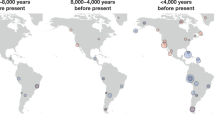
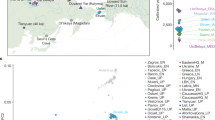
Abstract
Advances in the sequencing and the analysis of the genomes of both modern and ancient peoples have facilitated a number of breakthroughs in our understanding of human evolutionary history. These include the discovery of interbreeding between anatomically modern humans and extinct hominins; the development of an increasingly detailed description of the complex dispersal of modern humans out of Africa and their population expansion worldwide; and the characterization of many of the genetic adaptions of humans to local environmental conditions. Our interpretation of the evolutionary history and adaptation of humans is being transformed by analyses of these new genomic data.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Vigilant, L., Stoneking, M., Harpending, H., Hawkes, K. & Wilson, A. C. African populations and the evolution of human mitochondrial DNA. Science 253, 1503–1507 (1991).
Nordborg, M. On the probability of Neanderthal ancestry. Am. J. Hum. Genet. 63, 1237–1240 (1998).
Meltzer, D. J. First Peoples in a New World (University of California Press, 2009).
Ammerman, A. J. & Biagi, P. The Widening Harvest (Archaeological Institute of America, 2003).
Rasmussen, M. et al. Ancient human genome sequence of an extinct Palaeo-Eskimo. Nature 463, 757–762 (2010). This paper presents the first whole-genome sequence of an ancient human and demonstrates that there is a lack of genetic continuity between Arctic populations.
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010). This paper reports the first whole-genome sequence of a Neanderthal and supplies the first convincing evidence for introgression between humans and Neanderthals.
White, T. D. et al. Pleistocene Homo sapiens from Middle Awash, Ethiopia. Nature 423, 742–747 (2003).
McDougall, I., Brown, F. H. & Fleagle, J. G. Stratigraphic placement and age of modern humans from Kibish, Ethiopia. Nature 433, 733–736 (2005).
Grün, R. et al. U-series and ESR analyses of bones and teeth relating to the human burials from Skhul. J. Hum. Evol. 49, 316–334 (2005).
Liu, W. et al. The earliest unequivocally modern humans in southern China. Nature 526, 696–699 (2015).
Higham, T. et al. The timing and spatiotemporal patterning of Neanderthal disappearance. Nature 512, 306–309 (2014).
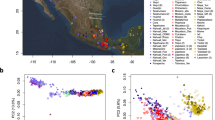
Rosenberg, N. A. et al. Genetic structure of human populations. Science 298, 2381–2385 (2002). This study demonstrates that the human population structure worldwide is correlated with geography and human history.
Tishkoff, S. A. et al. The genetic structure and history of Africans and African Americans. Science 324, 1035–1044 (2009). This article is a comprehensive investigation of genetic variation in Africa.
Busby, G. B. et al. Admixture into and within sub-Saharan Africa. eLife 5, e15266 (2016).
Pickrell, J. K. et al. The genetic prehistory of southern Africa. Nature Commun. 3, 1143 (2012).
Gurdasani, D. et al. The African Genome Variation Project shapes medical genetics in Africa. Nature 517, 327–332 (2015).
Bryc, K. et al. Genome-wide patterns of population structure and admixture in West Africans and African Americans. Proc. Natl Acad. Sci. USA 107, 786–791 (2010).
Schlebusch, C. M. et al. Genomic variation in seven Khoe-San groups reveals adaptation and complex African history. Science 338, 374–379 (2012).
Beltrame, M. H., Rubel, M. A. & Tishkoff, S. A. Inferences of African evolutionary history from genomic data. Curr. Opin. Genet. Dev. 41, 159–166 (2016).
Gronau, I., Hubisz, M. J., Gulko, B., Danko, C. G. & Siepel, A. Bayesian inference of ancient human demography from individual genome sequences. Nature Genet. 43, 1031–1034 (2011).
Schlebusch, C. M. & Soodyall, H. Extensive population structure in San, Khoe, and mixed ancestry populations from southern Africa revealed by 44 short 5-SNP haplotypes. Hum. Biol. 84, 695–724 (2012).
Veeramah, K. R. et al. An early divergence of KhoeSan ancestors from those of other modern humans is supported by an ABC-based analysis of autosomal resequencing data. Mol. Biol. Evol. 29, 617–630 (2012).
