Abstract
Protease-activated receptors (PARs) are a family of G-protein-coupled receptors (GPCRs) that are irreversibly activated by proteolytic cleavage of the N terminus, which unmasks a tethered peptide ligand that binds and activates the transmembrane receptor domain, eliciting a cellular cascade in response to inflammatory signals and other stimuli. PARs are implicated in a wide range of diseases, such as cancer and inflammation1,2,3. PARs have been the subject of major pharmaceutical research efforts3 but the discovery of small-molecule antagonists that effectively bind them has proved challenging. The only marketed drug targeting a PAR is vorapaxar4, a selective antagonist of PAR1 used to prevent thrombosis. The structure of PAR1 in complex with vorapaxar has been reported previously5. Despite sequence homology across the PAR isoforms, discovery of PAR2 antagonists has been less successful, although GB88 has been described as a weak antagonist6. Here we report crystal structures of PAR2 in complex with two distinct antagonists and a blocking antibody. The antagonist AZ8838 binds in a fully occluded pocket near the extracellular surface. Functional and binding studies reveal that AZ8838 exhibits slow binding kinetics, which is an attractive feature for a PAR2 antagonist competing against a tethered ligand. Antagonist AZ3451 binds to a remote allosteric site outside the helical bundle. We propose that antagonist binding prevents structural rearrangements required for receptor activation and signalling. We also show that a blocking antibody antigen-binding fragment binds to the extracellular surface of PAR2, preventing access of the tethered ligand to the peptide-binding site. These structures provide a basis for the development of selective PAR2 antagonists for a range of therapeutic uses.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Adams, M. N. et al. Structure, function and pathophysiology of protease activated receptors. Pharmacol. Ther. 130, 248–282 (2011)
Nystedt, S., Emilsson, K., Larsson, A. K., Strömbeck, B. & Sundelin, J. Molecular cloning and functional expression of the gene encoding the human proteinase-activated receptor 2. Eur. J. Biochem. 232, 84–89 (1995)
Ramachandran, R., Noorbakhsh, F., Defea, K. & Hollenberg, M. D. Targeting proteinase-activated receptors: therapeutic potential and challenges. Nat. Rev. Drug Discov. 11, 69–86 (2012)
Chackalamannil, S. et al. Discovery of a novel, orally active himbacine-based thrombin receptor antagonist (SCH 530348) with potent antiplatelet activity. J. Med. Chem. 51, 3061–3064 (2008)
Zhang, C. et al. High-resolution crystal structure of human protease-activated receptor 1. Nature 492, 387–392 (2012)
Suen, J. Y. et al. Modulating human proteinase activated receptor 2 with a novel antagonist (GB88) and agonist (GB110). Br. J. Pharmacol . 165, 1413–1423 (2012)
Hanson, M. A. et al. Crystal structure of a lipid G protein-coupled receptor. Science 335, 851–855 (2012)
Chrencik, J. E. et al. Crystal structure of antagonist bound human lysophosphatidic acid receptor 1. Cell 161, 1633–1643 (2015)
Ballesteros, J. A. & Weinstein, H. in Methods in Neurosciences Vol. 25 (ed. Sealfon, S. C. ) Ch. 19 (Elsevier, 1995)
Krissinel, E. & Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D Biol. Crystallogr . 60, 2256–2268 (2004)
Blackhart, B. D. et al. Extracellular mutations of protease-activated receptor-1 result in differential activation by thrombin and thrombin receptor agonist peptide. Mol. Pharmacol. 58, 1178–1187 (2000)
Zhang, D. et al. Two disparate ligand-binding sites in the human P2Y1 receptor. Nature 520, 317–321 (2015)
Jazayeri, A. et al. Extra-helical binding site of a glucagon receptor antagonist. Nature 533, 274–277 (2016)
Venkatakrishnan, A. J. et al. Molecular signatures of G-protein-coupled receptors. Nature 494, 185–194 (2013)
Borrelli, K. W., Vitalis, A., Alcantara, R. & Guallar, V. PELE: protein energy landscape exploration. A novel Monte Carlo based technique. J. Chem. Theory Comput. 1, 1304–1311 (2005)
Edman, K. et al. Ligand binding mechanism in steroid receptors: from conserved plasticity to differential evolutionary constraints. Structure 23, 2280–2290 (2015)
Giblin, P. et al. Fully human antibodies against the protease-activated receptor-2 (PAR-2) with anti-inflammatory activity. Hum. Antibodies 20, 83–94 (2011)
Caffrey, M. & Porter, C. Crystallizing membrane proteins for structure determination using lipidic mesophases. J. Vis. Exp. 45, 1712–1723 (2010)
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010)
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D Biol. Crystallogr . 69, 1204–1214 (2013)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr . 66, 213–221 (2010)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr . 66, 486–501 (2010)
Terwilliger, T. C. et al. Iterative model building, structure refinement and density modification with the PHENIX AutoBuild wizard. Acta Crystallogr. D Biol. Crystallogr . 64, 61–69 (2008)
Bricogne, G. et al. BUSTER version 2.9. Cambridge, United Kingdom: Global Phasing Ltd. (2010)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr . 66, 12–21 (2010)
Carlson, H. A. et al. CSAR 2014: A benchmark exercise using unpublished data from pharma. J. Chem. Inf. Model. 56, 1063–1077 (2016)
Grebner, C. et al. Binding mode and induced fit predictions for prospective computational drug design. J. Chem. Inf. Model. 56, 774–787 (2016)
Acknowledgements
We acknowledge support at AstraZeneca from J. Zhang, F. Wu, T.-M. Dracka, J. Cumming, S. Cowen, M. Duggan and H. Chen for contributions in chemistry, J. Claesson for support with cell cultivation, A. Ferguson for support on discovery of AZ3451; and at Heptares from B. Teobald for chemistry input at early stages of the project, A. Brown, J. Brown and O. Mace for contributions on receptor pharmacology and assay development; and from V. Guallar at the Barcelona Supercomputing Center, Spain for development of the PELE software to enable our studies.
