Abstract
With age, haematopoietic stem cells lose their ability to regenerate the blood system, and promote disease development. Autophagy is associated with health and longevity, and is critical for protecting haematopoietic stem cells from metabolic stress. Here we show that loss of autophagy in haematopoietic stem cells causes accumulation of mitochondria and an activated metabolic state, which drives accelerated myeloid differentiation mainly through epigenetic deregulations, and impairs haematopoietic stem-cell self-renewal activity and regenerative potential. Strikingly, most haematopoietic stem cells in aged mice share these altered metabolic and functional features. However, approximately one-third of aged haematopoietic stem cells exhibit high autophagy levels and maintain a low metabolic state with robust long-term regeneration potential similar to healthy young haematopoietic stem cells. Our results demonstrate that autophagy actively suppresses haematopoietic stem-cell metabolism by clearing active, healthy mitochondria to maintain quiescence and stemness, and becomes increasingly necessary with age to preserve the regenerative capacity of old haematopoietic stem cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Niccoli, T. & Partridge, L. Ageing as a risk factor for disease. Curr. Biol. 22, R741–R752 (2012)
Rando, T. A. Stem cells, ageing and the quest for immortality. Nature 441, 1080–1086 (2006)
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013)
Orkin, S. H. & Zon, L. I. Hematopoiesis: an evolving paradigm for stem cell biology. Cell 132, 631–644 (2008)
Kohli, L. & Passegué, E. Surviving change: the metabolic journey of hematopoietic stem cells. Trends Cell Biol. 24, 479–487 (2014)
Geiger, H., de Haan, G. & Florian, M. C. The ageing haematopoietic stem cell compartment. Nature Rev. Immunol. 13, 376–389 (2013)
Ergen, A. V., Boles, N. C. & Goodell, M. A. Rantes/Ccl5 influences hematopoietic stem cell subtypes and causes myeloid skewing. Blood 119, 2500–2509 (2012)
Kusumbe, A. P. et al. Age-dependent modulation of vascular niches for haematopoietic stem cells. Nature 532, 380–384 (2016)
He, C. & Klionsky, D. J. Regulation mechanisms and signaling pathways of autophagy. Annu. Rev. Genet. 43, 67–93 (2009)
Liu, F. et al. FIP200 is required for the cell-autonomous maintenance of fetal hematopoietic stem cells. Blood 116, 4806–4814 (2010)
Mortensen, M. et al. The autophagy protein Atg7 is essential for hematopoietic stem cell maintenance. J. Exp. Med. 208, 455–467 (2011)
Leveque-El Mouttie, L. et al. Autophagy is required for stem cell mobilization by G-CSF. Blood 125, 2933–2936 (2015)
Warr, M. R. et al. FOXO3A directs a protective autophagy program in haematopoietic stem cells. Nature 494, 323–327 (2013)
Rubinsztein, D. C., Mariño, G. & Kroemer, G. Autophagy and aging. Cell 146, 682–695 (2011)
García-Prat, L. et al. Autophagy maintains stemness by preventing senescence. Nature 529, 37–42 (2016)
Levine, B., Mizushima, N. & Virgin, H. W. Autophagy in immunity and inflammation. Nature 469, 323–335 (2011)
Codogno, P., Mehrpour, M. & Proikas-Cezanne, T. Canonical and non-canonical autophagy: variations on a common theme of self-eating? Nature Rev. Mol. Cell Biol. 13, 7–12 (2011)
Youle, R. J. & Narendra, D. P. Mechanisms of mitophagy. Nature Rev. Mol. Cell Biol. 12, 9–14 (2011)
Goldberg, M. S. et al. Parkin-deficient mice exhibit nigrostriatal deficits but not loss of dopaminergic neurons. J. Biol. Chem. 278, 43628–43635 (2003)
Vannini, N. et al. Specification of haematopoietic stem cell fate via modulation of mitochondrial activity. Nature Commun. 7, 13125 (2016)
Ito, K. et al. Self-renewal of a purified Tie2+ hematopoietic stem cell population relies on mitochondrial clearance. Science 354, 1156–1160 (2016)
Mizushima, N., Yamamoto, A., Matsui, M., Yoshimori, T. & Ohsumi, Y. In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol. Biol. Cell 15, 1101–1111 (2004)
Mohrin, M. et al. Stem cell aging. A mitochondrial UPR-mediated metabolic checkpoint regulates hematopoietic stem cell aging. Science 347, 1374–1377 (2015)
