Abstract
In the absence of specialized immune cells, the need for plants to reprogram transcription to transition from growth-related activities to defence is well understood1,2. However, little is known about translational changes that occur during immune induction. Using ribosome footprinting, here we perform global translatome profiling on Arabidopsis exposed to the microbe-associated molecular pattern elf18. We find that during this pattern-triggered immunity, translation is tightly regulated and poorly correlated with transcription. Identification of genes with altered translational efficiency leads to the discovery of novel regulators of this immune response. Further investigation of these genes shows that messenger RNA sequence features are major determinants of the observed translational efficiency changes. In the 5′ leader sequences of transcripts with increased translational efficiency, we find a highly enriched messenger RNA consensus sequence, R-motif, consisting of mostly purines. We show that R-motif regulates translation in response to pattern-triggered immunity induction through interaction with poly(A)-binding proteins. Therefore, this study provides not only strong evidence, but also a molecular mechanism, for global translational reprogramming during pattern-triggered immunity in plants.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Pajerowska-Mukhtar, K. M. et al. The HSF-like transcription factor TBF1 is a major molecular switch for plant growth-to-defense transition. Curr. Biol. 22, 103–112 (2012)
Huot, B., Yao, J., Montgomery, B. L. & He, S. Y. Growth-defense tradeoffs in plants: a balancing act to optimize fitness. Mol. Plant 7, 1267–1287 (2014)
Couto, D. & Zipfel, C. Regulation of pattern recognition receptor signalling in plants. Nature Rev. Immunol. 16, 537–552 (2016)
Wu, S., Shan, L. & He, P. Microbial signature-triggered plant defense responses and early signaling mechanisms. Plant Sci. 228, 118–126 (2014)
Dunbar, T. L., Yan, Z., Balla, K. M., Smelkinson, M. G. & Troemel, E. R. C. elegans detects pathogen-induced translational inhibition to activate immune signaling. Cell Host Microbe 11, 375–386 (2012)
Zipfel, C. et al. Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation. Cell 125, 749–760 (2006)
Lei, L. et al. Ribosome profiling reveals dynamic translational landscape in maize seedlings under drought stress. Plant J. 84, 1206–1218 (2015)
Merchante, C. et al. Gene-specific translation regulation mediated by the hormone-signaling molecule EIN2. Cell 163, 684–697 (2015)
Liu, M. J. et al. Translational landscape of photomorphogenic Arabidopsis. Plant Cell 25, 3699–3710 (2013)
Juntawong, P., Girke, T., Bazin, J. & Bailey-Serres, J. Translational dynamics revealed by genome-wide profiling of ribosome footprints in Arabidopsis. Proc. Natl Acad. Sci. USA 111, E203–E212 (2014)
Lukoszek, R., Feist, P. & Ignatova, Z. Insights into the adaptive response of Arabidopsis thaliana to prolonged thermal stress by ribosomal profiling and RNA-seq. BMC Plant Biol. 16, 221 (2016)
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. S. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009)
Zipfel, C. Combined roles of ethylene and endogenous peptides in regulating plant immunity and growth. Proc. Natl Acad. Sci. USA 110, 5748–5749 (2013)
von Arnim, A. G., Jia, Q. & Vaughn, J. N. Regulation of plant translation by upstream open reading frames. Plant Sci. 214, 1–12 (2014)
Barbosa, C., Peixeiro, I. & Romão, L. Gene expression regulation by upstream open reading frames and human disease. PLoS Genet. 9, e1003529 (2013)
Hinnebusch, A. G., Ivanov, I. P. & Sonenberg, N. Translational control by 5′-untranslated regions of eukaryotic mRNAs. Science 352, 1413–1416 (2016)
Patel, G. P., Ma, S. & Bag, J. The autoregulatory translational control element of poly(A)-binding protein mRNA forms a heteromeric ribonucleoprotein complex. Nucleic Acids Res. 33, 7074–7089 (2005)
Dufresne, P. J., Ubalijoro, E., Fortin, M. G. & Laliberté, J. F. Arabidopsis thaliana class II poly(A)-binding proteins are required for efficient multiplication of turnip mosaic virus. J. Gen. Virol. 89, 2339–2348 (2008)
