Abstract
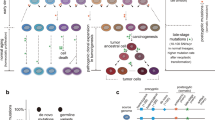
Somatic cells acquire mutations throughout the course of an individual’s life. Mutations occurring early in embryogenesis are often present in a substantial proportion of, but not all, cells in postnatal humans and thus have particular characteristics and effects1. Depending on their location in the genome and the proportion of cells they are present in, these mosaic mutations can cause a wide range of genetic disease syndromes2 and predispose carriers to cancer3,4. They have a high chance of being transmitted to offspring as de novo germline mutations and, in principle, can provide insights into early human embryonic cell lineages and their contributions to adult tissues5. Although it is known that gross chromosomal abnormalities are remarkably common in early human embryos6, our understanding of early embryonic somatic mutations is very limited. Here we use whole-genome sequences of normal blood from 241 adults to identify 163 early embryonic mutations. We estimate that approximately three base substitution mutations occur per cell per cell-doubling event in early human embryogenesis and these are mainly attributable to two known mutational signatures7. We used the mutations to reconstruct developmental lineages of adult cells and demonstrate that the two daughter cells of many early embryonic cell-doubling events contribute asymmetrically to adult blood at an approximately 2:1 ratio. This study therefore provides insights into the mutation rates, mutational processes and developmental outcomes of cell dynamics that operate during early human embryogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Samuels, M. E. & Friedman, J. M. Genetic mosaics and the germ line lineage. Genes 6, 216–237 (2015)
Erickson, R. P. Recent advances in the study of somatic mosaicism and diseases other than cancer. Curr. Opin. Genet. Dev. 26, 73–78 (2014)
Laurie, C. C. et al. Detectable clonal mosaicism from birth to old age and its relationship to cancer. Nat. Genet. 44, 642–650 (2012)
Ruark, E. et al. Mosaic PPM1D mutations are associated with predisposition to breast and ovarian cancer. Nature 493, 406–410 (2013)
Behjati, S. et al. Genome sequencing of normal cells reveals developmental lineages and mutational processes. Nature 513, 422–425 (2014)
Vanneste, E. et al. Chromosome instability is common in human cleavage-stage embryos. Nat. Med. 15, 577–583 (2009)
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013)
Oron, E. & Ivanova, N. Cell fate regulation in early mammalian development. Phys. Biol. 9, 045002 (2012)
Genovese, G. et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N. Engl. J. Med. 371, 2477–2487 (2014)
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371, 2488–2498 (2014)
Xie, M. et al. Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat. Med. 20, 1472–1478 (2014)
Bruce, A. W. & Zernicka-Goetz, M. Developmental control of the early mammalian embryo: competition among heterogeneous cells that biases cell fate. Curr. Opin. Genet. Dev. 20, 485–491 (2010)
Plusa, B. et al. The first cleavage of the mouse zygote predicts the blastocyst axis. Nature 434, 391–395 (2005)
Zernicka-Goetz, M., Morris, S. A. & Bruce, A. W. Making a firm decision: multifaceted regulation of cell fate in the early mouse embryo. Nat. Rev. Genet. 10, 467–477 (2009)
Plachta, N., Bollenbach, T., Pease, S., Fraser, S. E. & Pantazis, P. Oct4 kinetics predict cell lineage patterning in the early mammalian embryo. Nat. Cell Biol. 13, 117–123 (2011)
Bedzhov, I., Graham, S. J., Leung, C. Y. & Zernicka-Goetz, M. Developmental plasticity, cell fate specification and morphogenesis in the early mouse embryo. Phil. Trans. R. Soc. Lond. B 369, 20130538 (2014)
Morris, S. A., Guo, Y. & Zernicka-Goetz, M. Developmental plasticity is bound by pluripotency and the Fgf and Wnt signaling pathways. Cell Reports 2, 756–765 (2012)
Hardy, K., Handyside, A. H. & Winston, R. M. The human blastocyst: cell number, death and allocation during late preimplantation development in vitro . Development 107, 597–604 (1989)
Strnad, P. et al. Inverted light-sheet microscope for imaging mouse pre-implantation development. Nat. Methods 13, 139–142 (2016)
Rahbari, R. et al. Timing, rates and spectra of human germline mutation. Nat. Genet. 48, 126–133 (2016)
Acuna-Hidalgo, R. et al. Post-zygotic point mutations are an underrecognized source of de novo genomic variation. Am. J. Hum. Genet. 97, 67–74 (2015)
Huang, A. Y. et al. Postzygotic single-nucleotide mosaicisms in whole-genome sequences of clinically unremarkable individuals. Cell Res. 24, 1311–1327 (2014)
