Abstract
Zika virus (ZIKV) has recently emerged as a pandemic associated with severe neuropathology in newborns and adults1. There are no ZIKV-specific treatments or preventatives. Therefore, the development of a safe and effective vaccine is a high priority. Messenger RNA (mRNA) has emerged as a versatile and highly effective platform to deliver vaccine antigens and therapeutic proteins2,3. Here we demonstrate that a single low-dose intradermal immunization with lipid-nanoparticle-encapsulated nucleoside-modified mRNA (mRNA–LNP) encoding the pre-membrane and envelope glycoproteins of a strain from the ZIKV outbreak in 2013 elicited potent and durable neutralizing antibody responses in mice and non-human primates. Immunization with 30 μg of nucleoside-modified ZIKV mRNA–LNP protected mice against ZIKV challenges at 2 weeks or 5 months after vaccination, and a single dose of 50 μg was sufficient to protect non-human primates against a challenge at 5 weeks after vaccination. These data demonstrate that nucleoside-modified mRNA–LNP elicits rapid and durable protective immunity and therefore represents a new and promising vaccine candidate for the global fight against ZIKV.
Similar content being viewed by others
Main
ZIKV, first identified in 1947 (ref. 4), is a mosquito-borne and sexually transmitted flavivirus that has recently been associated with microcephaly and other birth defects in newborns and Guillain–Barré syndrome in adults1. Effective vaccines have been approved for other closely related flaviviruses5,6,7, but vaccine candidates for ZIKV have only recently been developed8,9,10,11,12. Multiple vaccine formats have been shown to protect mice or non-human primates against ZIKV infection, including plasmid DNA9,10,11,12, purified inactivated virus10,11, protein subunit8 and adenovirus vectors8,10. The ideal vaccine is safe and induces protective immunity after a single immunization, regardless of prior serologic history. Of the candidate Zika vaccines described to date, only a rhesus adenovirus platform (RhAd52) has been shown to confer protection after a single immunization in non-human primates; however, the efficacy of the RhAd52 vector in humans is currently unknown. Additionally, pre-existing immunity to adenovirus serotypes can limit the efficacy of such vectors13,14, and low neutralizing titres to rhesus adenoviruses, including RhAd52, have been detected in humans15.
mRNA has emerged as a promising vaccine modality that can elicit potent immune responses (reviewed in refs 2, 3), while avoiding the safety risks and anti-vector immunity associated with some live virus vaccines (reviewed in ref. 16). Vaccination with mRNA offers several advantages over other vaccine platforms: (i) it is a non-integrating, non-infectious gene vector that can be readily designed to express any protein with high efficiency; (ii) it has the potential for cost-effective and highly scalable manufacturing; and (iii) small doses are sufficient to induce protective immune responses.
Here, we designed a potent anti-ZIKV vaccine in which the pre-membrane and envelope (prM–E) glycoproteins of ZIKV H/PF/2013 (ref. 17) are encoded by mRNA (Extended Data Fig. 1a) containing the modified nucleoside 1-methylpseudouridine (m1Ψ), which prevents innate immune sensing and increases mRNA translation in vivo18. Nucleoside-modified ZIKV prM–E mRNA was prepared for vaccination by encapsulation in lipid nanoparticles (LNPs), which have been shown to mediate efficient and prolonged protein expression by mRNA in vivo19. Studies of ZIKV and other flaviviruses have demonstrated that co-expression of pre-membrane and envelope proteins is sufficient to assemble and secrete subviral particles12,20. ZIKV prM–E-encoding mRNA was first characterized by transfecting HEK293T cells and human and mouse dendritic cells. ZIKV envelope (E) protein was produced by all cell types and was secreted into the supernatant of HEK293T cells (Extended Data Fig. 1b). We hypothesize that dendritic cells also secrete E protein and that it is rapidly internalized by endocytosis, as has been proposed for HIV gag21. E protein in the supernatant of HEK293T cells was pelleted by ultracentrifugation when incubated with PBS, but not with 0.5% Triton X-100, consistent with subviral particle production from prM–E mRNA20 (Extended Data Fig. 1c).
The immune response induced by the nucleoside-modified ZIKV prM–E mRNA–LNP vaccine was first analysed in C57BL/6 mice. Animals were immunized by intradermal injection of 30 μg of ZIKV prM–E mRNA–LNP or polycytidylic acid (poly(C)) RNA–LNP as a negative control. No inflammation or other adverse events were observed at the sites of injection. Polyfunctional E-protein-specific CD4+ T cell responses were detected on the basis of intracellular IFNγ, TNF (also known as TNFα) and IL-2 production by splenocytes stimulated with ZIKV E protein at week 2 after vaccination (Fig. 1a and Extended Data Fig. 2). ZIKV E-protein-specific immunoglobulin G (IgG) in the serum developed quickly in vaccinated mice and stabilized at an endpoint titre of 180,000 (90 μg ml−1) at weeks 8–12 (Fig. 1b and Extended Data Fig. 3a). Anti-ZIKV neutralizing antibodies (nAb) were measured using two independent assays: a standard plaque-reduction neutralization test (PRNT) and a ZIKV reporter viral particle (RVP) assay12. The mean PRNT50 titre against ZIKV MR-766 peaked at around 1,300 at week 8 (Fig. 1c) and was relatively stable until week 12. The mean RVP nAb titre (EC50) against ZIKV H/PF/2013 reached approximately 105 at weeks 8–12 (Fig. 1d). The detection of higher nAb titres in the RVP assay compared to other assay formats similar to PRNT has previously been reported12. In addition, we noted that the ratio of RVP to PRNT titres was not fixed and varied with the animal model and viral isolate.
a–d, C57BL/6 mice were immunized by intradermal injection of 30 μg of nucleoside-modified ZIKV prM–E mRNA–LNP (n = 8) or control poly(C) RNA–LNP (n = 4). a, At week 2, splenic antigen-specific CD4+ T cells were detected by intracellular cytokine staining. b–d, The antibody response was analysed by ELISA (b), PRNT using ZIKV MR-766 (c) and RVP using ZIKV H/PF/2013 (d). e–g, BALB/c mice were immunized similarly with ZIKV mRNA–LNP (n = 10) or poly(C) RNA–LNP (n = 5) and immune responses were analysed by ELISA (e), PRNT using MR-766 (f) and RVP using H/PF/2013 (g). Points represent individual mice; horizontal lines show the mean; dotted lines indicate the limit of detection. The controls in d and g are from the week 8 time point. *P < 0.05 (unpaired t-test) in a; antibody responses in vaccine and control groups were compared at each time point by Mann–Whitney test, P < 0.01 for all comparisons (b–g).