Gallie, D. R. The role of the poly(A) binding protein in the assembly of the Cap-binding complex during translation initiation in plants. Translation 2, e959378 (2014)
Hinnebusch, A. G. Translational regulation of GCN4 and the general amino acid control of yeast. Annu. Rev. Microbiol. 59, 407–450 (2005)
Gilbert, W. V., Zhou, K., Butler, T. K. & Doudna, J. A. Cap-independent translation is required for starvation-induced differentiation in yeast. Science 317, 1224–1227 (2007)
Nakagawa, T. et al. Development of series of gateway binary vectors, pGWBs, for realizing efficient construction of fusion genes for plant transformation. J. Biosci. Bioeng. 104, 34–41 (2007)
Curtis, M. D. & Grossniklaus, U. A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol. 133, 462–469 (2003)
Xu, G. et al. One-step, zero-background ligation-independent cloning intron-containing hairpin RNA constructs for RNAi in plants. New Phytol. 187, 240–250 (2010)
Li, J. T. et al. Modification of vectors for functional genomic analysis in plants. Genet. Mol. Res. 13, 7815–7825 (2014)
Alonso, J. M. et al. Five components of the ethylene-response pathway identified in a screen for weak ethylene-insensitive mutants in Arabidopsis. Proc. Natl Acad. Sci. USA 100, 2992–2997 (2003)
Hua, J. et al. EIN4 and ERS2 are members of the putative ethylene receptor gene family in Arabidopsis. Plant Cell 10, 1321–1332 (1998)
Stepanova, A. N., Hoyt, J. M., Hamilton, A. A. & Alonso, J. M. A link between ethylene and auxin uncovered by the characterization of two root-specific ethylene-insensitive mutants in Arabidopsis. Plant Cell 17, 2230–2242 (2005)
Galon, Y. et al. Calmodulin-binding transcription activator 1 mediates auxin signaling and responds to stresses in Arabidopsis. Planta 232, 165–178 (2010)
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998)
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nature Protocols 7, 1534–1550 (2012)
Mustroph, A., Juntawong, P. & Bailey-Serres, J. Isolation of plant polysomal mRNA by differential centrifugation and ribosome immunopurification methods. Methods Mol. Biol. 553, 109–126 (2009)
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nature Methods 9, 357–359 (2012)
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015)
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014)
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–8 (2009)
Nicol, J. W., Helt, G. A., Blanchard, S. G., Jr, Raja, A. & Loraine, A. E. The Integrated Genome Browser: free software for distribution and exploration of genome-scale datasets. Bioinformatics 25, 2730–2731 (2009)
Miettinen, T. P. & Björklund, M. Modified ribosome profiling reveals high abundance of ribosome protected mRNA fragments derived from 3′ untranslated regions. Nucleic Acids Res. 43, 1019–1034 (2015)
Acknowledgements
This study was supported by grants from National Institutes of Health 5R01 GM069594-11 and the Howard Hughes Medical Institute and the Gordon and Betty Moore Foundation (through grant GBMF3032) to X. Dong. We thank J. M. Alonso for ein4-1, wei7-4, and ers1-10 seeds; T. Girke, M. Hummel, and J. Bailey-Serres for providing the Ribo-Seq workflow package for data analyses; W. Wang, P. Y. Hsu, and P. N. Benfey for discussing the protocol; R. Zavaliev for the callose staining method; and the Arabidopsis Information Resource for gcn2, erf7, eicbp.b, pab2/4, and pab2/8 seeds. We thank P. Zwack and S. Zebell for comments on the manuscript.
Author information
Authors and Affiliations
Contributions
G.X. and X.D. designed the research. G.X., H.Y. and J. Marqués optimized the footprinting protocol. H.Y. and G.X. performed ethylene-related and polysome profiling experiments. L.L. and G.X. generated the reporter lines. G.X. and J. Motley performed eIF2α phosphorylation assay. G.X. performed the rest of the experiments. G.G. and G.X. performed the bioinformatic analyses and prepared the figures. G.X. and X.D. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
A patent based on this study has been filed by Duke University with G.X., G.G. and X.D. as inventors.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Translational activities during elf18-induced PTI.