Dal, G. M. et al. Early postzygotic mutations contribute to de novo variation in a healthy monozygotic twin pair. J. Med. Genet. 51, 455–459 (2014)
Lynch, M. Rate, molecular spectrum, and consequences of human mutation. Proc. Natl Acad. Sci. USA 107, 961–968 (2010)
Martincorena, I. & Campbell, P. J. Somatic mutation in cancer and normal cells. Science 349, 1483–1489 (2015)
Nik-Zainal, S. et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature 534, 47–54 (2016)
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010)
Stephens, P. J. et al. The landscape of cancer genes and mutational processes in breast cancer. Nature 486, 400–404 (2012)
Ju, Y. S. et al. Origins and functional consequences of somatic mitochondrial DNA mutations in human cancer. eLife 3, e02935 (2014)
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012)
Van Loo, P . et al. Allele-specific copy number analysis of tumors. Proc. Natl Acad. Sci. USA 107, 16910–16915 (2010)
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011)
Skinner, M. E., Uzilov, A. V., Stein, L. D., Mungall, C. J. & Holmes, I. H. JBrowse: a next-generation genome browser. Genome Res. 19, 1630–1638 (2009)
Holstege, H. et al. Somatic mutations found in the healthy blood compartment of a 115-yr-old woman demonstrate oligoclonal hematopoiesis. Genome Res. 24, 733–742 (2014)
Marikawa, Y. & Alarcón, V. B. Establishment of trophectoderm and inner cell mass lineages in the mouse embryo. Mol. Reprod. Dev. 76, 1019–1032 (2009)
Laurent, L. et al. Dynamic changes in the human methylome during differentiation. Genome Res. 20, 320–331 (2010)
Acknowledgements
We thank M. Zernicka-Goetz at Gurdon Institute, K. J. Dawson at Wellcome Trust Sanger Institute and T. Bleazard at University of Manchester for discussion and assistance with manuscript preparation. This work was supported by the Wellcome Trust (grant reference 077012/Z/05/Z). Y.S.J. is supported by EMBO long-term fellowship (LTF 1203_2012), by KAIST (G04150052), and by a grant of the Korea Health Technology R&D project through the Korea Health Industry Development Institute (KHIDI) funded by the Ministry of Health & Welfare, Republic of Korea (HI16C2387). P.J.C. is a Wellcome Trust Senior Clinical Fellow. The ICGC Breast Cancer Consortium was supported by a grant from the European Union (BASIS) and the Wellcome Trust. For the family study, Generation Scotland received core support from the Chief Scientist Office of the Scottish Government Health Directorates (CZD/16/6) and the Scottish Funding Council (HR03006).
Author information
Authors and Affiliations
Contributions
M.R.S. designed and directed the project. Y.S.J. performed the overall study with bioinformatics analyses for detection of early embryonic mutations. I.M. and M.G. performed statistical testing to confirm unequal contributions of early cells and early mutation rates. L.B.A. carried out mutational signature analyses. R.R. and M.E.H. designed and directed family studies. D.C.W., H.R.D., M.R. and S.N.-Z. performed cancer genome analyses and provided conceptual advice. M.P., A.F., C.A., N.P., S.G. and S.O. carried out laboratory analyses. S.M. supported clinical data analysis and curation. D.D.G., T.S. and S.E.P. performed pathology review for breast cancer tissues. C.A.P., A.B., H.S., M.v.d.V., B.K.T.T., C.C., A.T., N.T.U., L.J.v.V., J.W.M.M., C.S., S.K., P.N.S., S.R.L., J.E.E., A.-L.B.-D., A.R., A.M.T. and A.V. provided clinical samples and commented on the manuscript. P.J.C. supervised overall analyses. Y.S.J., I.M., M.G., L.B.A. and M.R.S. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks M. Horowitz, S. Orkin and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Filters to exclude mutation candidates in regions with copy number variation.
a, For every blood sample, we assessed the distribution of coverage of around 3 million inherited SNP loci. Using this distribution, we determined a cut-off value that is used for inter-sample CNV filtering (see Methods). In the case of PD3989b shown in the figure, candidate mutation loci with >51× coverage were considered to be located on copy number gain thus removed. b, An example of inter-sample CNV filtering (see Methods). Normalized coverage for chr11:14,446,619 region of PD4116b is located in the normal copy number (CN = 2) cluster. c, Copy number gain was identified in a candidate mutation locus (chr6:285,671) from PD4116b by the inter-sample CNV filtering method. Therefore, this mutation candidate was removed from further downstream analyses.