Immunogenicity of the nucleoside-modified ZIKV prM–E mRNA–LNP vaccine was next evaluated in BALB/c mice. E-protein-specific IgG in the serum peaked at week 8 and remained stable between weeks 8 and 20 (endpoint titres around 200,000; 90–130 μg ml−1) (Fig. 1e and Extended Data Fig. 3b). PRNT50 nAb increased to a maximum of about 1,100 at week 16 and remained stable until week 20 (Fig. 1f). The RVP nAb titre rose to 50,000 at week 8 and remained above 20,000 until week 20 (Fig. 1g).
A challenge study was conducted in BALB/c mice immunized by intradermal injection of 30 μg of nucleoside-modified ZIKV prM–E mRNA–LNP or poly(C) RNA–LNP. Mice were challenged by intravenous injection at week 2 (short term) or week 20 (long term) after immunization with 200 plaque-forming units (PFU) of ZIKV PRVABC59. In the short-term protection study, 8 out of 9 control mice showed the presence of viral RNA in the blood (viraemia) by day 3, with a median peak of around 14,000 copies per ml. All ZIKV mRNA-immunized mice (n = 9) were protected against detectable viraemia (Fig. 2a). In the long-term study, all control mice (n = 5) showed viraemia on day 3, with a median peak of 1,200 copies per ml, whereas none of the ZIKV mRNA-immunized mice (n = 10) had detectable viraemia at days 3 or 7 (Fig. 2b). These data demonstrate that a single immunization with nucleoside-modified ZIKV prM–E mRNA–LNP rapidly elicits durable protection against detectable viraemia with a heterologous ZIKV strain in mice.
a, b, BALB/c mice immunized by intradermal injection of 30 μg of ZIKV prM–E mRNA–LNP or control poly(C) RNA–LNP were challenged by intravenous injection of 200 PFU ZIKV PRVABC59 at 2 weeks (a; n = 9 per group) or 20 weeks (b; n = 5 control mice; n = 10 ZIKV mRNA–LNP mice) after vaccination and plasma viral loads were measured by qRT–PCR for ZIKV capsid RNA. Two overlapping curves are indicated by a dagger symbol. Dotted lines indicate the limit of detection (200 copies per ml), with undetectable curves staggered to show individual mice. Day 3 viraemia in vaccine and control groups was compared by Mann–Whitney test, P < 0.001 for both challenges.
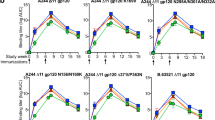
We next evaluated the efficacy of the nucleoside-modified ZIKV mRNA–LNP vaccine in rhesus macaques (Macaca mulatta), a non-human primate species that shows several features of ZIKV infection in humans22. Macaques were immunized by intradermal injection with doses of 600 μg, 200 μg or 50 μg of nucleoside-modified ZIKV prM–E mRNA–LNP. Similar to mice, no inflammation or other adverse events were observed. E-protein-specific IgG and nAb were efficiently induced by all three vaccine doses, with no statistically significant differences between groups. Endpoint IgG titres rose to >300,000 in all groups at week 4 and were maintained at ≥100,000 until week 12 (Fig. 3a). PRNT50 nAb titres against MR-766 peaked at about 400 at week 2 (Extended Data Fig. 4a). To facilitate comparison of nAb titres across laboratories, the neutralization curve is shown for a human ZIKV-neutralizing monoclonal antibody, A3594, in the PRNT assay (Extended Data Fig. 5). nAb titres obtained with a focus-reduction neutralization test (FRNT), which has a format similar to PRNT, were stable around 400 against ZIKV MEX I-44 at weeks 2–12 (Fig. 3b). The RVP assay showed nAb titres against H/PF/2013 of around 10,000 at week 2 and around 17,000 at week 4 (Fig. 3c), and a titre of around 3,000 against MR-766 at week 4 (Extended Data Fig. 4b). The neutralization of viruses from both Asian and African lineages is consistent with a previous report demonstrating the existence of only one serotype of ZIKV23. The absence of a significant dose-dependent effect in the antibody response in any assay (Kruskal–Wallis test, P > 0.05) suggests that a low dose of 50 μg (approximately 0.02 mg kg−1) was sufficient, or possibly more than sufficient, to induce robust anti-ZIKV immunity in macaques.
a–c, Rhesus macaques were immunized with 600 μg (n = 4), 200 μg (n = 3) or 50 μg (n = 3) of ZIKV prM–E mRNA–LNP and the antibody response was quantified by ELISA (a), FRNT using ZIKV MEX I-44 (b) and RVP using ZIKV H/PF/2013 (c). Data from pre-challenge (weeks 0–4 in a–c) and unchallenged (weeks 9–12 in a, b) animals are shown. Points represent individual monkeys; dotted lines indicate the limit of detection; horizontal lines indicate the mean. Immune responses in dose groups were compared by Kruskal–Wallis test, P > 0.05 for all comparisons.
