Abstract
CRISPR–Cas systems provide microbes with adaptive immunity by employing short DNA sequences, termed spacers, that guide Cas proteins to cleave foreign DNA1,2. Class 2 CRISPR–Cas systems are streamlined versions, in which a single RNA-bound Cas protein recognizes and cleaves target sequences3,4. The programmable nature of these minimal systems has enabled researchers to repurpose them into a versatile technology that is broadly revolutionizing biological and clinical research5. However, current CRISPR–Cas technologies are based solely on systems from isolated bacteria, leaving the vast majority of enzymes from organisms that have not been cultured untapped. Metagenomics, the sequencing of DNA extracted directly from natural microbial communities, provides access to the genetic material of a huge array of uncultivated organisms6,7. Here, using genome-resolved metagenomics, we identify a number of CRISPR–Cas systems, including the first reported Cas9 in the archaeal domain of life, to our knowledge. This divergent Cas9 protein was found in little-studied nanoarchaea as part of an active CRISPR–Cas system. In bacteria, we discovered two previously unknown systems, CRISPR–CasX and CRISPR–CasY, which are among the most compact systems yet discovered. Notably, all required functional components were identified by metagenomics, enabling validation of robust in vivo RNA-guided DNA interference activity in Escherichia coli. Interrogation of environmental microbial communities combined with in vivo experiments allows us to access an unprecedented diversity of genomes, the content of which will expand the repertoire of microbe-based biotechnologies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007)
Sorek, R., Kunin, V. & Hugenholtz, P. CRISPR—a widespread system that provides acquired resistance against phages in bacteria and archaea. Nat. Rev. Microbiol. 6, 181–186 (2008)
Makarova, K. S. et al. An updated evolutionary classification of CRISPR–Cas systems. Nat. Rev. Microbiol. 13, 722–736 (2015)
Shmakov, S. et al. Discovery and functional characterization of diverse class 2 CRISPR–Cas systems. Mol. Cell 60, 385–397 (2015)
Barrangou, R. & Doudna, J. A. Applications of CRISPR technologies in research and beyond. Nat. Biotechnol. 34, 933–941 (2016)
Brown, C. T. et al. Unusual biology across a group comprising more than 15% of domain Bacteria. Nature 523, 208–211 (2015)
Sharon, I. & Banfield, J. F . Genomes from metagenomics. Science 342, 1057–1058 (2013)
Levy, A. et al. CRISPR adaptation biases explain preference for acquisition of foreign DNA. Nature 520, 505–510 (2015)
Yosef, I., Goren, M. G. & Qimron, U. Proteins and DNA elements essential for the CRISPR adaptation process in Escherichia coli. Nucleic Acids Res. 40, 5569–5576 (2012)
Nuñez, J. K., Lee, A. S. Y., Engelman, A. & Doudna, J. A. Integrase-mediated spacer acquisition during CRISPR–Cas adaptive immunity. Nature 519, 193–198 (2015)
Chylinski, K., Makarova, K. S., Charpentier, E. & Koonin, E. V. Classification and evolution of type II CRISPR–Cas systems. Nucleic Acids Res. 42, 6091–6105 (2014)
Baker, B. J. et al. Enigmatic, ultrasmall, uncultivated Archaea. Proc. Natl Acad. Sci. USA 107, 8806–8811 (2010)
Baker, B. J. et al. Lineages of acidophilic Archaea revealed by community genomic analysis. Science 314, 1933–1935 (2006)
Comolli, L. R. & Banfield, J. F. Inter-species interconnections in acid mine drainage microbial communities. Front. Microbiol. 5, 367 (2014)
Yelton, A. P. et al. Comparative genomics in acid mine drainage biofilm communities reveals metabolic and structural differentiation of co-occurring archaea. BMC Genomics 14, 485 (2013)
Vagin, V. V. et al. A distinct small RNA pathway silences selfish genetic elements in the germline. Science 313, 320–324 (2006)
Stern, A., Keren, L., Wurtzel, O., Amitai, G. & Sorek, R. Self-targeting by CRISPR: gene regulation or autoimmunity? Trends Genet. 26, 335–340 (2010)
Zegans, M. E. et al. Interaction between bacteriophage DMS3 and host CRISPR region inhibits group behaviors of Pseudomonas aeruginosa. J. Bacteriol. 191, 210–219 (2009)
Shah, S. A., Erdmann, S., Mojica, F. J. M. & Garrett, R. A. Protospacer recognition motifs: mixed identities and functional diversity. RNA Biol. 10, 891–899 (2013)
Anders, C., Niewoehner, O., Duerst, A. & Jinek, M. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 513, 569–573 (2014)
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012)
Deltcheva, E. et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471, 602–607 (2011)
Zhang, Y., Rajan, R., Seifert, H. S., Mondragón, A. & Sontheimer, E. J. DNase H Activity of Neisseria meningitidis Cas9. Mol. Cell 60, 242–255 (2015)
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR–Cas system. Cell 163, 759–771 (2015)
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016)