Extended Data Figure 2 Features of ultrahigh-depth targeted amplicon sequencing used for validation.
a, Estimation of the effect of potential PCR allelic bias from targeted amplicon sequencing. Using inherited heterozygous SNP sites that were PCR amplified and ultra-deep sequenced, we assessed potential PCR bias (that is, preferential amplification of one allele compared to the other): the distribution of VAFs was broader than expected from a binomial distribution (theoretical maximum), but the PCR bias was not substantial as a clear peak at VAF = 0.5 was present. The estimated overdispersion level (theta value in beta-binomial distribution) was 223.88. The estimate was used in the simulation studies for assessment of cell-doubling asymmetry in early embryogenesis (see Methods for more details). b, High precision of ultrahigh-depth amplicon sequencing in assessment of VAF of a mutation. For the 14 early embryonic mutations, we quantified their VAFs from the second blood samples using the same strategy (that is, PCR amplification and deep sequencing). The VAF estimates from the first and the second sequencings were highly correlated. c, Background error rate of targeted amplicon sequencing (see Methods). The background mutation rate showed sequence context dependency. Error bars denote 2× interquartile range. We used these background mutation rates in a filtering step.
Extended Data Figure 3 Features of a blood sample with a neoplastic clonal expansion in the blood.
a, This hypothetical scenario illustrates the expectation in a normal blood sample when there is no obvious neoplastic clonal expansion. Each white-filled black circle represents an embryonic cell. White-filled red and red-filled circles are adult haematopoietic stem cells and adult blood cells, respectively. Here, for simplicity, we assumed a uniform mutation rate of one substitution per cell per cell doubling. Each mutation during cell doubling is represented by a number in a black-filled rectangle. Mutations accumulated in a specific early cell are shown with numbers next to the cell. The final mutations acquired at an early cell of cell generation IV (16-cell stage) and their expected relative contribution to adult blood tissues (1 out of 16 or 6.6%) is summarized in the box below the cellular phylogenetic tree. We assumed that breast cancer (green-filled circles) cells are descended from the embryonic cell of the leftmost lineage (which has mutations 1, 3, 7 and 15). In the circumstances, the expected features of early embryonic mutations (VAFs, chance to be shared with breast cancer) are summarized in the right table. b, An alternative scenario with a neoplastic clonal expansion in the blood (here we assumed a haematopoietic stem cell contributes 40% of all blood cells). We assumed that additional 100 somatic mutations were further acquired during late cell doublings. The expected features are summarized in the right table.
Extended Data Figure 4 Features of mutations in blood samples with neoplastic clonal expansions.
Mutations from samples with evidence of neoplastic clonal expansions display more similar VAFs to (the right violin plot) each other compared to mutations from samples without neoplastic clonal expansions (the left violin plot).
Extended Data Figure 5 Features of the early embryonic mutations identified in this study.
a, As expected for early embryonic mutations, we observe no relationship between the age of individuals and the number of mutations found in an individual. In case of late mutations, we find more mutations in the aged individuals (Fig. 1f). b, Features of mutations in the samples (n = 7) with four early embryonic mutations suggest that these mutations are not likely to be related with a neoplastic clonal expansion: VAFs of mutations are diverse and a fraction of these mutations are shared with the matched cancer. The corresponding VAFs in the matched tumour tissues are shown in numbers above the bars. c, Samples with neoplastic clonal expansions (that is, PD9568b, PD9752b and PD9569b) show different features: mutations show similar VAFs each other and are not shared by cancer cells. d, Enrichment of early mutations according to ENCODE dataset. We find higher mutation frequency in transcriptionally repressed (R) than active (T) regions, but the difference is non-significant in our study (χ2 test, degrees of freedom = 1, P value = 0.4696), presumably due to the insufficient number of early embryonic mutations (n = 163). R, repressed chromatin; T, transcribed chromatin; CTCF, CTCF-bound regions; E, enhancer related; TSS/PF, promoter related. e, From a simulation study using 1,000 in silico embryonic mutations, we assessed the detection sensitivity of early embryonic mutations from 32× whole-genome sequencing (see Methods). This sensitivity was used in downstream analyses (for example, likelihood tests for understanding the asymmetry of cell doublings and tests for the calculation of the early embryonic mutation rates. Error bars denote 95% confidence interval using exact Poisson tests.
Extended Data Figure 6 Expected proportion of early embryonic mutations shared by cancer according to the cell generation gap between the MRCA cell of adult blood cells and the MRCA cell of all somatic cells.
See Supplementary Discussion 4. a, A scenario in which there is no cell generation gap. Early mutations are represented by asterisks in colours. A summary of the expected proportion of mutations shared with cancer cells is shown in the table: the chance is twice the VAF of each early embryonic mutation. b, A scenario in which the MRCA cell of adult blood cells is formed one cell generation later than the MRCA cell of all somatic cells. The chance is identical to the VAF of each early embryonic mutation. c, A scenario in which the MRCA cell of adult blood cells is formed two cell generations later than the MRCA cell of all somatic cells. The chance is half the VAF of each early embryonic mutation.
