustal Omega (http://www.ebi.ac.uk/Tools/msa/clustalo). The figure was prepared with ESPript (http://espript.ibcp.fr). Residue numbers are labelled according to the FnCpf1 sequence. Similar residues are shown in red and identical residues in white over red background. Blue frames indicate homologous regions. The different structural domains of FnCpf1 and their amino acid composition are depicted as boxes above the sequences and labelled with the same names and colour code as in Fig. 1a. In addition, thin boxes above the domains, following the same colour code, mark the positions of relevant regions mentioned in the manuscript. The mutations performed in key residues are marked according to their region: PI rail conformation (square), catalytic (circle), unzipping cleft (triangle), LKL (star) and ‘septum’ (cross). The protein sequence identity and similarity (in percentage) of FnCpf1 (1,300 amino acids) with AsCpf1, LbCpf1, FtCpf1 and PcCpf1 are FnCpf1:AsCpf1, 36:42; FnCpf1:LbCpf1, 43:49; FnCpf1:FtCpf1, 99:100; and FnCpf1:PcCpf1, 43:52.
Extended Data Figure 2 DNA saturation cleavage assay.
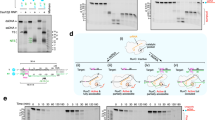
a, Scheme of the oligonucleotides used in the assay. In DNA1 both strands are labelled with 6-FAM in order to visualize the cleaved products of both non-target and target DNA. In DNA2 the non-target strand is labelled with 6-FAM in order to visualize the cleaved products from the non-target DNA. In DNA3 the target strand is labelled with 6-FAM to visualize the cleaved products from the target strand. b, Target DNA cleavage depends on Mg2+. Depletion of the cation by EDTA abrogates phosphodiester hydrolysis. c, Time course of the cleavage assay shows that FnCpf1 endonuclease completes the reaction in approximately 60 min. Quantification is shown in the graph below, representing the intensity of the corresponding bands (AU, arbitrary units) at different time points. d, Cleavage assay using DNA1, DNA2 and DNA3, shows the cleavage products of the different strands. For the saturation assay an increasing amount of DNA1 was used. Quantification of the cleaved and non-cleaved DNA1 is shown in the chart below (see Methods). The chart represents the intensity of the corresponding bands (AU) at increasing concentrations of the DNA1 substrate. The activity curve shows an asymptotic behaviour after 3 nmol of substrate. Each experiment was repeated 4 times. Error bars, s.d.
Extended Data Figure 3 The crRNA–DNA hybrid and the displaced non-target strand.
a, Detailed view of the 2mFo − DFc map of the R-loop inside FnCpf1 contoured at 1σ. b, Superposition of the FnCpf1–R-loop with the crRNA–DNA hybrid containing the PAM sequence of AsCpf1.
Extended Data Figure 4 The RNAase site and the LKL helix insertion.
a, Inset of the 5′ RNA region. Cpf1 can process its own crRNA. The RNA catalysis involves residues F873, H843, K869 and K852. 2mFo−DFc sigmaA map contoured at 1σ is displayed on the RNA molecule. The dashed lines indicate the flexible loop not built in the model. H843 is included in this highly mobile six residues loop and cannot be observed in our structure, showing the high flexibility of this region after RNA processing. The alanine mutants of these residues abolished RNA nuclease activity17. b, Overall view of the sliced FnCpf1 from the RuvC–Nuc interface, the key residues in each domain are surrounded by a pink and cyan oval. The dashed lines depict the distances between E1006 and R1218, and the dT–7 of the nt-strand and the dG+20 the t-strand to the RuvC and Nuc centres, respectively. The lower panel shows a detailed view of the RuvC–Nuc interface where the catalysis of the nt-strand and the t-strand occur after DNA unzipping. c, Cleavage assay of the E1006A and R1218A FnCpf1 mutants. Both mutations abrogate DNA cleavage. The experiment was repeated 4 times. d, Detailed view of the LKL helix insertion in the dsDNA. The dashed red lines show the 45° between the longitudinal DNA axis and the helix axis.
