Abstract
Recurrent chromosomal translocations producing a chimaeric MLL oncogene give rise to a highly aggressive acute leukaemia associated with poor clinical outcome1. The preferential involvement of chromatin-associated factors as MLL fusion partners belies a dependency on transcription control2. Despite recent progress made in targeting chromatin regulators in cancer3, available therapies for this well-characterized disease remain inadequate, prompting the need to identify new targets for therapeutic intervention. Here, using unbiased CRISPR–Cas9 technology to perform a genome-scale loss-of-function screen in an MLL-AF4-positive acute leukaemia cell line, we identify ENL as an unrecognized gene that is specifically required for proliferation in vitro and in vivo. To explain the mechanistic role of ENL in leukaemia pathogenesis and dynamic transcription control, a chemical genetic strategy was developed to achieve targeted protein degradation. Acute loss of ENL suppressed the initiation and elongation of RNA polymerase II at active genes genome-wide, with pronounced effects at genes featuring a disproportionate ENL load. Notably, an intact YEATS chromatin-reader domain was essential for ENL-dependent leukaemic growth. Overall, these findings identify a dependency factor in acute leukaemia and suggest a mechanistic rationale for disrupting the YEATS domain in disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Krivtsov, A. V. & Armstrong, S. A. MLL translocations, histone modifications and leukaemia stem-cell development. Nat. Rev. Cancer 7, 823–833 (2007)
Slany, R. K. When epigenetics kills: MLL fusion proteins in leukemia. Hematol. Oncol. 23, 1–9 (2005)
Cai, S. F., Chen, C.-W. & Armstrong, S. A. Drugging chromatin in cancer: recent advances and novel approaches. Mol. Cell 60, 561–570 (2015)
Ott, C. J. et al. BET bromodomain inhibition targets both c-Myc and IL7R in high-risk acute lymphoblastic leukemia. Blood 120, 2843–2852 (2012)
Delmore, J. E. et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell 146, 904–917 (2011)
Zuber, J. et al. RNAi screen identifies Brd4 as a therapeutic target in acute myeloid leukaemia. Nature 478, 524–528 (2011)
Bernt, K. M. et al. MLL-rearranged leukemia is dependent on aberrant H3K79 methylation by DOT1L. Cancer Cell 20, 66–78 (2011)
Daigle, S. R. et al. Potent inhibition of DOT1L as treatment of MLL-fusion leukemia. Blood 122, 1017–1025 (2013)
Daigle, S. R. et al. Selective killing of mixed lineage leukemia cells by a potent small-molecule DOT1L inhibitor. Cancer Cell 20, 53–65 (2011)
Guenther, M. G. et al. Aberrant chromatin at genes encoding stem cell regulators in human mixed-lineage leukemia. Genes Dev. 22, 3403–3408 (2008)
Krivtsov, A. V. et al. H3K79 methylation profiles define murine and human MLL-AF4 leukemias. Cancer Cell 14, 355–368 (2008)
Okada, Y. et al. hDOT1L links histone methylation to leukemogenesis. Cell 121, 167–178 (2005)
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014)
Luo, B. et al. Highly parallel identification of essential genes in cancer cells. Proc. Natl Acad. Sci. USA 105, 20380–20385 (2008)
Levis, M. et al. A FLT3-targeted tyrosine kinase inhibitor is cytotoxic to leukemia cells in vitro and in vivo . Blood 99, 3885–3891 (2002)
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015)
Li, Y. et al. AF9 YEATS domain links histone acetylation to DOT1L-mediated H3K79 methylation. Cell 159, 558–571 (2014)
Li, Y. et al. Molecular coupling of histone crotonylation and active transcription by AF9 YEATS domain. Mol. Cell 62, 181–193 (2016)