Rhesus macaques were challenged at week 5 by subcutaneous injection of 104 TCID50 of ZIKV PRVABC59 in five vaccinated animals and six control animals (Extended Data Table 1). All control animals became infected, with median peak plasma viraemia of 7,000 ZIKV RNA copies per ml (Fig. 4). By contrast, vaccinated macaques were highly protected against ZIKV infection. Four out of five animals—including three that received the lowest dose of 50 μg and one that received the medium dose of 200 μg—had no detectable viraemia (<50 copies per ml) at all time points. We detected a low and transient viral blip of 100 copies per ml at day 3 after challenge in one animal that received the highest dose of 600 μg of ZIKV prM–E mRNA–LNP, representing a 99% reduction in peak viraemia compared to the control animals. This animal exhibited among the lowest nAb titres in multiple assays at week 4 after vaccination: PRNT50 of 36 to MR-766, FRNT50 of 226 to MEX I-44 and RVP titres of 986 to MR-766 and 5,812 to H/PF/2013 (Fig. 3 and Extended Data Fig. 4). The significance of a low-level viral blip in one animal, including implications for a correlate of protection, are as yet uncertain and warrant further study with a greater number of animals.
Six unvaccinated control macaques and five vaccinated macaques that received 50 μg (n = 3), 200 μg (n = 1) or 600 μg (n = 1) of ZIKV mRNA–LNP at week 0 were challenged by subcutaneous injection of 104 TCID50 of ZIKV PRVABC59 at week 5. Viral loads were measured in plasma by qRT–PCR for ZIKV capsid RNA. Dotted lines indicate the threshold beneath which values are below the limit of detection (50 copies per ml), and undetectable values were staggered to show individual animals. Day 3 and 5 viraemia in vaccine and control groups was compared by Mann–Whitney test, P < 0.001.
In this report, we demonstrate that a single, low-dose intradermal immunization with nucleoside-modified ZIKV prM–E mRNA–LNP is protective in both mice and rhesus macaques and elicits higher nAb responses than a single immunization of multiple ZIKV vaccine candidates that have recently been reported, including purified inactivated virus (PIV) and plasmid DNA vaccines encoding prM–E or M–E10,11,12. In mice, PRNT50 nAb increased steadily over several months, rising to levels 50–100 times higher than those induced by a single immunization with PIV or DNA vaccines11,12. The ZIKV mRNA–LNP vaccine conferred complete, rapid and durable protection in mice that was maintained for at least 5 months, and probably much longer, since nAb titres were stable. The challenge studies in mice also showed that ZIKV PRVABC59 replicated more efficiently (P = 0.02, Mann–Whitney test) in 8-week-old BALB/c mice compared to 25-week-old mice, when two identical aliquots and doses of challenge virus stock were used. A previous report has shown that ZIKV-related mortality in immune-competent mice decreases between 1 and 4 weeks of age24, but ZIKV replication in adult mice has not yet been well described.
In rhesus macaques, a single immunization with 50 μg ZIKV prM–E mRNA–LNP induced RVP nAb titres that were 50 times higher than those induced by one immunization with 1 mg DNA vaccine and over two times higher than those induced by two immunizations of DNA12, as measured by the same assay in the same laboratory. ZIKV mRNA–LNP nAb titres may overlap with those elicited by one injection of PIV or RhAd52 ZIKV vaccines in macaques, although differences in assay formats prevent a precise comparison. The FRNT nAb titres elicited by ZIKV mRNA–LNP in macaques were maintained at a stable level until 12 weeks after immunization, suggesting that protection may be durable.
Future studies on the nucleoside-modified ZIKV mRNA–LNP vaccine will allow us to explore the effect of a boost, study the effect of this vaccine on other flavivirus infections and determine the efficacy in preventing fetal ZIKV infection and disease.
Methods
Ethics statement
Animals
The investigators adhered to the Guide for the Care and Use of Laboratory Animals by the Committee on Care of Laboratory Animal Resources Commission on Life Sciences, National Research Council. Mouse studies were conducted under protocols approved by the University of Pennsylvania (UPenn) IACUCs. Rhesus macaques (Macaca mulatta) were housed at Bioqual, Inc. (Rockville, MD). Macaque experiments were reviewed and approved by Bioqual and UPenn Animal Care and Use Committees. All animals were housed and cared for according to local, state and federal policies in an Association for Assessment and Accreditation of Laboratory Animal Care International (AAALAC)-accredited facility.
Human cells
Research involving human cells complied with the Declaration of Helsinki. De-identified leukapheresis cells were obtained from the UPenn Immunology Core under their Institutional Review Board (IRB)-approved protocol, and were deemed exempt by the UPenn IRB.
Antibody reagents
The pan-flavivirus mouse monoclonal antibody 4G2, clone D1-4G2-4-15 (EMD Millipore MAB10216) was used to detect ZIKV E protein by western blot. The following antibodies were used for flow cytometry: anti-CD4 PerCP/Cy5.5 (clone GK1.5, Biolegend), anti-CD3 APC-Cy7 (clone 145-2C11, BD Biosciences), anti-CD27 PE (clone LG.3A10, BD Biosciences), anti-TNF PE-Cy7 (clone MP6-XT22, BD Biosciences), anti-IFNγ AF700 (clone XMG1.2, BD Biosciences), anti-IL-2 APC (clone JES6-5H4, BD Biosciences). The live/dead fixable aqua dead cell stain kit (Life Technologies) was used to discriminate dead cells and debris. The following antibodies were used for ELISA assays: goat anti-mouse IgG HRP (Sigma 4416), goat anti-monkey IgG HRP (Sigma 2054) and ZIKV E-protein-specific monoclonal antibody NR-4747 clone E19 (BEI Resources). ZIKV-neutralizing human monoclonal antibody Ab3594 was provided by the Duke University, Duke-NUS Graduate Medical School, National University of Singapore team of C.E.M, G.D.S., R.P., E. E. Ooi, B.F.H., M. A. Moody, S. Lok, and H.-X. Liao.