Rodgers, J. T. et al. mTORC1 controls the adaptive transition of quiescent stem cells from G0 to GAlert . Nature 510, 393–396 (2014)
Flach, J. et al. Replication stress is a potent driver of functional decline in ageing haematopoietic stem cells. Nature 512, 198–202 (2014)
Chandel, N. S., Jasper, H., Ho, T. T. & Passegué, E. Metabolic regulation of stem cell function in tissue homeostasis and organismal ageing. Nature Cell Biol. 18, 823–832 (2016)
Owusu-Ansah, E. & Banerjee, U. Reactive oxygen species prime Drosophila haematopoietic progenitors for differentiation. Nature 461, 537–541 (2009)
Chambers, S. M. et al. Hematopoietic fingerprints: an expression database of stem cells and their progeny. Cell Stem Cell 1, 578–591 (2007)
Kohli, R. M. & Zhang, Y. TET enzymes, TDG and the dynamics of DNA demethylation. Nature 502, 472–479 (2013)
Fuso, A., Cavallaro, R. A., Orrù, L., Buttarelli, F. R. & Scarpa, S. Gene silencing by S-adenosylmethionine in muscle differentiation. FEBS Lett. 508, 337–340 (2001)
Owen, M. R., Doran, E. & Halestrap, A. P. Evidence that metformin exerts its anti-diabetic effects through inhibition of complex 1 of the mitochondrial respiratory chain. Biochem. J. 348, 607–614 (2000)
Puleston, D. J. et al. Autophagy is a critical regulator of memory CD8+ T cell formation. eLife 3, e03706 (2014)
Tang, A. H. & Rando, T. A. Induction of autophagy supports the bioenergetic demands of quiescent muscle stem cell activation. EMBO J. 33, 2782–2797 (2014)
Kühn, R., Schwenk, F., Aguet, M. & Rajewsky, K. Inducible gene targeting in mice. Science 269, 1427–1429 (1995)
Hara, T. et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441, 885–889 (2006)
Pietras, E. M. et al. Functionally distinct subsets of lineage-biased multipotent progenitors control blood production in normal and regenerative conditions. Cell Stem Cell 17, 35–46 (2015)
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 (2003)
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015)
Akalin, A. et al. Base-pair resolution DNA methylation sequencing reveals profoundly divergent epigenetic landscapes in acute myeloid leukemia. PLoS Genet. 8, e1002781 (2012)
Akalin, A. et al. methylKit: a comprehensive R package for the analysis of genome-wide DNA methylation profiles. Genome Biol. 13, R87 (2012)
Meldi, K. et al. Specific molecular signatures predict decitabine response in chronic myelomonocytic leukemia. J. Clin. Invest. 125, 1857–1872 (2015)
Acknowledgements
We thank J. Debnath for help with these studies, A. Brunet and S. Villeda for providing some old C57Bl/6 mice, J. Cox and D. Ruggero for the gift of Park2−/− mice and Ink128, respectively, S. Y. Zhang for overall assistance, J. Wong for help with Seahorse studies, J. Wong for electron microscopy analyses, J. Pollack for assistance with bioinformatics, P. Nyugen for contribution to confocal imaging, M. Lee for management of the Flow Cytometry Core Facility, and all members of the Passegué laboratory for insights and suggestions. T.T.H. is supported by an AHA Predoctoral Fellowship and T32GM008284, M.R.W by a LLS Special Fellow Award, E.A. by T32AG000114, and E.V. by a Netherlands Organisation for Scientific Research (NWO) Rubicon Fellowship and a BD Biosciences Stem Cell grant. This work was supported by National Institutes of Health R01HL126947 to M.E.F., and National Institutes of Health R01CA184014 and P30DK063720, a Program for Breakthrough Biomedical Research New Frontier Research Award, a Glenn Foundation Research Award, and a Leukemia & Lymphoma Society Scholar Award to E.P.
Author information
Authors and Affiliations
Contributions
T.T.H. performed all of the experiments with help from M.R.W. for the initial Atg12cKO mice analyses, E.R.A. and M.E.F. for DNA methylation studies, O.M.L. and E.V.V. for technical assistance, and J.F. for O-propargyl-puromycin experiments. T.T.H., M.R.W., and E.P. designed the experiments and interpreted the results. T.T.H. and E.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Characterization of autophagy-deficient Atg12cKO mice.