Andrews, F. H. et al. The Taf14 YEATS domain is a reader of histone crotonylation. Nat. Chem. Biol. 12, 396–398 (2016)
Shanle, E. K. et al. Association of Taf14 with acetylated histone H3 directs gene transcription and the DNA damage response. Genes Dev. 29, 1795–1800 (2015)
Mueller, D. et al. A role for the MLL fusion partner ENL in transcriptional elongation and chromatin modification. Blood 110, 4445–4454 (2007)
Yokoyama, A., Lin, M., Naresh, A., Kitabayashi, I. & Cleary, M. L. A higher-order complex containing AF4 and ENL family proteins with P-TEFb facilitates oncogenic and physiologic MLL-dependent transcription. Cancer Cell 17, 198–212 (2010)
He, N. et al. Human polymerase-associated factor complex (PAFc) connects the super elongation complex (SEC) to RNA polymerase II on chromatin. Proc. Natl Acad. Sci. USA 108, E636–E645 (2011)
Rubnitz, J. E., Morrissey, J., Savage, P. A. & Cleary, M. L. ENL, the gene fused with HRX in t(11;19) leukemias, encodes a nuclear protein with transcriptional activation potential in lymphoid and myeloid cells. Blood 84, 1747–1752 (1994)
Zeisig, D. T. et al. The eleven-nineteen-leukemia protein ENL connects nuclear MLL fusion partners with chromatin. Oncogene 24, 5525–5532 (2005)
Winter, G. E. et al. Phthalimide conjugation as a strategy for in vivo target protein degradation. Science 348, 1376–1381 (2015)
Clackson, T. et al. Redesigning an FKBP-ligand interface to generate chemical dimerizers with novel specificity. Proc. Natl Acad. Sci. USA 95, 10437–10442 (1998)
Rathert, P. et al. Transcriptional plasticity promotes primary and acquired resistance to BET inhibition. Nature 525, 543–547 (2015)
Peterlin, B. M. & Price, D. H. Controlling the elongation phase of transcription with P-TEFb. Mol. Cell. 23, 297–305 (2006)
Wilkinson, A. C. et al. RUNX1 is a key target in t(4;11) leukemias that contributes to gene activation through an AF4-MLL complex interaction. Cell Reports 3, 116–127 (2013)
Shi, J. et al. Discovery of cancer drug targets by CRISPR–Cas9 screening of protein domains. Nat. Biotechnol. 33, 661–667 (2015)
Acknowledgements
The authors thank S. A. Armstrong, C. D. Allis, and X. Shi for transparent and supportive dialogue. We also thank J. A. Perry for editing the manuscript and N. S. Gray and C. J. Ott for suggestions. Quantitative proteomics studies were performed by R. Kunz. This research was supported by philanthropic gifts from K. Lubin and E. Woods, as well as NIH grants (R01-CA176745 and P01-CA109901 to J.E.B.). G.E.W. was supported by an EMBO long-term fellowship. D.L.B. is a Merck Fellow of the Damon Runyon Cancer Research Foundation (DRG-2196-14). N.E.S. is supported by a Pathway to Independence Award (R00-HG008171) from the NHGRI.
Author information
Authors and Affiliations
Contributions
M.A.E. performed experiments and analysed data. G.E.W. designed plasmids for the dTAG system with J.M.R. and performed CRISPR–Cas9 screens collaboratively with N.E.S., O.S. and F.Z. T.G.S. assisted cellular assays. B.E.L., H.X. and S.H.O. performed experiments on HSPCs. J.P., H.-S.S., N.K.O.-A. and S.D.-P. performed protein biochemistry. A.S. performed mouse experiments. S.D. and D.L.B. designed and synthesized dTAG molecules. B.N. assisted in sgRNA validation. R.Z. assisted in exon-scanning CRISPR–Cas9. M.A.E., G.E.W. and J.E.B. designed the experimental strategy and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.E.B. is now an employee, shareholder, and executive of Novartis Pharmaceuticals; G.E.W. is a consultant for C4 Therapeutics; and N.E.S., O.S. and F.Z. are inventors on a patent application related to the CRISPR screening technology.
