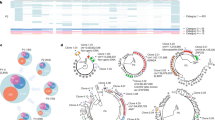
Abstract
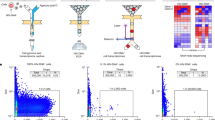
The persistence of the HIV reservoir in infected individuals is a major obstacle to the development of a cure for HIV1,2,3. Here, using an in vitro model of HIV-infected quiescent CD4 T cells, we reveal a gene expression signature of 103 upregulated genes that are specific for latently infected cells, including genes for 16 transmembrane proteins. In vitro screening for surface expression in HIV-infected quiescent CD4 T cells shows that the low-affinity receptor for the immunoglobulin G Fc fragment, CD32a, is the most highly induced, with no detectable expression in bystander cells. Notably, productive HIV-1 infection of T-cell-receptor-stimulated CD4 T cells is not associated with CD32a expression, suggesting that a quiescence-dependent mechanism is required for its induction. Using blood samples from HIV-1-positive participants receiving suppressive antiretroviral therapy, we identify a subpopulation of 0.012% of CD4 T cells that express CD32a and host up to three copies of HIV DNA per cell. This CD32a+ reservoir was highly enriched in inducible replication-competent proviruses and can be predominant in some participants. Our discovery that CD32a+ lymphocytes represent the elusive HIV-1 reservoir may lead to insights that will facilitate the specific targeting and elimination of this reservoir.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Chun, T. W. et al. Presence of an inducible HIV-1 latent reservoir during highly active antiretroviral therapy. Proc. Natl Acad. Sci. USA 94, 13193–13197 (1997)
Chun, T. W., Davey, R. T., Jr, Engel, D., Lane, H. C. & Fauci, A. S. Re-emergence of HIV after stopping therapy. Nature 401, 874–875 (1999)
Finzi, D. et al. Identification of a reservoir for HIV-1 in patients on highly active antiretroviral therapy. Science 278, 1295–1300 (1997)
Zack, J. A. et al. HIV-1 entry into quiescent primary lymphocytes: molecular analysis reveals a labile, latent viral structure. Cell 61, 213–222 (1990)
Stevenson, M., Stanwick, T. L., Dempsey, M. P. & Lamonica, C. A. HIV-1 replication is controlled at the level of T cell activation and proviral integration. EMBO J . 9, 1551–1560 (1990)
Chomont, N. et al. HIV reservoir size and persistence are driven by T cell survival and homeostatic proliferation. Nat. Med . 15, 893–900 (2009)
Laguette, N. et al. SAMHD1 is the dendritic- and myeloid-cell-specific HIV-1 restriction factor counteracted by Vpx. Nature 474, 654–657 (2011)
Berger, A. et al. SAMHD1-deficient CD14+ cells from individuals with Aicardi-Goutières syndrome are highly susceptible to HIV-1 infection. PLoS Pathog . 7, e1002425 (2011)
Hrecka, K. et al. Vpx relieves inhibition of HIV-1 infection of macrophages mediated by the SAMHD1 protein. Nature 474, 658–661 (2011)
Descours, B. et al. SAMHD1 restricts HIV-1 reverse transcription in quiescent CD4+ T-cells. Retrovirology 9, 87 (2012)
Baldauf, H. M. et al. SAMHD1 restricts HIV-1 infection in resting CD4+ T cells. Nat. Med . 18, 1682–1687 (2012)
Cribier, A., Descours, B., Valadão, A. L., Laguette, N. & Benkirane, M. Phosphorylation of SAMHD1 by cyclin A2/CDK1 regulates its restriction activity toward HIV-1. Cell Reports 3, 1036–1043 (2013)
Nimmerjahn, F. & Ravetch, J. V. Fcγ receptors as regulators of immune responses. Nat. Rev. Immunol . 8, 34–47 (2008)
Guilliams, M., Bruhns, P., Saeys, Y., Hammad, H. & Lambrecht, B. N. The function of Fcγ receptors in dendritic cells and macrophages. Nat. Rev. Immunol . 14, 94–108 (2014)
Laird, G. M. et al. Rapid quantification of the latent reservoir for HIV-1 using a viral outgrowth assay. PLoS Pathog . 9, e1003398 (2013)
Ho, Y. C. et al. Replication-competent noninduced proviruses in the latent reservoir increase barrier to HIV-1 cure. Cell 155, 540–551 (2013)
Cohn, L. B. et al. HIV-1 integration landscape during latent and active infection. Cell 160, 420–432 (2015)
Bournazos, S. et al. Broadly neutralizing anti-HIV-1 antibodies require Fc effector functions for in vivo activity. Cell 158, 1243–1253 (2014)
Bruel, T. et al. Elimination of HIV-1-infected cells by broadly neutralizing antibodies. Nat. Commun . 7, 10844 (2016)