Tishkoff, S. A. et al. History of click-speaking populations of Africa inferred from mtDNA and Y chromosome genetic variation. Mol. Biol. Evol. 24, 2180–2195 (2007).
Stringer, C. Human evolution: out of Ethiopia. Nature 423, 692–695 (2003).
Hammer, M. F., Woerner, A. E., Mendez, F. L., Watkins, J. C. & Wall, J. D. Genetic evidence for archaic admixture in Africa. Proc. Natl Acad. Sci. USA 108, 15123–15128 (2011).
Hsieh, P. et al. Model-based analyses of whole-genome data reveal a complex evolutionary history involving archaic introgression in Central African Pygmies. Genome Res. 26, 291–300 (2016).
Lachance, J. et al. Evolutionary history and adaptation from high-coverage whole-genome sequences of diverse African hunter-gatherers. Cell 150, 457–469 (2012).
Hsieh, P. et al. Whole-genome sequence analyses of Western Central African Pygmy hunter-gatherers reveal a complex demographic history and identify candidate genes under positive natural selection. Genome Res. 26, 279–290 (2016).
Ramachandran, S. et al. Support from the relationship of genetic and geographic distance in human populations for a serial founder effect originating in Africa. Proc. Natl Acad. Sci. USA 102, 15942–15947 (2005).
Jakobsson, M. et al. Genotype, haplotype and copy-number variation in worldwide human populations. Nature 451, 998–1003 (2008).
Clark, J. D. et al. Stratigraphic, chronological and behavioural contexts of Pleistocene Homo sapiens from Middle Awash, Ethiopia. Nature 423, 747–752 (2003).
Pickrell, J. K. et al. Ancient west Eurasian ancestry in southern and eastern Africa. Proc. Natl Acad. Sci. USA 111, 2632–2637 (2014).
Henn, B. M. et al. Hunter-gatherer genomic diversity suggests a southern African origin for modern humans. Proc. Natl Acad. Sci. USA 108, 5154–5162 (2011).
Lahr, M. M. & Foley, R. A. Towards a theory of modern human origins: geography, demography, and diversity in recent human evolution. Am. J. Phys. Anthropol. 27 (suppl.), 137–176 (1998).
Groucutt, H. S. et al. Rethinking the dispersal of Homo sapiens out of Africa. Evol. Anthropol. 24, 149–164 (2015).
Pugach, I., Delfin, F., Gunnarsdottir, E., Kayser, M. & Stoneking, M. Genome-wide data substantiate Holocene gene flow from India to Australia. Proc. Natl Acad. Sci. USA 110, 1803–1808 (2013).
Reed, F. A. & Tishkoff, S. A. African human diversity, origins and migrations. Curr. Opin. Genet. Dev. 16, 597–605 (2006).
Quintana-Murci, L. et al. Genetic evidence of an early exit of Homo sapiens sapiens from Africa through eastern Africa. Nature Genet. 23, 437–441 (1999).
Gravel, S. et al. Demographic history and rare allele sharing among human populations. Proc. Natl Acad. Sci. USA 108, 11983–11988 (2011).
Li, H. & Durbin, R. Inference of human population history from individual whole-genome sequences. Nature 475, 493–496 (2011).
Harris, K. & Nielsen, R. Inferring demographic history from a spectrum of shared haplotype lengths. PLoS Genet. 9, e1003521 (2013).
Schiffels, S. & Durbin, R. Inferring human population size and separation history from multiple genome sequences. Nature Genet. 46, 919–925 (2014).
Scally, A. & Durbin, R. Revising the human mutation rate: implications for understanding human evolution. Nature Rev. Genet. 13, 745–753 (2012).
Malaspinas, A. S. et al. A genomic history of Aboriginal Australia. Nature 538, 207–214 (2016).
Mallick, S. et al. The Simons Genome Diversity Project: 300 genomes from 142 diverse populations. Nature 538, 201–206 (2016).
Pagani, L. et al. Genomic analyses inform on migration events during the peopling of Eurasia. Nature 538, 238–242 (2016).
Rasmussen, M. et al. An Aboriginal Australian genome reveals separate human dispersals into Asia. Science 334, 94–98 (2011).