Author information
Authors and Affiliations
Contributions
R.K.Y.C. carried out construct design, protein purification, LCP crystallizations, designed crystallization optimization, harvested crystals, collected diffraction data and solved the small molecule crystal complexes. A.J. and G.W. carried out the conformational thermostabilization. R.P. provided pharmacology profiling of PAR2 variants. O.S. contributed to construct design, and established together with C.F.-V. the protein expression and purification. A.S. performed protein purification. A.Z. developed the Biacore protocols and performed compound profiling, S.G. performed Biacore compound profiling in support of the discovery of AZ3451. L.S. developed assays and performed small molecule pharmacological profiling. N.-O.H. provided cell support and performed radioligand binding assays. P.T. carried out the pharmacological profiling of blocking antibody. A.S.D. and M.R. collected diffraction data and were involved in structure determinations. K.E. and P.J. solved and refined the structure of PAR2 in complex with Fab. B.T. carried out the computational analysis of the structure and ligand binding. C.G. carried out the molecular simulations of ligand exit and entry. B.A. and L.L. performed the sequence analysis of MAB3949 from the hybridoma cell line. D.T., H.S. and C.E.D. discovered the novel chemotype by DNA-encoded library screening leading to AZ3451. G.A.B. managed the chemistry program at Heptares. D.G.B. led the chemistry optimization leading to AZ3451. R.M.C. managed the structural biology at Heptares. Overall project management was carried out by N.D. and F.H.M. The manuscript was prepared by R.K.Y.C., A.S.D., K.E., A.J., F.H.M. and N.D.
Corresponding author
Ethics declarations
Competing interests
R.K.Y.C., C.F.-V., O.S., G.A.B., R.M.C., A.S.D., A.J., R.P., M.R., B.T., G.W., A.Z. and F.H.M. are employees of Heptares Therapeutics limited and are shareholders in Sosei Group Corporation. K.E., D.G.B., S.G., C.G., N.-O.H., P.J., A.S., L.S., P.T. and N.D. are employees and shareholders of AstraZeneca. B.A. and L.L. are employees and shareholders of Bio-techne. C.E.D., H.S. and D.T. are employees and shareholders of X-Chem Inc.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Comparison between PAR2 and PAR1.
a, PAR1 (PDB code 3VW7 in purple) with vorapaxar (yellow) bound was superimposed onto PAR2 (in green) with the positions of AZ8838 (in magenta) and AZ3451 (in orange) indicated. b, c, Differences in the helical trajectories of the extracellular and intracellular side are indicated by red arrows. d, The inward movement of TM5 and TM6 places residues Tyr2425.38, Phe2435.39, His3106.58 and Tyr3116.59 into steric clashes with vorapaxar when superimposed onto PAR1. e, The AZ8838 allosteric pocket is also present in PAR1. His1352.64 in PAR2 is replaced by Tyr1622.64 in PAR1 which would clash with AZ8838.
Extended Data Figure 2 Overview of the crystal asymmetric unit.
a, PAR2 receptor is shown with the location of the StaR mutations (in orange stick representation) and the glycosylation site mutation N222Q (in red stick representation). The location of the sodium ion (in purple sphere) is indicated. b, b562RIL is inserted between residue Leu2695.65 and Glu276ICL3 of ICL3 of PAR2. c, The N-terminal lobe of T4L is not involved in crystal contacts and is less well ordered compared to the C-terminal lobe. d, Electron density for AZ8838; view from extracellular space looking down the receptor TMD bundle. For all panels, density shown (blue mesh) is from a composite omit map calculated using the final model of PAR2 double fusion complexed with AZ8838 and contoured at 1.0σ.