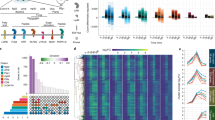
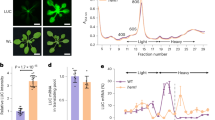
a, Schematic of the 35S:uORFsTBF1–LUC reporter, which is a fusion between the TBF1 exon1 (uORF1/2 and sequence of the N-terminal 73 amino acids) and the firefly luciferase gene (LUC) expressed constitutively by the CaMV 35S promoter. R, R-motif. b–e, Translation (b, d) and transcript levels (c, e) of the 35S:uORFsTBF1–LUC reporter in WT after Mock or elf18 treatment (b, c) or in WT and efr-1 upon elf18 treatment (d, e). LUC activity, mean ± s.e.m. (b, n = 12; d, n = 9) after normalization to time 0; transcript levels, mean of fold changes normalized to time 0 with individual biological replicates shown as solid circles (n = 3); hpi, hours post infiltration. f–i, Polysome profiling of global translational activity (f, h) and TBF1 mRNA translational activity calculated as ratios of polysomal/total mRNA (g, i) in WT after Mock or elf18 treatment (f, g) or in WT and efr-1 upon elf18 treatment (h, i). Lower-case letters indicate polysomal fractions in polysome profiling indicated by sucrose gradient absorbance (A254 nm). Expression levels of TBF1 were normalized against UBQ5 level determined by RT–PCR in total mRNA and in polysomal fractions respectively. Data are shown as the relative TBF1 mRNA level in polysomal fractions after normalization to total TBF1 mRNA level. Bar with solid circles, mean with individual biological replicates (n = 3). j, Schematic of RNA-seq (RS) and Ribo-seq (RF) library construction using uORFsTBF1–LUC/WT plants. RNase I and alkaline are two methods of generating RNA fragments.
Extended Data Figure 2 Improvement made in the library construction protocol.
a, Addition of 5′ deadenylase and RecJf to remove excess 5′ pre-adenylylated linker. mRNA fragments of RNA-seq and Ribo-seq were size-selected and dephosphorylated by PNK treatment, followed by 5′ pre-adenylylated linker ligation. The original method used gel purification to remove the excess linker. In the new method (pink background), 5′ deadenylase was used to remove pre-adenylylated group (Ap) from the unligated linker allowing cleavage by RecJf. The resulting sample could then be used directly for reverse transcription. b, The original (Original) and new (New) methods to remove excess linker were compared. Synthetic RNA markers of 26 and 34 nucleotides (nt) were used for linker ligation. RNA markers without the linker were used as controls. Arrow indicates the excess linkers. DNA ladder, 10 base pairs (bp). c, Reverse transcription (RT) showed the improvement of the new method over the original one. Half of the ligation mixture (O) was gel purified to remove excess linkers before reverse transcription (loaded twice). The other half (N) was treated with 5′ deadenylase and RecJf, and directly used as template for reverse transcription (loaded once). Reverse transcription primers were loaded as control. Arrow indicates excess reverse transcription primers.
Extended Data Figure 3 Quality and reproducibility of RNA-seq and Ribo-seq libraries.
Related to Fig. 1. a, BioAnalyzer profile showed high quality of RNA-seq (RS) and Ribo-seq (RF) libraries. In addition to internal standards (35 bp and 10,380 bp), a single ~170 bp peak is present for RNA-seq and Ribo-seq libraries for Mock and elf18 treatments with both biological replicates (Rep1/2). b, Length distribution of total reads from four RNA-seq and four Ribo-seq libraries. c, Fraction of 30-nucleotide reads in total reads from four RNA-seq and four Ribo-seq libraries. Bar with solid circles, mean with individual biological replicates (n = 4) of percentage of reads with 5′ aligning to A (frame1), U (frame2), and G (frame3) of the initiation codon. d, Read density along 5′ UTR, CDS, and 3′ UTR of total reads from four RNA-seq and four Ribo-seq libraries. Expressed genes with RPKM in CDS ≥ 1 and length of UTR ≥ 1 nucleotide were used for box plots. The top, middle, and bottom lines of the box indicate the 25th, 50th, and 75th percentiles, respectively. Filled circles represent RPKM values for individual outlier genes. e, Nucleotide resolution of the coverage around start and stop codons using the 15th nucleotide of 30-nucleotide reads of Ribo-seq. Reads in 3′ UTR may be due to digestion conditions that might favour the capture of ribosomes in different conformations associated with UTRs as previously observed10 and explained38. f, Correlation between two replicates (Rep1/2) of RNA-seq and Ribo-seq samples. Data are shown as the correlation of log2(RPKM) in CDS for expressed genes with RPKM in CDS ≥ 1. Pearson correlation coefficient r is shown. g, h, Hierarchical clustering showing the reproducibility between RNA-seq (g) and Ribo-seq (h) within two replicates (Rep1/2). Darker colour means greater correlation.