Prüfer, K. et al. The complete genome sequence of a Neanderthal from the Altai Mountains. Nature 505, 43–49 (2014).
Wall, J. D. et al. Higher levels of neanderthal ancestry in East Asians than in Europeans. Genetics 194, 199–209 (2013).
Vernot, B. & Akey, J. M. Complex history of admixture between modern humans and Neandertals. Am. J. Hum. Genet. 96, 448–453 (2015).
Sankararaman, S. et al. The genomic landscape of Neanderthal ancestry in present-day humans. Nature 507, 354–357 (2014).
Sankararaman, S., Patterson, N., Li, H., Pääbo, S. & Reich, D. The date of interbreeding between Neandertals and modern humans. PLoS Genet. 8, e1002947 (2012).
Vernot, B. & Akey, J. M. Resurrecting surviving Neandertal lineages from modern human genomes. Science 343, 1017–1021 (2014).
Kim, B. Y. & Lohmueller, K. E. Selection and reduced population size cannot explain higher amounts of Neandertal ancestry in East Asian than in European human populations. Am. J. Hum. Genet. 96, 454–461 (2015).
Fu, Q. et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature 524, 216–219 (2015).
Skoglund, P. et al. Origins and genetic legacy of Neolithic farmers and hunter-gatherers in Europe. Science 336, 466–469 (2012). The first multi-individual study of ancient genomes; it supports the idea that migration drove the Neolithic transition in Europe.
Lazaridis, I. et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513, 409–413 (2014).
Haak, W. et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522, 207–211 (2015). This study uses SNP data from 69 Europeans who lived 3–8 kyr ago to point to the origin of Indo-European languages.
Gamba, C. et al. Genome flux and stasis in a five millennium transect of European prehistory. Nature Commun. 5, 5257 (2014).
Allentoft, M. E. et al. Population genomics of Bronze Age Eurasia. Nature 522, 167–172 (2015). This study uses the sequencing of 101 ancient genomes to identify population movements in the Bronze Age.
Günther, T. & Jakobsson, M. Genes mirror migrations and cultures in prehistoric Europe — a population genomic perspective. Curr. Opin. Genet. Dev. 41, 115–123 (2016).
Benazzi, S. et al. Early dispersal of modern humans in Europe and implications for Neanderthal behaviour. Nature 479, 525–528 (2011).
Fu, Q. et al. The genetic history of Ice Age Europe. Nature 534, 200–205 (2016). By sequencing the genomes of 51 Eurasian people who lived 7–45 kyr ago, this study finds turnover in the population composition of Europeans.
Asouti, E. & Fuller, D. Q. A contextual approach to the emergence of agriculture in Southwest Asia. Curr. Anthropol. 54, 299–345 (2013).
Günther, T. et al. Ancient genomes link early farmers from Atapuerca in Spain to modern-day Basques. Proc. Natl Acad. Sci. USA 112, 11917–11922 (2015).
Skoglund, P. et al. Genomic diversity and admixture differs for Stone-Age Scandinavian foragers and farmers. Science 344, 747–750 (2014).
Cohen, M. N. & Armelagos, G. J. (eds) Paleopathology at the Origins of Agriculture (Academic, 1984).
Jones, E. R. et al. Upper Palaeolithic genomes reveal deep roots of modern Eurasians. Nature Commun. 6, 8912 (2015).
Bouckaert, R. et al. Mapping the origins and expansion of the Indo-European language family. Science 337, 957–960 (2012).
Novembre, J. et al. Genes mirror geography within Europe. Nature 456, 98–101 (2008). This study identifies fine-scale population structure in Europe that is highly correlated with geography.
Lao, O. et al. Correlation between genetic and geographic structure in Europe. Curr. Biol. 18, 1241–1248 (2008).
Auton, A. et al. Global distribution of genomic diversity underscores rich complex history of continental human populations. Genome Res. 19, 795–803 (2009).
Raghavan, M. et al. Upper Palaeolithic Siberian genome reveals dual ancestry of Native Americans. Nature 505, 87–91 (2014). This study sequences the genome of a 24-kyr-old Eurasian individual and shows that modern Native Americans are the descendants of a population admixed between Western Eurasians (represented by the sequenced sample) and an ancestral East Asian population.