Extended Data Figure 7 The MRCA cell of adult blood cells is the MRCA cell of all somatic cells (or the fertilized egg).
See Supplementary Discussion 4. Using the expected proportion of mutations shared with cancer (Extended Data Fig. 6), we estimated the timing when the MRCA cell of adult blood cells is formed. The four orange boxes show the expected proportions from four scenarios, when there are 0, 1, 2 and 3 cell generation gaps between the MRCA cells. The observed proportion (26%; green horizontal line) in this study is closest to the expectation from the model of 0 cell generation gap. Error bars are interquartile range × 2 (from the simulation study).
Extended Data Figure 8 The simulation study to understand potential stochasticity in the embryoblast formation.
See Methods, ‘A stochastic model of embryoblast formation’ for more details. a, The expected distribution of VAF of early embryonic mutations in a stochastic model in which n cells (y axis) are randomly selected as epiblasts from the 32-cell stage embryo. The size of circle is proportional to the relative frequency of mutations at each VAF. b, The stochastic model estimates the number of founder epiblast cells and the timing (cell stage) of their commitment. The maximum likelihood is selection of 11 cells in 64-cell stage. c, The VAF distribution of early embryonic mutations expected from the maximum likelihood stochastic model. The maximum likelihood estimation (MLE) and the posterior probability by a Bayesian approach are shown by green and purple curves, respectively. Our observation of the 163 early embryonic mutations is represented by the histogram. d, Unequal contribution of the first two cells to ICM cells by direct observation of 12 mouse-embryos using inverted light-sheet microscope (see ref. 19). Schematic diagram (cell phylogeny) is shown above the bar graph. We reanalysed their observation, counting the relative contribution to ICM (black dots indicate the observed asymmetry in each embryo). These unequal contribution levels ranged from 0.5:0.5 to 0.74:0.26 and the average was 0.6:0.4.
Extended Data Figure 9 Early embryonic mutations (n = 7) identified from three large families.
a–g, Sequencing reads (using IGV images) for the seven mutation loci are shown. All mutations are subclonal to a specific allele of a heterozygous SNP in the vicinity. As expected for early embryonic mutations, the VAFs of mutant alleles are lower than 0.5 and the mutant alleles are not found in the genomes of all the parents and the siblings. It was possible to perform ultrahigh-depth targeted amplicon sequencing (by MiSeq) on three mutations, and all were successfully validated.
Extended Data Figure 10 Signatures of early embryonic mutations.
a, The mutational spectrum for 163 early embryonic mutations is displayed according to the 96 substitution classes (defined by 6 substitution classes (C>A, C>G, C>T, T>A, T>C, T>G) and 16 sequence contexts (immediate 5′ and 3′ bases to the mutated pyrimidine bases; see ref. 7 for more details). The observed spectrum can be decomposed into two known mutational signatures (signatures 5 and 1), suggesting that endogenous mutational processes are dominantly operative in early human embryogenesis (see Supplementary Discussion 6 for more details). b, The methylation status of 28 C>T early embryonic mutations occurred at  sequence contexts. Methylation levels were obtained from a previous report38. The vast majority of the 28 loci were methylated, which is higher than background (right).
sequence contexts. Methylation levels were obtained from a previous report38. The vast majority of the 28 loci were methylated, which is higher than background (right).
Supplementary information
Supplementary Information
This file contains the Supplementary Discussion and additional references. (PDF 200 kb)
Supplementary Table 1
A list of samples sequenced in this study. (XLSX 71 kb)
Supplementary Table 2
The list of primers used for targeted amplicon sequencing. (XLSX 127 kb)
Supplementary Table 3
A list of somatic mutations catalogued in this study. (XLSX 353 kb)
Rights and permissions
About this article
Cite this article
Ju, Y., Martincorena, I., Gerstung, M. et al. Somatic mutations reveal asymmetric cellular dynamics in the early human embryo. Nature 543, 714–718 (2017). https://doi.org/10.1038/nature21703
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21703
This article is cited by
-
Human embryonic genetic mosaicism and its effects on development and disease
Nature Reviews Genetics (2024)
-
Spatial architectures of somatic mutations in normal prostate, benign prostatic hyperplasia and coexisting prostate cancer
Experimental & Molecular Medicine (2024)
-
Cell-type-resolved mosaicism reveals clonal dynamics of the human forebrain
Nature (2024)
-
Grave-to-cradle: human embryonic lineage tracing from the postmortem body
Experimental & Molecular Medicine (2023)
-
Somatic genetic variation in healthy tissue and non-cancer diseases
European Journal of Human Genetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.