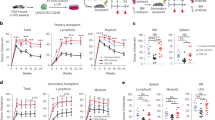
a, Complete blood count analyses of lymphocytes and haematocrit in control (Cnt) and Atg12cKO (cKO) mice after pIC treatment. b, Lineage distribution in peripheral blood of control and cKO mice at 2 months after pIC, and in young and old mice. My, myeloid; Ly, lymphoid. c, Total cell numbers in peripheral blood, spleen (Spl), and bone marrow of control and cKO mice at 2 months after pIC. d, Gating strategy for mature populations. ImGr, imature granulocytes/monocytes; Gr, granulocyte; B, B cells. e, f, Mature populations in (e) bone marrow and (f) spleen of control and cKO mice at 2 months after pIC. g, Gating strategy for immature bone marrow populations. Lin, lineage negative; MP, myeloid progenitors; CMP, common myeloid progenitor; MEP, megakaryocyte/erythrocyte progenitor. h, Quantification of immature bone marrow populations in control and cKO mice at 2 months after pIC. CLP, common lymphoid progenitor. i, HSC frequency over time in control and cKO mice after pIC. j, Quantification of MP bone marrow populations in control and cKO mice at 2 months after pIC. k, Colony formation in methylcellulose from control and cKO bone marrow at 2 months after pIC. Data are mean ± s.d. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001.
Extended Data Figure 2 Characterization of autophagy-deficient Atg12cKO mice and regenerative capacity of Atg12cKO HSCs.
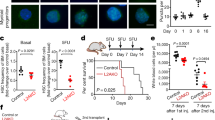
a–e, Haematopoietic features of control (Cnt-5) and Atg5cKO (cKO-5) mice after pIC treatment: (a) scheme for deleting Atg5 in the adult blood system, and (b) neutrophil counts in peripheral blood, (c) total cell numbers in bone marrow and spleen, and quantification of (d) MP and (e) immature bone marrow populations at 2 months after pIC. f, Scheme for control and cKO HSC primary (1°) and secondary (2°) transplantation (tplx). g, Engraftment of young (Y) and old (O) HSCs with donor chimaerism in peripheral blood over time (left), and lineage distribution in peripheral blood (centre) with HSC chimaerism (right) at the indicated time after transplantation. h, Scheme for ageing recipients transplanted with non-pIC-treated control and cKO bone marrow cells and subsequently deleted for Atg12. i, Atg12 deletion in recipients transplanted with 2 × 106 bone marrow cells from 2- or 24-month-old non-pIC-treated cKO donors with donor chimaerism in peripheral blood after PBS or pIC (left, ± s.e.m.), and lineage distribution in peripheral blood at 125 d after transplantation (right). Data are mean ± s.d. except where indicated. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001.
Extended Data Figure 3 Altered biology of Atg12cKO HSCs and characterization of Park2−/− mice.
a, Representative FACS plot of MTG staining in control and cKO HSCs. b, Representative electron micrographs depicting expanded small vesicles (top) and endoplasmic reticulum/Golgi (bottom) in control and cKO HSCs. Scale bar, 1 μm. c, Representative FACS plot and quantification of endoplasmic reticulum mass measured by ER-Tracker flow cytometry staining in control and cKO HSCs. d, e, Representative immunofluorescence images and quantification of (d) LAMP1 and (e) KDEL staining in control and cKO HSCs. Scale bars, 10 μm. f, Levels of p62 in control and cKO HSCs. g, Mitochondria parameters in Park2+/+ and Park2−/− HSCs. h–m, Characterization of Park2−/− mice: (h) scheme for analyses, (i) complete blood count parameters (left) and lineage distribution in peripheral blood (right), (j) bone marrow total cell numbers, (k) bone marrow mature populations, and (l, m) bone marrow immature populations. n, Transplantation of Park2−/− HSCs showing donor chimaerism (left) and lineage distribution (centre) in peripheral blood, and HSC chimaerism (right) at the indicated time after transplantation in primary and secondary recipients. Data are mean ± s.d. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001.
Extended Data Figure 4 Comparison between aHSCs, Atg12cKO HSCs and oHSCs.
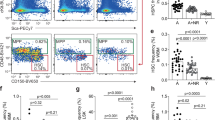
a, Representative FACS plot of pS6 levels in HSC cultured for 1 h (hr) with or without cytokines (±cyt). b, Representative FACS plot of GFP–LC3 in freshly isolated HSCs (t = 0) and HSCs cultured for 6 h with or without cytokines. c, Inactivation of autophagy in +cyt measured by p62 levels. d, Drug modulation of autophagy levels in Gfp–Lc3 HSCs cultured for 3 h with or without cytokines. Rap, rapamycin; I128, Ink128; CC, compound C. Results are expressed as percentage GFP–LC3 levels upon 3 h culture in +cyt conditions. e, Cell size measured by forward scatter (FSC) in control and cKO HSCs, and yHSCs and oHSCs. f, g, Glycolysis activity measured by extracellular acidification rate (ECAR) in (f) control and cKO LSKs, and (g) yHSCs and oHSCs. Oligo, oligomycin; 2-DG, 2-deoxy-d-glucose. h, Mitochondria parameters in yHSCs and oHSCs. i, ROS levels in aHSCs. j, Representative FACS plot and quantification of ROS levels in control and cKO HSCs. k, Representative examples of three independent experiments showing immunofluorescence co-staining of γH2AX with fibrillarin (FBN), 53BP1, and RAD51 in control and cKO HSCs. Sscale bar, 10 μm. Data are mean ± s.d. except for line graphs, and are expressed relative to 0 h HSC (c, i), control HSC (e, j), or yHSC (e, h) levels. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001.