Protein reagents
Purified recombinant ZIKV E protein (Aalto Bioreagents AZ 6312) was used in ELISAs to detect E-protein-specific IgG, in western blots as a positive control and in mouse splenocyte stimulation.
mRNA production
mRNA was produced as previously described25 using T7 RNA polymerase on linearized plasmid (pTEV-ZIKVprM-E-A101) encoding codon-optimized26 ZIKV strain H/PF/2013 (Asian lineage, French Polynesia, 2013, GenBank: KJ776791) prM–E glycoproteins. mRNA was transcribed to contain a 101 nucleotide-long poly(A) tail. 1-methylpseudouridine-5′-triphosphate (TriLink) was used instead of UTP to generate modified nucleoside-containing mRNA. mRNA was capped using a m7G capping kit with 2′-O-methyltransferase to obtain cap1 and was purified by a fast protein liquid chromatography (FPLC) method, as described27. mRNA was analysed by agarose gel electrophoresis and stored frozen at −20 °C.
Cell culture
Human embryonic kidney (HEK)293T cells (ATCC) were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 2 mM l-glutamine (Life Technologies) and 10% fetal calf serum (FCS) (HyClone) (complete medium). The HEK293T cell line was tested for mycoplasma contamination after receipt from ATCC and before expansion and cryopreservation. Human dendritic cells were generated from monocytes, as described28, and grown in RPMI1640 medium containing 2 mM l-glutamine (Life Technologies) and 10% FCS (HyClone) (complete medium) supplemented with 50 μg ml−1 recombinant human GM-CSF and 100 μg ml−1 recombinant human IL-4 (R&D systems). Cells were maintained by adding fresh medium containing IL-4 and GM-CSF every 3 days and were used on day 7. Mouse dendritic cells were generated from bone marrow cells obtained from the femurs of animals and grown in complete medium supplemented with 50 μg ml−1 mouse GM-CSF (R&D systems). Cells were maintained by adding fresh medium containing mouse GM-CSF every 3 days and were used on day 7.
mRNA transfection
Transfection of human and mouse dendritic cells and HEK293T cells was performed with TransIT-mRNA (Mirus Bio) according to the manufacturer’s instructions: mRNA (0.3 μg) was combined with TransIT-mRNA reagent (0.34 μl) and boost reagent (0.22 μl) in 17 μl of serum-free medium, and the complex was added to 2 × 105 cells in 183 μl complete medium. Supernatant was collected and cells were lysed for 1 h on ice in RIPA buffer (Sigma) at 18 h after transfection.
Western blot analysis of envelope protein expression
Whole-cell lysates and supernatants from cells transfected with ZIKV prM–E were assayed for ZIKV E protein by non-denaturing SDS–PAGE western blot. Samples were combined with 4 × Laemmli buffer (Bio-Rad) and separated on a 4–15% precast polyacrylamide Criterion TGX gel (Bio-Rad) for 45 min at 200 V. Transfer to PVDF membrane was performed using a semi-dry apparatus (Ellard Instrumentation, Ltd) at 10 V for 1 h. The membrane was blocked with 5% non-fat dry milk in TBS buffer containing 0.5% Tween-20. E protein was detected using 1:10,000 4G2 ascites for 1 h, followed by secondary goat anti-mouse IgG HRP 1:10,000 for 1 h. Antibody incubations were performed at room temperature in blocking buffer. Blots were developed using Luminata Forte substrate (Millipore) and a Kodak X-OMAT 1000A processor. At least 2 independent experiments were performed.
Characterization of E protein in supernatant
Supernatant from HEK293T cells transfected with ZIKV prM–E mRNA was tested for whether E protein could be pelleted and disrupted with detergent, consistent with subviral particles. Supernatant was incubated in PBS alone or PBS with 0.5% Triton X-100 for 1 h on ice. Samples were then spun at 42,000 r.p.m. for 2.5 h in a Beckman TLA-55 rotor. The supernatant was then removed from the pellet, which was resuspended in 50 μl of PBS. Equal volumes of the input, pellet and post-centrifugation supernatant fractions were then analysed by western blot, as described above.
Lipid-nanoparticle encapsulation of the mRNA
FPLC-purified mRNA and polycytidylic acid (poly(C) RNA) (Sigma) were encapsulated in LNPs using a self-assembly process in which an aqueous solution of mRNA at pH 4.0 is rapidly mixed with a solution of lipids dissolved in ethanol29. LNPs used in this study were similar in composition to those described previously29,30, which contain an ionizable cationic lipid (proprietary to Acuitas), phosphatidylcholine, cholesterol and PEG-lipid (with a ratio of 50:10:38.5:1.5 mol/mol) and were encapsulated at an RNA to total lipid ratio of around 0.05 (wt/wt). The LNPs had a diameter of around 80 nm as measured by dynamic light scattering using a Zetasizer Nano ZS instrument (Malvern Instruments Ltd). RNA–LNP formulations were stored at −80 °C at a concentration of RNA of about 1 μg μl−1.
Administration of LNPs to mice and rhesus monkeys
Mice
Female BALB/c and C57BL/6 mice aged 8 weeks were purchased from Charles River Laboratories, and cages of mice were randomly allocated to groups. Power analysis was used to calculate the size of all animal groups to ensure statistically significant results. RNA–LNP was diluted in PBS and injected into animals intradermally with a 3/10cc 29½G insulin syringe (BD Biosciences). Four sites of injection (30 μl each) over the lower back were used.
Monkeys
Ketamine-anaesthetized animals were shaved on their back and injected with mRNA–LNP diluted in PBS. Ten sites of injection (60 μl each) were used. Animals of similar age and weight were allocated to each group.
Blood collection from mice and rhesus macaques
Mice
Blood was collected from the orbital sinus under isoflurane anaesthesia. Blood was centrifuged for 10 min at 13,000 r.p.m. and the serum was stored at −20 °C and used for ELISA and virus neutralization assays. EDTA-plasma was collected to isolate RNA for quantitative PCR with reverse transcription (qRT–PCR) analysis.
Monkeys
Blood was collected by femoral venipuncture under ketamine anaesthesia, and serum and EDTA–plasma were collected and stored at −80 °C for ELISA, neutralization analysis and to isolate RNA for qRT–PCR.