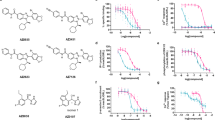
Extended Data Figure 3 Saturation binding of [3H]-radiolabelled antagonist and agonist to membranes from HEK293 cells overexpressing wild-type and StaR variant of human PAR2.
Data analysis using a single-binding-site model gave a Kd of 344 ± 74 nM and 356 ± 72 for [3H]-AZ8838 antagonist to wild-type and StaR, respectively. High-affinity binding of agonist [3H]-acetyl-GB110 was observed to wild-type PAR2 with a Kd of 34 ± 5 nM, whereas no binding could be detected to PAR2-StaR, indicative of an antagonist-stabilized state. Data were fitted to a single-binding-site model and the reported values are the mean ± s.e.m. of two independent experiments, except in b, which is based on three experiments. Error bars represent s.e.m.
Extended Data Figure 4 Inhibition of PAR2 signalling by MAB3949.
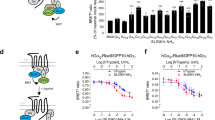
a–d, Blocking activity of MAB3949 or Fab3949 of PAR2 activation by trypsin (a, c) or activating peptide SLIGRL (b, d) measured using FLIPR in human A549 cells. The IC50 of MAB3949 is 286 ± 143 nM for trypsin activation (n = 3) and 253 ± 36.7 nM for activating peptide (n = 2). Isotype controls (red curves) are mouse IgG2A clone NIP228 as full IgG or Fab fragment. Minor non-specific inhibition seen with high concentrations of full-length mouse IgG2A isotype control maybe related to IgG Fc domains as is not apparent with IgG2A Fab. Error bars represent s.e.m. e, f, Representative sensorgrams for interaction of Fab with full-length (e) and NΔ57 (f) PAR2-StaR variants. The mean Kd values determined from four replicate runs are 7.6 ± 1.9 nM and 11 ± 2.7 nM, respectively.
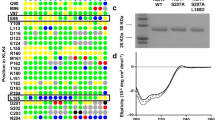
Extended Data Figure 5 Schematic representation of PAR2 constructs used in this study.
Residues are labelled with the sequence number and Ballesteros–Weinstein nomenclature (Ballesteros and Weinstein, 1995)9 . Insertion positions of T4L and b562RIL at the N terminus and ICL3 are indicated. StaR mutations are coloured in green, residues involved in AZ8838 interaction in purple, AZ3451 in blue and the most conserved residue (x.50 in Ballesteros–Weinstein nomenclature) in each transmembrane helix in orange. The disulfide bond Cys1483.25–Cys226ECL2 is indicated with a yellow dotted line. Residues not observed in the crystal structure are coloured in grey. Residues involved in more than one interaction have more than one colour.
Extended Data Figure 6 The triclinic P1 crystal lattice of PAR2-StaR T4L-NΔ57/CΔ20 N222Q b562RIL-C04.
a, b, d, Three orientations of the crystal lattice looking down the bc (a), ac (b) and ab (d) plane. c, Magnified view of the dotted area in b. e, Magnified view of the dotted area in c. f, Magnified view of the dotted area in d. In f, T4L is semi-transparent for clarity.
Supplementary information
Supplementary Information
This file contains Supplementary Text. (PDF 217 kb)
Video of the AZ8838 entry simulations
The video starts with an overview of the X-ray structure of PAR2 (green) in complex with AZ8838 (magenta) with the His227ECL2 highlighted in orange sticks. Then the simulated entry trajectory of AZ8838 (cyan) is shown for accepted Monte Carlo step. The video finally transitions into the refinement trajectory within the allosteric pocket. (MP4 22082 kb)
Rights and permissions
About this article
Cite this article
Cheng, R., Fiez-Vandal, C., Schlenker, O. et al. Structural insight into allosteric modulation of protease-activated receptor 2. Nature 545, 112–115 (2017). https://doi.org/10.1038/nature22309
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature22309
This article is cited by
-
G protein-coupled receptors (GPCRs): advances in structures, mechanisms, and drug discovery
Signal Transduction and Targeted Therapy (2024)
-
Structural insights into the activation and inhibition of CXC chemokine receptor 3
Nature Structural & Molecular Biology (2024)
-
Control of the antitumour activity and specificity of CAR T cells via organic adapters covalently tethering the CAR to tumour cells
Nature Biomedical Engineering (2023)
-
Structural basis of hydroxycarboxylic acid receptor signaling mechanisms through ligand binding
Nature Communications (2023)
-
Small-molecule discovery through DNA-encoded libraries
Nature Reviews Drug Discovery (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.