Extended Data Figure 5 Properties of autophagy-deficient and oHSCs, and effect of ROS scavenging in Atg12cKO mice.
a–c, Characteristics of control and cKO HSCs: (a) apoptosis measured by cleaved caspase 3 (CC3) activity, (b) protein synthesis measured by O-propargyl-puromycin (OP-puro) immunofluorescence staining, and (c) cycling activity measured by EdU incorporation. d, e, Characteristics of yHSCs and oHSCs: (d) ROS levels and (e) protein synthesis with representative OP-puro immunofluorescence staining (left) and quantification (right). f, Scheme for NAC in vitro treatment and representative example of colony formation in methylcellulose from NAC-treated control and cKO HSCs. CFU, colony-forming unit; Mk(or)E and G(or)M, mature megakaryocyte, erythroid, granulocyte, or macrophage colonies; GM/Mix, immature GM or GMMkE colonies. Results are expressed as percentage of plated cells. g–i, Scavenging ROS levels in Atg12cKO mice: (g) scheme for NAC in vivo treatment after pIC deletion of Atg12, (h) neutrophil counts (left) and lineage distribution (right) in peripheral blood, and (i) quantification of immature bone marrow populations. Data are mean ± s.d., and are expressed relative to control HSC (a, b), or yHSC (d, e) levels. *P ≤ 0.05, **P ≤ 0.01.
Extended Data Figure 6 Differential gene expression in Atg12cKO HSCs and GMPs, and regulation of DNA methylation in HSCs.
a, Heatmap of differentially expressed genes (DEG) in cKO versus control HSC microarrays (n = 4). b, Volcano plot of DEGs in cKO versus control HSCs. c, Gene set enrichment analyses (GSEA) of DEGs in cKO versus control HSCs: NES: normalized enrichment score. d, Heatmap of DEGs in cKO versus control GMP microarrays (n = 4). e, SAM levels in control and cKO HSCs, and aHSCs. f, aKG levels in c-Kit-enriched control and cKO bone marrow (left, P = 0.0672), and freshly isolated and activated c-Kit-enriched bone marrow (right). Data are mean ± s.d. *P ≤ 0.05, ***P ≤ 0.001.
Extended Data Figure 7 Additional analyses of old Gfp–Lc3 HSCs.
a, Lineage distribution in peripheral blood (left) and frequency of the indicated populations (right) in young and old Gfp–Lc3 mice (related to Fig. 5a, b). b, pS6 levels in yHSCs and oHSCs. c, d, Representative FACS plots and quantification of Cyto-ID dye levels in (c) HSCs cultured for 3 h with or without cytokines, and (d) yHSCs and oHSCs. e, Representative FACS plots of autophagy low (ATlo) and autophagy high (AThi) oHSC subsets. f, Representative examples of three independent experiments showing immunofluorescence staining of FOXO3a in ATlo and AThi oHSCs. Scale bar, 10 μm. g, Representative electron micrographs of oHSCs with or without autophagosomes (AP) showing vesicles (top) and endoplasmic reticulum/Golgi (bottom) compartments. Scale bar, 1 μm. h, Representative immunofluorescence staining and quantification of TOM20 in ATlo and AThi oHSCs. Scale bar, 10 μm. i, j, Characteristics of ATlo and AThi oHSCs: (i) TMRE levels, (j) ATP levels, and (k) ROS levels measured by Mitosox Red (MSR) staining. l, Expansion of ATlo and AThi oHSCs in self-renewal culture conditions. m, Colony formation in methylcellulose from ATlo and AThi oHSCs. Results are expressed as percentage of 100 plated cells. Data are mean ± s.d., and are expressed relative to +cyt HSC (c), yHSC (b, d), or ATlo oHSC (h–j, l) levels. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001.