Stimulation and staining of splenocytes
Single-cell suspensions from spleens were made in complete medium. Splenocytes were washed once in PBS and resuspended in complete medium at 2 × 107 cells per ml. Subsequently, 2 × 106 cells (100 μl) per sample were stimulated for 6 h at 37 °C using 2 μg ml−1 purified, recombinant ZIKV E protein. GolgiPlug (brefeldin A, BD Biosciences) and GolgiStop (monensin, BD Biosciences) were diluted 1:100 and 1:143 in complete medium, respectively, and 20 μl from both diluted reagents was added to each sample to inhibit the secretion of intracellular cytokines after 1 h. An unstimulated sample for each animal was included. Samples stimulated with PMA (10 ng ml−1)–ionomycin (250 ng ml−1) (Sigma) were used as positive controls.
After stimulation, cells were washed in PBS and stained using the live/dead fixable aqua dead cell stain kit (Life Technologies) and then surface stained for CD4 and CD27. Antibodies were incubated with cells for 30 min at room temperature. Following surface staining, cells were washed in FACS buffer and fixed using the Cytofix/Cytoperm kit (BD Biosciences) according to the manufacturer’s instructions. Following fixation, the cells were washed in the appropriate perm buffer and incubated with antibodies against CD3, TNF, IFNγ and IL-2 for 1 h at room temperature. Following staining, the cells were washed with the appropriate perm buffer, fixed (PBS containing 1% paraformaldehyde) and stored at 4 °C until analysis. Results are obtained from one technical replicate.
Flow cytometry
Splenocytes were analysed on a modified LSR II flow cytometer (BD Biosciences). One hundred thousand events were collected per sample. After the gates for each function were created, the Boolean gate platform was used to create the full array of possible combination of cytokines, equating to seven response patterns when testing three cytokines. Data were expressed by subtracting the per cent positive unstimulated cells from the per cent positive cells stimulated with E protein.
Enzyme-linked immunosorbent assays (ELISA) for ZIKV E-specific IgG
Immulon 4HXB ELISA plates were coated with 6 μg ml−1 purified, recombinant ZIKV E protein in 0.1 M sodium bicarbonate buffer overnight at 4 °C. The plate was blocked with 2% BSA in PBS for 1 h, and washed three times with wash buffer (PBS with 0.05% Tween-20). Mouse or rhesus macaque serum was diluted in blocking buffer and incubated on the plate for 1 h at room temperature, followed by four washes. Secondary HRP-conjugated antibody was diluted 1:10,000 in blocking buffer and incubated on the plate for 1 h, followed by four washes. TMB substrate (KPL) was applied to the plate and the reaction was stopped with 2 Normal sulfuric acid. The absorbance was measured at 450 nm using an MRX Revelation microplate reader. ZIKV E-protein-specific IgG was analysed in two ways: as an endpoint dilution titre, defined as the highest reciprocal dilution of serum to give an OD greater than the sum of the background plus 0.01 OD units; and as an estimate of the absolute IgG concentration, which was based on the mouse monoclonal antibody NR-4747 as a standard (applicable only to mouse samples). All samples were run in at least technical duplicates.
ZIKV MR-766 plaque reduction neutralization tests (PRNT)
ZIKV strain MR-766 (African lineage, Uganda, 1947, GenBank: AY632535) (UTMB Arbovirus Reference Collection) was produced in Vero cells (ATCC CCL-81) and 50 PFU were incubated with increasing dilutions of heat-inactivated serum in serum-free DMEM (Corning) medium for 1 h at 37 °C. The virus–serum mixture (200 μl) was added to a confluent monolayer of Vero cells in 6-well format and incubated for 1.5 h at 37 °C with intermittent rocking. Then, 3 ml of overlay, containing a final concentration of 0.5% methylcellulose (4,000 centipoise) (Sigma), 1 × DMEM (Gibco), 16 mM HEPES, 0.56% sodium bicarbonate, 1.6× GlutaMAX (Gibco), 1 × penicillin/streptomycin (Corning), and 4 μg ml−1 amphotericin B (Gibco), was added to each well, and plates were incubated for 5 days at 37 °C in 5% CO2. The overlay was aspirated and cells were fixed and stained with 0.5% crystal violet (Sigma) in 25% methanol, 75% deionized water. Wells were rinsed with deionized water to visualize plaques. Neutralization titres (EC50) were determined by plotting a line through the linear portion of the curve that crossed 50% inhibition and calculating the reciprocal dilution of serum required for 50% neutralization of infection. EC50 titres are reported as the mean of one or two technical replicates and values below the limit of detection are reported as half of the limit of detection.
ZIKV MEX I-44 focus reduction neutralization tests (FRNT)
ZIKV MEX I-44 (Asian lineage, Mexico, 2016, GenBank: KX856011) stocks were generated in Vero 76 cells (ATCC CRL-1587) and collected as clarified cell-culture lysate/supernatant. FRNT was performed by combining a standard dose of ZIKV with twofold serial dilutions of heat-inactivated serum for 1 h at 37 °C. Virus–serum mixtures (100 μl) were then inoculated onto Vero 76 monolayers, incubated at 37 °C for 1 h and overlayed with an Avicel (FMC Biopolymer)-containing growth medium. After 3 days of incubation, plates were formalin-fixed, permeabilized, blocked and stained by sequential incubation with a biotin-conjugated 4G2 monoclonal antibody (ATCC HB-112), streptavidin-HRP (BD Biosciences) and TrueBlue peroxidase substrate (KPL). Virus input was verified in parallel (acceptable range: 20–60 foci). FRNT50 (EC50 titres) are reported as the highest reciprocal dilution giving a focus count ≤ the 50% neutralization cutoff, and the geometric mean was computed for technical duplicates.