Extended Data Figure 4 Global analyses of transcriptome, translatome, and translational efficiency upon elf18 treatment.
Related to Fig. 1. a, Flowchart for read processing and assignment. b, Reads after each processing. c, Statistical methods and criteria for transcriptome (RSfc), translatome (RFfc), and translational efficiency fold-change analyses. d, GO term enrichment analysis for RNA-seq upregulated genes. e, Normal distribution of log2(translational efficiency) for Mock and elf18 treatment. f, Translational efficiency changes in the endogenous TBF1 gene. Read coverage was normalized to uniquely mapped reads with IGB. Translational efficiencies for the TBF1 exon 2 in Mock and elf18 treatments were determined to calculate translational efficiency fold change. g, GO term enrichment found in TEup genes in response to elf18 treatment. A z score ≥1.5 was used. h, Correlation between translational efficiency fold change and exon length, 5′ UTR length, 3′ UTR length, and guanine–cytosine (GC) composition. TE, translational efficiency.
Extended Data Figure 5 Characterization of novel PTI regulators, related to Fig. 1f.
a, RNA-seq and translational efficiency changes in known or homologues of known components of the ethylene- and the damage-associated molecular pattern Pep-mediated PTI signalling pathways (top) and normalized distribution of RNA-seq and Ribo-seq reads of one example (that is, EIN4; bottom). The pathway was modified from ref. 13. In rectangular boxes: black, RNA-seq-changed; red, TEup; blue, TEdn. b, MAPK activation. Twelve-day-old ein4-1, eicbp.b, and erf7 seedlings were treated with 1 μM elf18 solution and collected at the indicated time points for immunoblot analysis using the phosphospecific antibody against MAPK3 and MAPK6. See Supplementary Text for gel source data. c, Callose deposition. Three-week-old plants were infiltrated with 1 μM elf18 or Mock. Leaves were stained 20 h later in aniline blue followed by confocal microscopy. Representative of five images. Scale bar, 100 μm. d, Schematic of the dual-LUC system. Test, 5′ leader sequence (including UTR) or 3′ UTR of the gene tested; LUC, firefly luciferase; RLUC, renilla luciferase, Ter, terminator. e, Dual-LUC assay of EIN4 UTRs on translational activity upon elf18 treatment in N. benthamiana (n = 4). f, Effects of EIN4 UTRs on ratios of LUC/RLUC mRNA upon elf18 treatment (two experiments with three technical replicates). EV, empty vector. g, EIN4 translational activity upon elf18 treatment calculated as ratios of polysomal/total mRNA (two experiments with three technical replicates). Bar with solid circles, mean with individual biological replicates.
Extended Data Figure 6 uORF–mediated translational control.
a, b, Flowcharts of steps used to identify predicted (a) and translated (b) uORFs. c, Read density of uORF and mORF. For those genes with reads assigning to uORF and with RPKM in its mORF ≥ 1, log2(RPKM) for individual uORFs and mORFs are plotted for Mock and elf18 treatment, respectively. r, Pearson correlation coefficient. d, Definition of mORF/uORF ratio shift between Mock and elf18 treatments. e, Histogram of mORF/uORF shift upon elf18 treatment. The ratio of mORF/uORF for elf18 divided by that for Mock was defined as the shift value. Data are shown as the distribution of the log2 transformation of shift values. uORFs with significant shift determined by z score are coloured and whose numbers are shown. f, Histogram of mORF/uORF shift upon hypoxia stress10. g, Venn diagrams showing overlapping uORFs with significant ribo-shift in responses to elf18 and hypoxia treatments. h, Normalized distribution of RNA-seq and Ribo-seq reads to show ribo-shift of GPS1 (AT2G34630) and GSTU16 (AT1G59700) upon elf18 treatment. Numbers on the right mean log2(mORF/uORF) of Ribo-seq. uORFs are boxed with blue colour.
Extended Data Figure 7 R-motif-mediated translational control in response elf18 induction.