Extended Data Figure 5 Cleavage assay of FnCpf1 Mutants.
a, SDS–PAGE gel of the purified proteins used in the assay. b, Substrate target DNA oligonucleotide used in this assay. In DNA1A both strands are labelled with 6FAM in order to visualize the cleaved products of both the non-target and the target DNA strands in Figs 2e, 3b and Extended Data Fig. 5c. The encircled region indicates the PAM sequence c, Wild-type FnCpf1 and K671A, K667A and K677A mutants cleavage with different PAM sequences. The PAMs are indicated in bold above the gels. No cleavage can be observed for the wild-type FnCpf1 or mutants with PAMs different than the TTA. Neither FnCpf1 nor the LKL mutants exhibit PAM promiscuity in this context. Each experiment was repeated 4 times.
Extended Data Figure 6 Electrostatic potential map of FnCpf1.
a, The close-up of the unzipping cleft shows the disposition of the hydrophobic and acidic amino acids to facilitate the formation of the initial hybridization between the target DNA and the crRNA. b, c, Two different views of the cavity where the displaced nt-DNA resides after target cleavage. The cavity consists of hydrophobic and basic residues that direct the nt-DNA to the DNA nuclease site. The PAM, and DNA nuclease site and the septum are encircled in a dashed yellow line.
Extended Data Figure 7 PAM-interacting domain conformation.
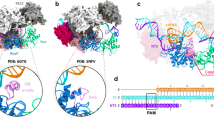
a, The PI domains of LbCpf1, AsCpf1 and FnCpf1 structure have been superimposed using the Lsq routine in O36 to observe the different conformations depending on the presence or absence of the non-target DNA strand. The dashed black line represents the path of the nt-DNA. b, View of the non-target PAM strand embraced by the ‘rail’ conformation of the PI domain. The AsCpf1 PI domain is superimposed for comparison. The electron density map (2mFo − DFc map) displayed on the non-target strand inside FnCpf1 is contoured at 1σ.
Extended Data Figure 8 Comparison of Cpf1 conformations upon target DNA binding.
a, The figure displays surface models after superimposition of LbCpf1, AsCpf1 and FnCpf1 conformations representative of different states. The comparison shows the differences in the Cpf1 structure from the less to the more compact conformation. Nucleic acids are not shown for clarity. b, The figure depicts the differences in the HLH section between the LbCpf1 apo structure and the FnCpf1–R-loop structure. The HLH domain adopts the ‘flap-on’ conformation when the PAM sequence is bound.
Supplementary information
Supplementary Figure
This file contains uncropped scans of all the UREA-PAGE gels in the main and in the Extended Data Figures. (PDF 8001 kb)
Overview of the FnCpf1 R-loop crystal structure
The protein-crRNA-target DNA ternary complex is rotated 180º around its longitudinal axis. The polypeptide is composed by the WED (I, II and II subdomains), REC1, REC2, PI, BH, RuvC (I, II and III subdomains) and Nuc domains. The colours assigned to FnCpf1 domains are the same in Fig. 1a. (MPG 16085 kb)
Morphing between the apo and R-loop structures reflecting conformational change for the HLH motif in the REC1 domain between these two states
The video shows the morphing between the apo and R-loop structures reflecting conformational change for the HLH motif in the REC1 domain between these two states (Flap-on conformation). (MOV 3930 kb)
Rights and permissions
About this article
Cite this article
Stella, S., Alcón, P. & Montoya, G. Structure of the Cpf1 endonuclease R-loop complex after target DNA cleavage. Nature 546, 559–563 (2017). https://doi.org/10.1038/nature22398
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22398
This article is cited by
-
An alpha-helical lid guides the target DNA toward catalysis in CRISPR-Cas12a
Nature Communications (2024)
-
Targeting miRNA by CRISPR/Cas in cancer: advantages and challenges
Military Medical Research (2023)
-
CRISPR applications in cancer diagnosis and treatment
Cellular & Molecular Biology Letters (2023)
-
CRISPR/Cas9 therapeutics: progress and prospects
Signal Transduction and Targeted Therapy (2023)
-
TnpB structure reveals minimal functional core of Cas12 nuclease family
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.