Fu, Q. et al. Genome sequence of a 45,000-year-old modern human from western Siberia. Nature 514, 445–449 (2014). This study sequences DNA from a 45 kyr old Siberian person who was more closely related to modern Asians than to modern Europeans.
Seguin-Orlando, A. et al. Genomic structure in Europeans dating back at least 36,200 years. Science 346, 1113–1118 (2014).
Clarkson, C. et al. The archaeology, chronology and stratigraphy of Madjedbebe (Malakunanja II): a site in northern Australia with early occupation. J. Hum. Evol. 83, 46–64 (2015).
O'Connell, J. F. & Allen, J. The process, biotic impact, and global implications of the human colonization of Sahul about 47,000 years ago. J. Archaeol. Sci. 56, 73–84 (2015).
Thorne, A. G. & Wolpoff, M. H. Regional continuity in Australasian Pleistocene hominid evolution. Am. J. Phys. Anthropol. 55, 337–349 (1981).
Wollstein, A. et al. Demographic history of Oceania inferred from genome-wide data. Curr. Biol. 20, 1983–1992 (2010).
Skoglund, P. et al. Genomic insights into the peopling of the Southwest Pacific. Nature 538, 510–513 (2016).
Storey, A. A. et al. Radiocarbon and DNA evidence for a pre-Columbian introduction of Polynesian chickens to Chile. Proc. Natl Acad. Sci. USA 104, 10335–10339 (2007).
Gongora, J. et al. Indo-European and Asian origins for Chilean and Pacific chickens revealed by mtDNA. Proc. Natl Acad. Sci. USA 105, 10308–10313 (2008).
Malaspinas, A.-S. et al. Two ancient human genomes reveal Polynesian ancestry among the indigenous Botocudos of Brazil. Curr. Biol. 24, R1035–R1037 (2014).
Moreno-Mayar, J. V. et al. Genome-wide ancestry patterns in Rapanui suggest pre-European admixture with Native Americans. Curr. Biol. 24, 2518–2525 (2014).
Jenkins, D. L. et al. Clovis age Western Stemmed projectile points and human coprolites at the Paisley Caves. Science 337, 223–228 (2012).
Pedersen, M. W. et al. Postglacial viability and colonization in North America's ice-free corridor. Nature 537, 45–49 (2016).
Heintzman, P. D. et al. Bison phylogeography constrains dispersal and viability of the Ice Free Corridor in western Canada. Proc. Natl Acad. Sci. USA 113, 8057–8063 (2016).
González-José, R. et al. Craniometric evidence for Palaeoamerican survival in Baja California. Nature 425, 62–65 (2003).
Stanford, D. J. & Bradley, B. A. Across Atlantic Ice (University of California Press, 2012).
Owsley, D. W. & Jantz, R. L. (eds) Kennewick Man (Texas A&M University Press, 2014).
Rasmussen, M. et al. The genome of a Late Pleistocene human from a Clovis burial site in western Montana. Nature 506, 225–229 (2014). This study sequences a 12.6 kyr old individual from the Clovis culture, finding that people from this culture are ancestors of modern Native Americans.
Rasmussen, M. et al. The ancestry and affiliations of Kennewick Man. Nature 523, 455–458 (2015).
Raghavan, M. et al. Genomic evidence for the Pleistocene and recent population history of Native Americans. Science 349, aab3884 (2015).
Skoglund, P. et al. Genetic evidence for two founding populations of the Americas. Nature 525, 104–108 (2015).
Reich, D. et al. Reconstructing Native American population history. Nature 488, 370–374 (2012).
Gilbert, M. T. P. et al. Paleo-Eskimo mtDNA genome reveals matrilineal discontinuity in Greenland. Science 320, 1787–1789 (2008).
Raghavan, M. et al. The genetic prehistory of the New World Arctic. Science 345, 1255832 (2014).
Reich, D. et al. Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature 468, 1053–1060 (2010). This study reports the draft Denisovan genome sequence.
Sawyer, S. et al. Nuclear and mitochondrial DNA sequences from two Denisovan individuals. Proc. Natl Acad. Sci. USA 112, 15696–15700 (2015).