Related to Fig. 2. a, Effects of R-motif containing 5′ leader sequences on basal translational activities after normalization to mRNA (n = 3). b, Effects of R-motif deletions (ΔR) on mRNA abundance (n = 6). c–f, Effects of R-motif deletion and R-motif substitution mutations on basal translation (c, e; n = 4) and mRNA levels (d, f, two experiments with three technical replicates) for IAA18 and BET10 (c, d) and TBF1 (e, f). g, mRNA levels in WT and R-motif deletion mutants with and without elf18 treatment (n = 9). h, Effects of R-motif deletions (ΔR) on translational responsiveness to elf18 measured using the dual-LUC assay (n = 3). i, Effects of GA, G(A)3, G(A)6, and G(A)n repeats on mRNA levels when inserted into 5′ UTR of the reporter in transient assay performed in N. benthamiana (two experiments with three technical replicates). j, k, Effects of R-motif deletion and/or uORF mutations on TBF1 mRNA abundance (j) and transcriptional responsiveness to Mock and elf18 treatments (k); n = 3 after normalization to WT (j) or WT with Mock treatment (k). l, Contributions of R-motif and uORFs to TBF1 translational response to elf18 in transgenic Arabidopsis plants. Numbers 1, 2, and 3 on the x axis represent individual transgenic lines tested (n = 6 after normalization to Mock). Bar with solid circles, mean with individual biological replicates.
Extended Data Figure 8 Effects of PABs on mRNA transcription and PTI-associated phenotypes.
Related to Fig. 3. a, Influence of co-expressing PAB2 on mRNA abundance (n = 9). b, The elf18-induced seedling growth inhibition in WT, efr-1, pab2 pab4 (pab2/4), and pab2 pab8 (pab2/8) (mean ± s.e.m., n = 5). c, MAPK activation in WT, pab2/4, pab2/8, and efr-1 seedlings after elf18 treatment measured by immunoblotting using a phosphospecific antibody against MAPK3 and MAPK6.
Extended Data Figure 9 Roles of GCN2 in PTI in plants.
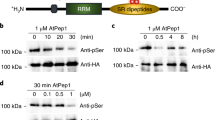
a–d, Effects of the gcn2 mutation on elf18-induced eIF2α phosphorylation (a), translational induction (b, mean ± s.e.m. of LUC activity, n = 8), and transcription of the uORFsTBF1–LUC reporter (c, n = 3; bar with solid circles, mean with individual biological replicates), and resistance to Psm ES4326 (d, mean ± s.e.m., n = 8). See Supplementary Text for gel source data.
Supplementary information
Supplementary Information
This file contains Supplementary Results and a Supplementary Figure showing the uncropped immunoblots. (PDF 206 kb)
Supplementary Table 1
This table shows RSfc, RFfc and TEfc upon elf18 treatment. (XLSX 4100 kb)
Supplementary Table 2
This table contains GO terms for TE-altered genes upon elf18 treatment. (XLSX 15 kb)
Supplementary Table 3
This table contains information regarding the genes involved in the elf18-ethylene-Peps signalling pathway. (XLSX 14 kb)
Supplementary Table 4
This table shows uORF-containing genes. (XLSX 1196 kb)
Supplementary Table 5
This table shows R-motif-containing genes in Arabidopsis, Drosophila, mouse and humans. (XLSX 1212 kb)
Supplementary Table 6
This table contains the plasmids, primers and antibodies used in this study. (XLSX 23 kb)
Rights and permissions
About this article
Cite this article
Xu, G., Greene, G., Yoo, H. et al. Global translational reprogramming is a fundamental layer of immune regulation in plants. Nature 545, 487–490 (2017). https://doi.org/10.1038/nature22371
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22371
This article is cited by
-
N-Acetyltransferase 10 represses Uqcr11 and Uqcrb independently of ac4C modification to promote heart regeneration
Nature Communications (2024)
-
Engineering disease-resistant plants with alternative translation efficiency by switching uORF types through CRISPR
Science China Life Sciences (2024)
-
Changing turn-over rates regulate abundance of tryptophan, GS biosynthesis, IAA transport and photosynthesis proteins in Arabidopsis growth defense transitions
BMC Biology (2023)
-
SPR9 encodes a 60 S ribosomal protein that modulates panicle spreading and affects resistance to false smut in rice (Oryza sativa. L)
BMC Plant Biology (2023)
-
Pervasive downstream RNA hairpins dynamically dictate start-codon selection
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.