Reporter virus particle (RVP) production
Pseudo-infectious RVPs were produced by complementation of a GFP-expressing WNV sub-genomic replicon23,31 with a plasmid encoding the viral structural proteins (capsid-prM–E). Briefly, ZIKV MR-766 and ZIKV H/PF/2013 RVPs were produced via co-transfection of HEK293T cells with the structural gene and replicon plasmids (3:1 ratio by mass) using Lipofectamine 3000 per the manufacturer’s protocol (Invitrogen). Transfected cells were incubated at 30 °C and RVP-containing supernatants were collected on days 3–6. Stocks were passed through a 0.2-μm filter and aliquots were stored at −80 °C until use. Stock titres were determined by infecting Raji-DCSIGNR cells with serial dilutions of filtered RVP supernatants. GFP-positive cells were assessed by flow cytometry at 48 h after infection and RVP titres were calculated.
RVP neutralization assay
Previously titred RVPs were diluted to ensure antibody excess at informative points on the dose–response curves and were incubated with serial dilutions of mouse or macaque serum for 1 h at 37 °C to allow for steady-state binding. Raji-DCSIGNR cells were then infected with antibody–RVP complexes in two technical replicates. Infections were carried out at 37 °C and GFP-positive infected cells were detected by flow cytometry 24–48 h later. Neutralization results were analysed by nonlinear regression to estimate the reciprocal dilution of serum required for half-maximal neutralization of infection (EC50 titre) (Prism 6, GraphPad). The initial dilution of serum (on the basis of the final volume of RVPs, cells, and serum) was set as the limit of confidence of the assay. Titres for which nonlinear regression was predicted to be below this threshold were reported as a titre half the limit of confidence. Individual EC50 titres are reported as the geometric mean of at least 2 technical replicates.
Preparation of challenge ZIKV virus
Mice
Challenge ZIKV strain PRVABC59 (Asian lineage, Puerto Rico, 2015, GenBank: KU501215) (BEI Resources NR-50240) was grown in Vero CCL81 cells. A T175 flask of cells at 75–90% confluency was inoculated with an MOI of 0.01 ZIKV in 10 ml of serum-free DMEM medium. The flask was incubated at 37 °C, 5% CO2 for 1.5 h with intermittent rocking, then warmed medium was added to a final concentration of 1.5% FCS, 1 × GlutaMAX (Gibco) and 1 × penicillin/streptomycin (Corning) in a final volume of 25 ml. The flask was incubated for 4 days or until cytopathic effects were visible. Then, the supernatant was collected, clarified by low-speed centrifugation and ultra-centrifuged at 20,000 r.p.m. for 1 h at 4 °C in a Sorvall SureSpin 630 rotor. The supernatant was removed and the pellet was resuspended in 1 ml of serum-free DMEM, aliquoted and stored at −80 °C. Before challenge, virus was thawed and diluted in PBS to 2,000 PFU per ml.
Monkeys
Challenge ZIKV strain PRVABC59 was grown in Vero 76 CRL-1587 cells. T150 flasks of cells at 80–85% confluency were used for propagation. Infection was performed with 100 μl stock virus diluted in 4 ml of fresh L-15 medium (Gibco) supplemented with 10% FCS (Gibco), 10% tryptose phosphate broth (Sigma Aldrich), 1 × penicillin/streptomycin (Gibco) and l-glutamine (Gibco) and adsorbed for 1 h at room temperature with gentle agitation every 15 min. Each flask received 7 ml of fresh L-15 medium after adsorption and was incubated for 4 days at 37 °C. Cellular debris was removed by centrifugation at 1,200 r.p.m. for 5 min at 4 °C in an Eppendorf A-4-62 rotor. Virus stocks were aliquoted and stored at −80 °C.
Zika virus challenge in mice and rhesus macaques
Mice
At 2 or 20 weeks after vaccination, mice were bled and then challenged intravenously with 200 PFU of ZIKV PRVABC59 in 100 μl of PBS. Blood was collected 3 and 7 days after the challenge to determine viral loads (ZIKV RNA copies per ml) in plasma.
Monkeys
Macaques were anaesthetized with ketamine and injected subcutaneously in the hind thigh with 104 TCID50 of ZIKV PRVABC59 in a volume of 1 ml in PBS. Blood was collected 1, 3, 5, and 7 days after the challenge to determine viral loads (ZIKV RNA copies per ml) in plasma.
Viral load quantification (qRT–PCR)
Using blinded samples, RNA was isolated from 200 μl (macaque) or 50 μl (mouse) plasma using the QIAamp MinElute Virus spin kit (Qiagen). Extracted RNA was used for amplification using the SensiFAST Probe Lo-ROX One-Step Kit (Bioline BIO-78005) on a 7500 Real-Time PCR system (Applied Biosystems). Primers and probe were designed to amplify a conserved region of the capsid gene from ZIKV BeH815744, as follows: fwd 5′-GGAAAAAAGAGGCTATGGAAATAATAAAG-3′; rev 5′-CTCCTTCCTAGCATTGATTATTCTCA-3′; probe 5′-AGTTCAAGAAAGATCTGGCTG-3′.
Primers and probe were used at a final concentration of 2 μM, and the following program was run: 48 °C for 30 min, 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 1 min at 60 °C. Assay sensitivity was 50 copies per ml for macaque and 200 copies per ml for mouse samples. Results are calculated from at least two technical replicates.
Statistical analysis
No statistical methods were used to predetermine sample size, unless indicated. The investigators were not blinded to allocation during experiments and outcome assessment unless indicated (qRT–PCR). GraphPad Prism 5.0f was used to perform Mann–Whitney and Kruskal–Wallis (with Dunn’s correction) tests to compare immune responses in vaccinated and control mice and in different dose groups of macaques, respectively. SPICE 5.35 and Microsoft Excel software was used to perform Student’s t-tests to compare T cell responses in vaccinated and control mice.
Data availability
All data and full plasmid sequences are available upon request from the corresponding author.