Extended Data Figure 8 Functionality of ATlo and AThi oHSC subsets.
a, Scheme for primary and secondary transplantations of ATlo or AThi yHSCs and oHSCs. b, Transplantation of ATlo and AThi oHSC subsets with 15% GFP–LC3 high/low expression cutoff showing donor chimaerism (left) and lineage distribution (centre) in peripheral blood, and HSC chimaerism (right) at the indicated times after transplantation in primary (top row) and secondary (bottom row) recipients. c, GFP–LC3 levels in ATlo and AThi oHSCs before transplantation, and in primary and secondary recipients. Results are expressed relative to GFP–LC3 levels in yHSCs (± s.e.m.). d, Transplantation of ATlo and AThi yHSC subsets with 33% GFP–LC3 high/low expression cutoff showing donor chimaerism (left) and lineage distribution (centre) in peripheral blood, and HSC chimaerism (right) at the indicated times after transplantation in primary recipients. Data are mean ± s.d. except when indicated. *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001.
Extended Data Figure 9 Autophagy capability of ATlo and AThi oHSC subsets.
a, Representative FACS plot and quantification of CD150 levels in ATlo and AThi oHSCs. b, Levels of pS6 measured by flow cytometry in HSCs cultured with or without cytokines for the indicated times. Results are normalized to IgG levels. c, Drug modulation of autophagy levels in Gfp–Lc3 oHSCs cultured for 3 h with or without cytokines. I128, Ink128; CC, compound C. Results are expressed as percentage GFP–LC3 levels upon 3 h culture in +cyt conditions. d, Scheme for the stress experiments assessing the autophagy capability of ATlo and AThi oHSCs. e, f, Response to cytokine starvation with (e) GFP–LC3 levels in freshly isolated (t = 0) and upon 6 h culture in ±cyt conditions, and (f) autophagy flux after 6 h culture in ±cyt ±BafA conditions. Percentage flux is calculated as [100 × (1 − (−BafA/+BafA))]. g, Response to glutamine (glu) deprivation. Data are mean ± s.d. and are expressed relative to ATlo oHSC (a, e) or freshly isolated (t = 0) oHSC (b) levels, or relative to +glu conditions (g). ***P ≤ 0.001.
Extended Data Figure 10 Model for the role of autophagy in HSC function and HSC ageing.
HSC activation is accompanied by mitochondria activation and a shift in metabolic activity from glycolysis to OXPHOS, which provides energy and increases the production of mitochondrial metabolites such as α-ketoglutarate (αKG) that act as substrates/co-factors for epigenetic enzymes. Metabolically, aHSCs are poised to undergo lineage priming and produce differentiated progeny to regenerate the blood system. However, aHSCs must also return to quiescence to maintain the stem cell pool. In this context, autophagy plays an essential role by clearing active mitochondria to allow OXPHOS-driven HSCs to efficiently revert to a mostly glycolysis-based metabolic quiescence. Without autophagy, HSCs display an overactive OXPHOS-driven metabolism that promotes myeloid-biased differentiation and loss of stemness as a consequence of epigenetic reprogramming. Other mechanisms of mitochondria elimination probably allow some autophagy-deficient HSCs to return to quiescence during homeostasis, but they do not substitute for autophagy in maintaining HSC function in conditions of intense regeneration stress such as transplantation. This role of autophagy becomes even more important with age as the inability of about two-thirds of oHSCs to activate autophagy results in an overactive OXPHOS metabolism that impairs self-renewal, promotes proliferation and myeloid differentiation, and contributes to replication stress. These unhealthy oHSCs drive most of the ageing blood phenotypes. In contrast, about one-third of oHSCs activate autophagy, control their metabolic activity, and are the fittest old stem cells that retain functional abilities in an adverse ageing bone marrow microenvironment. As all oHSCs remain competent for autophagy induction, it will be exciting to test whether rejuvenation interventions aimed at activating autophagy in unhealthy autophagy-inactivated oHSCs will improve the health of the ageing blood system.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-5. (PDF 265 kb)
Source data
Rights and permissions
About this article
Cite this article
Ho, T., Warr, M., Adelman, E. et al. Autophagy maintains the metabolism and function of young and old stem cells. Nature 543, 205–210 (2017). https://doi.org/10.1038/nature21388
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21388
This article is cited by
-
Autophagy induces hair follicle stem cell activation and hair follicle regeneration by regulating glycolysis
Cell & Bioscience (2024)
-
Characterization of gene regulatory networks underlying key properties in human hematopoietic stem cell ontogeny
Cell Regeneration (2024)
-
Autophagy regulates the maturation of hematopoietic precursors in the embryo
Nature Communications (2024)
-
Made to order: emergency myelopoiesis and demand-adapted innate immune cell production
Nature Reviews Immunology (2024)
-
mTOR pathway occupies a central role in the emergence of latent cancer cells
Cell Death & Disease (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.