Horwitz, J. A. et al. HIV-1 suppression and durable control by combining single broadly neutralizing antibodies and antiretroviral drugs in humanized mice. Proc. Natl Acad. Sci. USA 110, 16538–16543 (2013)
Avettand-Fènoël, V. et al. LTR real-time PCR for HIV-1 DNA quantitation in blood cells for early diagnosis in infants born to seropositive mothers treated in HAART area (ANRS CO 01). J. Med. Virol. 81, 217–223 (2009)
Pereira Bittencourt Passaes, C . et al. Ultrasensitive HIV-1 p24 detects single infected cells and differences in reservoir induction by latency-reversal agents. J. Virol. 91, e.02296-16 (2017)
Rosenbloom, D. I . et al. Designing and interpreting limiting dilution assays: general principles and applications to the latent reservoir for Human Immunodeficiency Virus-1. Open Forum Infect. Dis. 2, ofv123 (2015)
Acknowledgements
We thank the participants enrolled in this study. We thank members of the Molecular Virology laboratory for critical reading of the manuscript, J. Venables for editing the manuscript, MRI platform for FACSAria cell sorting. We are grateful to J.-F. Delfraissy and F. Barré-Sinoussi for their continuous support. This work was supported by grants from the European FP7 contract 305762, MSDAvenir, ANRS and FRM ‘équipe labéllisée’ and Labex EpiGenMed to M.B. G.P was supported by FRM fellowships; B.D. and R.R. by ANRS fellowships. T.B. was supported by a Vaccine Research Institute (VRI) fellowship. O.S was supported by grants from ANRS, VRI, Labex IBEID, Sidaction, European FP7 contract 305762 and Institut Pasteur.
Author information
Authors and Affiliations
Contributions
M.B. conceived the study. M.B., G.P. and B.D. designed experiments, interpreted data and wrote the paper. R.R. analysed the RNA-seq data. T.B. and O.S. performed highly sensitive p24 assays. J.D.L., J.-L.L.-Z., C.L., C.P., J.R. and Y.L. recruited participants and collected blood samples. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks N. Chomont, F. Nimmerjahn, D. Richman, G. Silvestri and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Cellular pathways significantly modulated in latently infected quiescent CD4 T cells compared to bystanders and non-infected cells.
a, b, Biological pathway analysis using REACTOME (a) or STRING (b) databases for the 103 genes differentially expressed in XH+ compared to XH− and non-infected cells.
Extended Data Figure 2 Flow cytometry dot plots and gating strategy for cell sorting of CD32ahi, CD32aint and CD32a− CD4 T lymphocytes subsets from 10 HIV-1 infected participants.
When available similar number of events were displayed in CD32a staining than in isotype control. Note that for patient 566, the cell-sorting strategy was designed by selecting a threshold on CD3 positivity.
Extended Data Figure 3 Contribution of CD32a+ CD4 T cells to the inducible viral reservoir contained in total CD4 T cells.
qVOA was performed using CD32a− and total CD4 T cell isolated from participant 769.
Related audio
Supplementary information
Supplementary Data
This file contains Supplementary Table 1. (XLSX 4517 kb)
Supplementary Data
This file contains Supplementary Table 2. (XLSX 16 kb)
Supplementary Data
This file contains Supplementary Tables 3 and 4. (PDF 166 kb)
Rights and permissions
About this article
Cite this article
Descours, B., Petitjean, G., López-Zaragoza, JL. et al. CD32a is a marker of a CD4 T-cell HIV reservoir harbouring replication-competent proviruses. Nature 543, 564–567 (2017). https://doi.org/10.1038/nature21710
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21710
This article is cited by
-
Targeted shock-and-kill HIV-1 gene therapy approach combining CRISPR activation, suicide gene tBid and retargeted adenovirus delivery
Gene Therapy (2024)
-
Phenotypic characterization of single CD4+ T cells harboring genetically intact and inducible HIV genomes
Nature Communications (2023)
-
The landscape of alternative splicing in HIV-1 infected CD4 T-cells
BMC Medical Genomics (2020)
-
High CD45 expression of CD8+ and CD4+ T cells correlates with the size of HIV-1 reservoir in blood
Scientific Reports (2020)
-
Fc-optimized antibodies elicit CD8 immunity to viral respiratory infection
Nature (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.