References
Pierson, T. C. & Graham, B. S. Zika virus: immunity and vaccine development. Cell 167, 625–631 (2016)
Weissman, D. mRNA transcript therapy. Expert Rev. Vaccines 14, 265–281 (2015)
Sahin, U., Karikó, K. & Türeci, Ö. mRNA-based therapeutics—developing a new class of drugs. Nat. Rev. Drug Discov. 13, 759–780 (2014)
Dick, G. W., Kitchen, S. F. & Haddow, A. J. Zika virus. I. Isolations and serological specificity. Trans. R. Soc. Trop. Med. Hyg. 46, 509–520 (1952)
Beck, A. S. & Barrett, A. D. Current status and future prospects of yellow fever vaccines. Expert Rev. Vaccines 14, 1479–1492 (2015)
Jarmer, J. et al. Variation of the specificity of the human antibody responses after tick-borne encephalitis virus infection and vaccination. J. Virol. 88, 13845–13857 (2014)
Guy, B. & Jackson, N. Dengue vaccine: hypotheses to understand CYD-TDV-induced protection. Nat. Rev. Microbiol. 14, 45–54 (2016)
Kim, E. et al. Preventative vaccines for Zika virus outbreak: preliminary evaluation. EBioMedicine 13, 315–320 (2016)
Muthumani, K. et al. In vivo protection against ZIKV infection and pathogenesis through passive antibody transfer and active immunisation with a prMEnv DNA vaccine. npj Vaccines 1, 16021 (2016)
Abbink, P. et al. Protective efficacy of multiple vaccine platforms against Zika virus challenge in rhesus monkeys. Science 353, 1129–1132 (2016)
Larocca, R. A. et al. Vaccine protection against Zika virus from Brazil. Nature 536, 474–478 (2016)
Dowd, K. A. et al. Rapid development of a DNA vaccine for Zika virus. Science 354, 237–240 (2016)
Ledgerwood, J. E. et al. A replication defective recombinant Ad5 vaccine expressing Ebola virus GP is safe and immunogenic in healthy adults. Vaccine 29, 304–313 (2010)
Sumida, S. M. et al. Neutralizing antibodies and CD8+ T lymphocytes both contribute to immunity to adenovirus serotype 5 vaccine vectors. J. Virol. 78, 2666–2673 (2004)
Abbink, P. et al. Construction and evaluation of novel rhesus monkey adenovirus vaccine vectors. J. Virol. 89, 1512–1522 (2015)
Minor, P. D. Live attenuated vaccines: historical successes and current challenges. Virology 479–480, 379–392 (2015)
Baronti, C. et al. Complete coding sequence of zika virus from a French Polynesia outbreak in 2013. Genome Announc. 2, e00500–14 (2014)
Andries, O. et al. N1-methylpseudouridine-incorporated mRNA outperforms pseudouridine-incorporated mRNA by providing enhanced protein expression and reduced immunogenicity in mammalian cell lines and mice. J. Control. Release 217, 337–344 (2015)
Pardi, N. et al. Expression kinetics of nucleoside-modified mRNA delivered in lipid nanoparticles to mice by various routes. J. Control. Release 217, 345–351 (2015)
Wang, P. G. et al. Efficient assembly and secretion of recombinant subviral particles of the four dengue serotypes using native prM and E proteins. PLoS One 4, e8325 (2009)
Weissman, D. et al. HIV gag mRNA transfection of dendritic cells (DC) delivers encoded antigen to MHC class I and II molecules, causes DC maturation, and induces a potent human in vitro primary immune response. J. Immunol. 165, 4710–4717 (2000)
Dudley, D. M. et al. A rhesus macaque model of Asian-lineage Zika virus infection. Nat. Commun. 7, 12204 (2016)
Dowd, K. A. et al. Broadly neutralizing activity of Zika virus-immune sera identifies a single viral serotype. Cell Reports 16, 1485–1491 (2016)
Lazear, H. M. et al. A mouse model of Zika virus pathogenesis. Cell Host Microbe 19, 720–730 (2016)
Pardi, N., Muramatsu, H., Weissman, D. & Karikó, K. In vitro transcription of long RNA containing modified nucleosides. Methods Mol. Biol. 969, 29–42 (2013)
Thess, A. et al. Sequence-engineered mRNA without chemical nucleoside modifications enables an effective protein therapy in large animals. Mol. Ther. 23, 1456–1464 (2015)
Weissman, D., Pardi, N., Muramatsu, H. & Karikó, K. HPLC purification of in vitro transcribed long RNA. Methods Mol. Biol. 969, 43–54 (2013)
Karikó, K. et al. Incorporation of pseudouridine into mRNA yields superior nonimmunogenic vector with increased translational capacity and biological stability. Mol. Ther. 16, 1833–1840 (2008)
Maier, M. A. et al. Biodegradable lipids enabling rapidly eliminated lipid nanoparticles for systemic delivery of RNAi therapeutics. Mol. Ther. 21, 1570–1578 (2013)
Jayaraman, M. et al. Maximizing the potency of siRNA lipid nanoparticles for hepatic gene silencing in vivo . Angew. Chem. Int. Edn Engl. 51, 8529–8533 (2012)
Pierson, T. C. et al. A rapid and quantitative assay for measuring antibody-mediated neutralization of West Nile virus infection. Virology 346, 53–65 (2006)
Acknowledgements
We thank R. Tesh at the UTMB World Reference Center for Emerging Viruses and Arboviruses for providing ZIKV stocks. We gratefully acknowledge the technical or administrative support of M. Bertrand, L. Arwood, C. Sample, M. J. Barr, C. Vivian, T. Gurley and M. A. Moody, E. E. Ooi, S. Lok, H.-X. Liao, S. Awasthi, L. Hook, F. Shaheen, and the UPenn CFAR. The Virology Unit of the Duke Regional Biocontainment Laboratory (Duke Human Vaccine Institute) received support from the NIH, UC6-AI058607. The Duke Center for HIV/AIDS Vaccine Immunology received support from NIH AI100645. D.V. received funding from the Department of Diagnostic Medicine and Pathobiology, College of Veterinary Medicine, Kansas State University. R.S.P., C.R.D., K.A.D., T.C.P. and B.S.G. were funded by the NIAID Division of Intramural Research and the Vaccine Research Center. S.E.H. received funding through NIH U19-AI057229. M.J.Hog. received funding from NIH T32 AI007632-14. D.W. received funding from NIH R01-AI050484, R01-AI124429 and R01-AI084860, and Takeda Pharmaceuticals, New Frontier Science.