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 ENL, but not AF9, is required for growth of acute leukaemia.
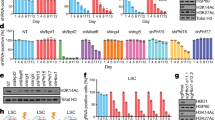
a, Summary of CRISPR–Cas9 knockout screen results in MV4;11 cells showing the number of hits (NES < 0.1 and P < 0.05 in both replicates) and the percentage of hits known to be essential genes. Right, top 15 hits in essential and non-essential protein-coding genes. b, Immunoblot of ENL 5 days after transduction with the indicated sgRNA. c, Immunoblot of FLT3 5 days after transduction with the indicated sgRNA. d, Competition-based CRISPR–Cas9 mutagenesis in MV4;11-Cas9 cells. Percentage GFP+ subpopulation (sgRNA+) after transduction with lentiviral constructs co-expressing GFP and the sgRNA indicated. Mean ± s.d., n = 3. e, ENL protein expression by immunoblot of all cell lines tested for ENL-dependent growth. f–j, As in d but for the cell lines and sgRNAs indicated. k, Indel quantification by TIDE (tracking of indels by sequence trace decomposition) analysis 5 days after transduction with the sgRNA indicated in MV4;11-Cas9 cells. l, As in d for the cell lines and sgRNAs indicated, n = 4 for MV4;11, n = 3 for MOLM-13.
Extended Data Figure 2 Loss of mouse Enl has minimal impact on LSK cells.
a, Validation of sgRNA targeting mouse Enl. Indel quantification of genomic Enl locus in NIH/3T3-Cas9 cells by TIDE analysis 5 days after transduction with control Rosa26-sg (left) or Enl-sg (right). b, Mac-1 and Gr-1 myeloid marker staining of viable doxycycline-inducible Cas9-expressing LSK cells 9 days after transduction with the sgRNA indicated.
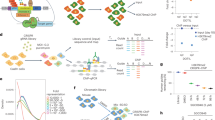
Extended Data Figure 3 Characterization of dTAG-ENL system.
a, Chemical structure of dTAG-7. b, Dose-responsive FKBP12(F36V)-ENL degradation detected by immunoblot after 16 h treatment of MV4;11 (Cas9+, ENL-FKBP12(F36V)–HA-ENL+) cells with DMSO (−) or dTAG-7 at the indicated concentration. c, As in b, but with dTAG-13. d, Immunoblot detection of ENL-FKBP12(F36V) degradation in MV4;11 (Cas9+, ENL-FKBP12(F36V)–HA+) after 16 h. e, Kinetic evaluation of FKBP12(F36V)-ENL degradation by dTAG-7 (500 nM) in MV4;11 (Cas9+, FKBP12(F36V)–HA-ENL+) cells. f, Degradation of ENL-FKBP12(F36V) in MOLM-13 cells after a 4-h treatment of ENL-FKBP12(F36V)-expressing cells with dTAG-13 (500 nM). g, As in f, but with NOMO-1 cells expressing ENL-FKBP12(F36V). h, As in f, but with JURKAT cells expressing ENL-FKBP12(F36V). i, Long-term kinetic evaluation of ENL-FKBP12(F36V) degradation by a single dose of dTAG-7 (500 nM) in MV4;11 (Cas9+, ENL-FKBP12(F36V)–HA+) cells. j, As in i, but with dTAG-13 (500 nM).
Extended Data Figure 4 Response to dTAG-7 and dTAG-13.