Stringer, C. B. & Barnes, I. Deciphering the Denisovans. Proc. Natl Acad. Sci. USA 112, 15542–15543 (2015).
Skoglund, P. & Jakobsson, M. Archaic human ancestry in East Asia. Proc. Natl Acad. Sci. USA 108, 18301–18306 (2011).
Vernot, B. et al. Excavating Neandertal and Denisovan DNA from the genomes of Melanesian individuals. Science 352, 235–239 (2016).
Huerta-Sánchez, E. et al. Altitude adaptation in Tibetans caused by introgression of Denisovan-like DNA. Nature 512, 194–197 (2014).
Kuhlwilm, M. et al. Ancient gene flow from early modern humans into Eastern Neanderthals. Nature 530, 429–433 (2016).
Wilde, S. et al. Direct evidence for positive selection of skin, hair, and eye pigmentation in Europeans during the last 5,000 y. Proc. Natl Acad. Sci. USA 111, 4832–4837 (2014).
Mathieson, I. et al. Genome-wide patterns of selection in 230 ancient Eurasians. Nature 528, 499–503 (2015).
Jablonski, N. G. & Chaplin, G. Human skin pigmentation as an adaptation to UV radiation. Proc. Natl Acad. Sci. USA 107, 8962–8968 (2010).
Lamason, R. L. et al. SLC24A5, a putative cation exchanger, affects pigmentation in zebrafish and humans. Science 310, 1782–1786 (2005).
Norton, H. L. et al. Genetic evidence for the convergent evolution of light skin in Europeans and East Asians. Mol. Biol. Evol. 24, 710–722 (2007).
Beall, C. M. Andean, Tibetan, and Ethiopian patterns of adaptation to high-altitude hypoxia. Integr. Comp. Biol. 46, 18–24 (2006).
Yi, X. et al. Sequencing of 50 human exomes reveals adaptation to high altitude. Science 329, 75–78 (2010).
Beall, C. M. et al. Natural selection on EPAS1 (HIF2α) associated with low hemoglobin concentration in Tibetan highlanders. Proc. Natl Acad. Sci. USA 107, 11459–11464 (2010).
Simonson, T. S. et al. Genetic evidence for high-altitude adaptation in Tibet. Science 329, 72–75 (2010).
Bigham, A. et al. Identifying signatures of natural selection in Tibetan and Andean populations using dense genome scan data. PLoS Genet. 6, e1001116 (2010).
Bersaglieri, T. et al. Genetic signatures of strong recent positive selection at the lactase gene. Am. J. Hum. Genet. 74, 1111–1120 (2004).
Tishkoff, S. A. et al. Convergent adaptation of human lactase persistence in Africa and Europe. Nature Genet. 39, 31–40 (2007).
Ranciaro, A. et al. Genetic origins of lactase persistence and the spread of pastoralism in Africa. Am. J. Hum. Genet. 94, 496–510 (2014).
Mathias, R. A. et al. Adaptive evolution of the FADS gene cluster within Africa. PLoS ONE 7, e44926 (2012).
Ameur, A. et al. Genetic adaptation of fatty-acid metabolism: a human-specific haplotype increasing the biosynthesis of long-chain omega-3 and omega-6 fatty acids. Am. J. Hum. Genet. 90, 809–820 (2012).
Fumagalli, M. et al. Greenlandic Inuit show genetic signatures of diet and climate adaptation. Science 349, 1343–1347 (2015).
Kothapalli, K. S. D. et al. Positive selection on a regulatory insertion–deletion polymorphism in FADS2 influences apparent endogenous synthesis of arachidonic acid. Mol. Biol. Evol. 33, 1726–1739 (2016).
Fumagalli, M. et al. Signatures of environmental genetic adaptation pinpoint pathogens as the main selective pressure through human evolution. PLoS Genet. 7, e1002355 (2011).
Traherne, J. A. Human MHC architecture and evolution: implications for disease association studies. Int. J. Immunogenet. 35, 179–192 (2008).
Kwiatkowski, D. P. How malaria has affected the human genome and what human genetics can teach us about malaria. Am. J. Hum. Genet. 77, 171–192 (2005).
Piel, F. B. et al. Global distribution of the sickle cell gene and geographical confirmation of the malaria hypothesis. Nature Commun. 1, 104 (2010).