Author information
Authors and Affiliations
Contributions
N.P., M.J.Hog., B.F.H., T.C.P., B.S.G., M.G.L. and D.W. designed the studies and evaluated data. N.P. and H.M. conducted mouse studies. M.J.Hog. performed virologic and immunologic assays for mouse and non-human primate studies. D.W., M.J.Hog., N.P. and K.K. developed the mRNA construct and platform. E.W. and S.E.H. provided virologic technical assistance and reagents. V.M.H. provided molecular assay assistance. C.E.M., R.P., G.D.S., R.S.P., C.R.D., K.A.D. and T.C.P. conducted nAb and IgG assays. B.L.M., Y.K.T., T.D.M. and M.J.Hop. developed and provided LNPs. Y.-J.H., D.V. and S.H. provided non-human primate challenge stock. H.A., J.G. and M.W. conducted viral load analysis. M.G.L., L.L.S., R.M.S., W.W. and A.G. carried out the non-human primate immunization and challenge trial. J.-S.Y., S.San., H.B., S.Sah. and M.L. processed non-human primate samples and conducted assays for non-human primate studies. N.P., M.J.Hog. and D.W. wrote the paper with all co-authors.
Corresponding author
Ethics declarations
Competing interests
N.P., M.J.Hog., K.K. and D.W. are named on patents that describe the use of nucleoside-modified mRNA as a platform to deliver therapeutic proteins and vaccines. We have disclosed those interests fully to the University of Pennsylvania, and have an approved plan in place for managing any potential conflicts arising from licensing of our patents.
Additional information
Reviewer Information Nature thanks A. Barrett, G. Screaton and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
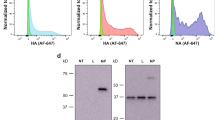
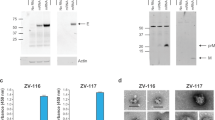
Extended Data Figure 1 Design and characterization of ZIKV prM–E mRNA.
a, The ZIKV mRNA encodes the signal peptide (SP) from MHC class II and pre-membrane (prM) and envelope (E) glycoproteins from ZIKV H/PF/2013. b, mRNA was transfected into HEK293T cells (n = 3), human (n = 3) or mouse dendritic cells (n = 2). E protein expression in cell lysate and supernatant was analysed by western blot, using firefly luciferase-encoding mRNA-transfected cells as a negative control. c, ZIKV mRNA supernatant from transfected HEK293T cells was characterized by ultracentrifugation in the presence and absence of 0.5% Triton X-100, followed by western blot of input (IN), pellet (P) and final supernatant (S) fractions (n = 3).
Extended Data Figure 2 Nucleoside-modified ZIKV mRNA–LNP immunization elicits polyfunctional ZIKV E-protein-specific CD4+ T cell responses.
C57BL/6 mice were immunized with 30 μg of nucleoside-modified ZIKV prM–E mRNA–LNP (n = 8) or control poly(C) RNA–LNP (n = 4). At week 2, antigen-specific CD4+ T cells were detected by intracellular cytokine staining. Graph shows mean ± s.e.m. of the frequencies of combinations of cytokines produced by CD4+ T cells. *P < 0.05 (Student’s t-test).
Extended Data Figure 3 Concentration of ZIKV E-protein-specific IgG in mice.
a, b, Serum from C57BL/6 mice (a; n = 4 control; n = 8 ZIKV mRNA–LNP) or BALB/c mice (b; n = 5 control; n = 10 ZIKV mRNA–LNP) was analysed by ELISA, and estimates of the concentration of ZIKV E-protein-specific IgG were calculated using the mouse monoclonal antibody NR-4747 as a standard. Points represent individual mice; horizontal lines indicate the mean. Responses in vaccine and control groups were compared at each time point by Mann–Whitney test, P < 0.01 for all comparisons.
Extended Data Figure 4 Neutralizing antibody responses against ZIKV MR-766 in macaques immunized with ZIKV prM–E mRNA–LNP.
Serum from immunized macaques was analysed for neutralization of ZIKV MR-766 using the PRNT assay (a) or the RVP assay (b) at the indicated time points. Shaded area indicates values below the limit of detection; horizontal bars indicate mean; symbols indicate individual animals. Immune responses in dose groups were compared by Kruskal–Wallis test, P > 0.05 for all comparisons.
Extended Data Figure 5 Neutralization curve for a human anti-ZIKV neutralizing monoclonal antibody.
ZIKV MR-766 was neutralized by Ab3594, a human ZIKV-neutralizing monoclonal antibody, as a positive control in the PRNT assay. Shown is a representative curve (n = 4). EC50 = 0.026 ± 5.4 μg ml−1 (mean ± s.d.).
Supplementary information
Supplementary Information
This file contains the uncropped Western blot scans with size marker indications. (PDF 241 kb)
Rights and permissions
About this article
Cite this article
Pardi, N., Hogan, M., Pelc, R. et al. Zika virus protection by a single low-dose nucleoside-modified mRNA vaccination. Nature 543, 248–251 (2017). https://doi.org/10.1038/nature21428
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21428
This article is cited by
-
Multivalent cytomegalovirus glycoprotein B nucleoside modified mRNA vaccines did not demonstrate a greater antibody breadth
npj Vaccines (2024)
-
Nobel Prize for physiology or medicine in 2023: how to dupe the cellular innate immune system using modified RNA for therapeutic treatment
Pflügers Archiv - European Journal of Physiology (2024)
-
Engineered exosomes enriched in netrin-1 modRNA promote axonal growth in spinal cord injury by attenuating inflammation and pyroptosis
Biomaterials Research (2023)
-
The use of RNA-based treatments in the field of cancer immunotherapy
Molecular Cancer (2023)
-
mRNA-based vaccines and therapeutics: an in-depth survey of current and upcoming clinical applications
Journal of Biomedical Science (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.