a, CRISPR–Cas9 knockout of endogenous ENL in MV4;11 (Cas9+, ENL-FKBP12(F36V)–HA+) cells. ENL/HA immunoblot in MV4;11 (Cas9+, ENL-FKBP12(F36V)–HA+) (P, parental) and MV4;11 (Cas9+, ENL-FKBP12(F36V)–HA+, ENL−/−) cells. b, CRISPR–Cas9 knockout of endogenous ENL in MV4;11 (Cas9+, FKBP12(F36V)–HA-ENL+) cells. c, DMSO-normalized cellular viability of MV4;11 (Cas9+, ENL-FKBP12(F36V)–HA+, ENL−/−) cells after 72-h treatment with dTAG-13 approximated by ATP-lite assay. Mean ± s.d., n = 4. d, DMSO-normalized cellular viability of MV4;11 (Cas9+, FKBP12(F36V)–HA-ENL+, ENL−/−) cells after 72-h treatment with dTAG-7 or dTAG-13 approximated by ATP-lite assay. Mean ± s.d., n = 4. e, Growth over time of MV4;11 (Cas9+, ENL-FKBP12(F36V)–HA+, ENL−/−) cells treated with DMSO or dTAG-13 (500 nM). Total number of viable cells as approximated by trypan blue exclusion is plotted over time. Mean ± s.d., n = 3. f, g, As in e, but with cell line indicated at top. h, As in d, but with wild-type MV4;11 cells. i, Representative plots of BrdU incorporation used for cell cycle analysis shown in Fig. 2f.
Extended Data Figure 5 Genomic localization of ENL and ENL-FKBP12(F36V) in MV4;11 cells.
a, Correlation of ENL and ENL-FKBP12(F36V) genomic enrichment by ChIP–seq. Top, scatterplot of ENL-FKBP12(F36V) and ENL ChIP–seq signal (log2 RPM per bp) at the union of all regions enriched with ENL-FKBP12(F36V) or ENL. Bottom, Venn diagram depicts the number of unique and overlapping regions of enrichment identified by ChIP–seq of ENL-FKBP12(F36V) (18,294 total) and ENL (5,397 total). b, ENL binding at active enhancers. Top, rank-ordered heat map of H3K27ac, ENL-FKBP12(F36V), and ENL at H3K27ac-defined active enhancers. Rows depict a single enhancer (centred on an H3K27ac peak) and are sorted by ENL-FKBP12(F36V) signal. ChIP–seq signal is depicted by scaled colour intensity. Bottom, meta plot representation of heat maps. Average read density is plotted against the distance from the enhancer centre. c, ENL binding at promoters by ChIP–seq. Top, rank-ordered heat map of ChIP–seq signals at ENL-FKBP12(F36V)-bound promoters, sorted by ENL-FKBP12(F36V) levels and aligned at TSS. ChIP–seq signal is depicted by scaled colour intensity. Bottom, meta-plot representation of heat maps. Average read density is plotted against the distance from the TSS. H3K9ac, ENL-FKBP12(F36V) and ENL samples are also shown in Fig. 3a. d–i, Correlation of ENL and ENL-FKBP12(F36V) with histone acetyl-lysine residues by ChIP–seq. Scatterplot of ChIP–seq signals (log2 RPM per bp) at the union of all enriched regions for the two samples indicated in each plot. j, Asymmetric localization of ENL-FKBP12(F36V). ChIP–seq signal at each location is plotted against its rank among all occupied regions. Points coloured red are loci with asymmetric ENL-FKBP12(F36V) load compared to typically loaded regions shown in grey. P values determined by Pearson correlation.
Extended Data Figure 6 Genomic localization of ENL in MOLM-13 cells.
a, ENL binding at enhancers and promoters. Top, mean H3K27ac and ENL ChIP–seq signal across all H3K27ac-defined active enhancers in MOLM-13 cells. Average read density is plotted against the distance from the enhancer centre. Bottom, mean H3K27ac and ENL ChIP–seq signal at ENL-bound TSS regions in MOLM-13 cells. Average read density is plotted against the distance from the TSS. Dataset for H3K27ac was obtained from GEO accession GSM1652920. b, Correlation of H3K27ac and ENL genomic enrichment by ChIP–seq. Scatterplot of H3K27ac and ENL ChIP–seq signal at the union of all H3K27ac and ENL enriched regions genome-wide. P value determined by Pearson correlation. c, Asymmetric localization of ENL in MOLM-13 cells. ChIP–seq signal at each location is plotted against its rank among all occupied regions. Points coloured red are loci with asymmetric ENL load compared to typically loaded regions shown in grey. d, Gene tracks of ChIP–seq signal at examples of asymmetrically (MYB) and typically enriched (ACTB) ENL target genes.