Fan, S., Hansen, M. E. B., Lo, Y. & Tishkoff, S. A. Going global by adapting local: a review of recent human adaptation. Science 354, 54–59 (2016).
Hamblin, M. T. & Di Rienzo, A. Detection of the signature of natural selection in humans: evidence from the Duffy blood group locus. Am. J. Hum. Genet. 66, 1669–1679 (2000).
Simonti, C. N. et al. The phenotypic legacy of admixture between modern humans and Neandertals. Science 351, 737–741 (2016).
Racimo, F., Sankararaman, S., Nielsen, R. & Huerta-Sanchez, E. Evidence for archaic adaptive introgression in humans. Nature Rev. Genet. 16, 359–371 (2015).
Abi-Rached, L. et al. The shaping of modern human immune systems by multiregional admixture with archaic humans. Science 334, 89–94 (2011).
Ding, Q. et al. Neanderthal origin of the haplotypes carrying the functional variant Val92Met in the MC1R in modern humans. Mol. Biol. Evol. 31, 1994–2003 (2014).
Karlsson, E. K., Kwiatkowski, D. P. & Sabeti, P. C. Natural selection and infectious disease in human populations. Nature Rev. Genet. 15, 379–393 (2014).
Bigham, A. W. & Lee, F. S. Human high-altitude adaptation: forward genetics meets the HIF pathway. Genes Dev. 28, 2189–2204 (2014).
Pritchard, J. K., Pickrell, J. K. & Coop, G. The genetics of human adaptation: hard sweeps, soft sweeps, and polygenic adaptation. Curr. Biol. 20, R208–R215 (2010).
Turchin, M. C. et al. Evidence of widespread selection on standing variation in Europe at height-associated SNPs. Nature Genet. 44, 1015–1019 (2012).
Berg, J. J. & Coop, G. A population genetic signal of polygenic adaptation. PLoS Genet. 10, e1004412 (2014).
Zoledziewska, M. et al. Height-reducing variants and selection for short stature in Sardinia. Nature Genet. 47, 1352–1356 (2015).
Field, Y. et al. Detection of human adaptation during the past 2,000 years. Science 354, 760–764 (2016).
Gibbons, A. A new view of the birth of Homo sapiens. Science 331, 392–394 (2011).
Harris, K. & Nielsen, R. The genetic cost of Neanderthal introgression. Genetics 203, 881–891 (2016).
Renfrew, C. Before Civilization: Radiocarbon Revolution and Prehistoric Europe (Pimlico, 1973).
Childe, V. G. The Dawn of European Civilization (Kegan Paul, 1925).
Acknowledgements
This work was supported in part by: US National Institutes of Health (NIH) grant R01GM110068 (J.M.A.); the European Research Council, the Knut and Alice Wallenberg Foundation and the Swedish Research Council (M.J.); the Danish National Research Foundation and the Lundbeck Foundation and KU2016 initiative (E.W.); and NIH grants R01GM116044 (R.N.), 1R01DK104339-01 and 1R01GM113657-01 (S.T.). We also thank S. Moon, S. Tucci, A. Sapfo-Malaspinas, M. Raghavan and M. W. Pedersen for discussions or for help in preparing the figures.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reprints and permissions information is available at www.nature.com/reprints.
Reviewer Information Nature thanks D. Lambert, J. Novembre, S. Schiffels and C. Stringer for their contribution to the peer review of this work.
Rights and permissions
About this article
Cite this article
Nielsen, R., Akey, J., Jakobsson, M. et al. Tracing the peopling of the world through genomics. Nature 541, 302–310 (2017). https://doi.org/10.1038/nature21347
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21347
This article is cited by
-
Reconstructing colonization dynamics to establish how human activities transformed island biodiversity
Scientific Reports (2024)
-
Physiography, foraging mobility, and the first peopling of Sahul
Nature Communications (2024)
-
Tumor phylogeography reveals block-shaped spatial heterogeneity and the mode of evolution in Hepatocellular Carcinoma
Nature Communications (2024)
-
Undiscovered bird extinctions obscure the true magnitude of human-driven extinction waves
Nature Communications (2023)
-
Haplotype-based inference of recent effective population size in modern and ancient DNA samples
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.