Extended Data Figure 7 Loss of ENL disrupts SEC recruitment and activity.
a, Boxplot of DMSO-normalized fold changes in gene expression of typically enriched (grey) and asymmetrically enriched (red) ENL-FKBP12(F36V) target genes with dTAG-13 (500 nM) treatment. P values from Welch’s two-tailed t-test. b, c, Gene set enrichment analysis (GSEA) of DMSO-normalized gene-expression changes with 8 h dTAG-13 (500 nM) treatment compared to the set of asymmetrically loaded ENL (b) and ENL-FKBP12(F36V) (c) target genes. d, Boxplots of change in RNA Pol II ChIP–Rx signal (RRPM) at TSS and gene-body regions of typically enriched (grey) and asymmetrically enriched (red) ENL-FKBP12(F36V) target genes after 24 h dTAG-13 (500 nM) treatment. P values from Welch’s two-tailed t-test. e, Cumulative distribution plot of RNA Pol II travelling ratio at asymmetrically loaded ENL (left) and ENL-FKBP12(F36V) (right) target genes determined by RNA Pol II ChIP–Rx after 24 h DMSO or dTAG-13 (500 nM) treatment. f, Waterfall plot (left) and boxplot (right) of change in AFF4 ChIP–Rx signal (RRPM) at promoters (TSS ± 5 kb) of ENL-FKBP12(F36V) target genes (typically enriched in grey, asymmetrically enriched in red) after 6 h dTAG-13 treatment (500 nM). P value from Welch’s two-tailed t-test. g, As in f, but for CDK9. h, Meta-gene representations of Pol II S2P ChIP–Rx signal at ENL and ENL-FKBP12(F36V) target genes after 24 h dTAG-13 (500 nM) treatment. i, Gene tracks of ChIP–seq signal at an example of an asymmetrically enriched ENL target gene, MEIS1.
Extended Data Figure 8 Comparison of MV4;11 response to ENL degradation and DOT1L inhibition.
a, Immunoblot for H3K79me2 after 96 h DMSO, dTAG-13 (500 nM), or EPZ-5676 (500 nM) treatment. Bottom, semi-quantitative representation of H3-normalized H3K79me2 signal as a percentage of DMSO treatment. b, Gene track view of H3K79me2 ChIP–seq signal at examples of ENL target genes (asymmetric load: MYB and MEIS1; typical load: ACTB) after 96 h DMSO or dTAG13 (500 nM) treatment. c, Meta gene representation of H3K79me2 ChIP–seq mean read density at typically loaded (left) and asymmetrically loaded (right) ENL target genes. d, Kinetic comparison of dTAG-13 (500 nM) and EPZ-5676 (500 nM) on gene expression changes in MV4;11 (Cas9+, ENL-FKBP12(F36V)–HA+, ENL−/−) cells. Gene expression quantified by quantitative PCR with reverse transcription (qRT–PCR) (ΔΔCt method). Student’s t-tests comparing DMSO to EPZ-5676 and dTAG-13 are in grey and red, respectively. Mean ± s.d., triplicate PCR analysis. e, Cell cycle analysis via propidium iodide staining of MV4;11 (Cas9+, ENL-FKBP12(F36V)–HA+, ENL−/−) cells treated with DMSO, dTAG-13 (500 nM), or EPZ-5676 (500 nM). Percentage of G1 cells were compared by two-tailed t-tests. Mean ± s.d., n = 3. f, Gene expression changes in MV4;11 (Cas9+, ENL-FKBP12(F36V)–HA+, ENL−/−) cells with combination treatments of dTAG-13 (500 nM) and EPZ-5676 (500 nM). Gene expression quantified by qRT–PCR (ΔΔCt method). P values by Student’s t-test. Mean ± s.d., triplicate PCR analysis. g, Overlap analysis of published MLL-AF4 target genes and asymmetrically loaded ENL or ENL-FKBP12 target genes. h, Heat map of DMSO-normalized fold changes in gene expression caused by dTAG-13 (500 nM) or EPZ-5676 (500 nM) treatment for the indicated amount of time. Raw expression values determined by RNA-seq were cell-count-normalized by spike-in of synthetic ERCC RNA standards. dTAG-13 data are redundant with data shown in Fig. 2g. i, GSEA of DMSO-normalized gene-expression changes with dTAG-13 (500 nM) or EPZ-5676 (500 nM) treatment compared to MLL-AF4 target genes. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Not significant (n.s.) P > 0.05.
Extended Data Figure 9 Characterization of ENL YEATS mutations.
a, Sequence alignment of AF9 and ENL YEATS domains. Identical residues are highlighted in grey. Two of these residues are coloured red to denote sites in in which mutations were engineered into ENL for interrogation of YEATS function. b, Histone peptide microarray probed with lysates from 293T cells overexpressing full-length wild-type ENL, or mutants F47A or Y78A, with a 3×HA epitope tag. Select residues known to be YEATS-domain substrates and showing YEATS-domain-dependent signals have been boxed in red, blue or orange. Several other spots showed high signal, but not enriched over the corresponding unmodified peptide, and not YEATS-dependent. These have not been boxed in. c, Quantification of histone peptide microarray spots containing H3K18ac or H3K27ac after probing with cellular lysates from 293T cells overexpressing wild-type or mutant 3×HA–ENL. Mean ± s.d., n = 2. d, Isothermal titration calorimetry (ITC) measurements showing wild-type ENL bound to H34–10K9ac in a 1:0.943 stoichiometry with binding affinity (dissociation constant, Kd) of 30.1 μM. Interaction was not detected with ENL Y78A. e, Subcellular localization of ENL. MV4;11 cells stably expressing 3×HA–ENL (wild-type, F47A or Y78A) were fractionated and probed by immunoblot. Cy, cytoplasm; N nucleoplasm; Ch, chromatin. f, Thermal stability of ENL in cells. MV4;11 cells stably expressing 3×HA–ENL (wild-type, F47A or Y78A) were heated to the indicated temperature to induce irreversible aggregation. Remaining soluble protein was isolated and probed by immunoblot. g, Quantification of band intensity in e showing the temperature of aggregation (Tagg). h, Immunoblot to determine expression level of exogenously expressed ENL constructs used for rescue and ChIP–qPCR experiments.
Supplementary information
Supplementary Figure 1
This file contains the uncropped versions of all gel images in the manuscript. (PDF 3128 kb)
Supplementary Table 1
This file contains the source data for the genome scale CRISPR/Cas9 screen. (XLSX 8391 kb)
Supplementary Information
This file contains Supplementary Methods, dTAG molecule characterizations and additional references. (PDF 1005 kb)
Rights and permissions
About this article
Cite this article
Erb, M., Scott, T., Li, B. et al. Transcription control by the ENL YEATS domain in acute leukaemia. Nature 543, 270–274 (2017). https://doi.org/10.1038/nature21688
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21688
This article is cited by
-
PROTAC-mediated vimentin degradation promotes terminal erythroid differentiation of pluripotent stem cells
Stem Cell Research & Therapy (2024)
-
High expression of YEATS2 as a predictive factor of poor prognosis in patients with hepatocellular carcinoma
Scientific Reports (2024)
-
RNA sequestration in P-bodies sustains myeloid leukaemia
Nature Cell Biology (2024)
-
CHSY1 promotes CD8+ T cell exhaustion through activation of succinate metabolism pathway leading to colorectal cancer liver metastasis based on CRISPR/Cas9 screening
Journal of Experimental & Clinical Cancer Research (2023)
-
Multi-landmark alignment of genomic signals reveals conserved expression patterns across transcription start sites
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.