Anantharaman, K. et al. Thousands of microbial genomes shed light on interconnected biogeochemical processes in an aquifer system. Nat. Commun. 7, 13219 (2016)
Godde, J. S. & Bickerton, A. The repetitive DNA elements called CRISPRs and their associated genes: evidence of horizontal transfer among prokaryotes. J. Mol. Evol. 62, 718–729 (2006)
Burstein, D. et al. Major bacterial lineages are essentially devoid of CRISPR-Cas viral defence systems. Nat. Commun. 7, 10613 (2016)
Hug, L. A. et al. A new view of the tree of life. Nat. Microbiol. 1, 16048 (2016)
Luef, B. et al. Diverse uncultivated ultra-small bacterial cells in groundwater. Nat. Commun. 6, 6372 (2015)
Kantor, R. S. et al. Small genomes and sparse metabolisms of sediment-associated bacteria from four candidate phyla. MBio 4, e00708–e00713 (2013)
Nelson, W. C. & Stegen, J. C. The reduced genomes of Parcubacteria (OD1) contain signatures of a symbiotic lifestyle. Front. Microbiol. 6, 713 (2015)
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013)
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–W37 (2011)
Nuñez, J. K. et al. Cas1–Cas2 complex formation mediates spacer acquisition during CRISPR–Cas adaptive immunity. Nat. Struct. Mol. Biol. 21, 528–534 (2014)
Denef, V. J. & Banfield, J. F. In situ evolutionary rate measurements show ecological success of recently emerged bacterial hybrids. Science 336, 462–466 (2012)
Miller, C. S., Baker, B. J., Thomas, B. C., Singer, S. W. & Banfield, J. F. EMIRGE: reconstruction of full-length ribosomal genes from microbial community short read sequencing data. Genome Biol. 12, R44 (2011)
Probst, A. J. et al. Genomic resolution of a cold subsurface aquifer community provides metabolic insights for novel microbes adapted to high CO2 concentrations. Environ. Microbiol. http://dx.doi.org/10.1111/1462-2920.13362 (2016)
Emerson, J. B., Thomas, B. C., Alvarez, W. & Banfield, J. F. Metagenomic analysis of a high carbon dioxide subsurface microbial community populated by chemolithoautotrophs and bacteria and archaea from candidate phyla. Environ. Microbiol. 18, 1686–1703 (2016)
Peng, Y., Leung, H. C. M., Yiu, S. M. & Chin, F. Y. L. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28, 1420–1428 (2012)
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012)
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11, 119 (2010)
Wu, Y.-W., Simmons, B. A. & Singer, S. W. MaxBin 2.0: an automated binning algorithm to recover genomes from multiple metagenomic datasets. Bioinformatics 32, 605–607 (2016)
Dick, G. J. et al. Community-wide analysis of microbial genome sequence signatures. Genome Biol. 10, R85 (2009)
Grissa, I., Vergnaud, G. & Pourcel, C. CRISPRFinder: a web tool to identify clustered regularly interspaced short palindromic repeats. Nucleic Acids Res. 35, W52–W57 (2007)
Enright, A. J., Van Dongen, S. & Ouzounis, C. A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 30, 1575–1584 (2002)
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009)
The UniProt Consortium. UniProt: a hub for protein information. Nucleic Acids Res. 43, D204–D212 (2015)
Remmert, M., Biegert, A., Hauser, A. & Söding, J. HHblits: lightning-fast iterative protein sequence searching by HMM–HMM alignment. Nat. Methods 9, 173–175 (2011)
Dong, D. et al. The crystal structure of Cpf1 in complex with CRISPR RNA. Nature 532, 522–526 (2016)
Yamano, T. et al. Crystal structure of Cpf1 in complex with guide RNA and target DNA. Cell 165, 949–962 (2016)
Drozdetskiy, A., Cole, C., Procter, J. & Barton, G. J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res. 43 (W1), W389–W394 (2015)
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocols 10, 845–858 (2015)
Skennerton, C. T., Imelfort, M. & Tyson, G. W. Crass: identification and reconstruction of CRISPR from unassembled metagenomic data. Nucleic Acids Res. 41, e105(2013)
Crooks, G. E., Hon, G., Chandonia, J.-M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004)
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003)
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol. 5, R12 (2004)
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012)
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013)
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014)
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44 (W1), W242–W245 (2016)
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009)
Esvelt, K. M. et al. Orthogonal Cas9 proteins for RNA-guided gene regulation and editing. Nat. Methods 10, 1116–1121 (2013)
Zhang, Y. et al. Processing-independent CRISPR RNAs limit natural transformation in Neisseria meningitidis. Mol. Cell 50, 488–503 (2013)
Sternberg, S. H., Haurwitz, R. E. & Doudna, J. A. Mechanism of substrate selection by a highly specific CRISPR endoribonuclease. RNA 18, 661–672 (2012)
Oakes, B. L. et al. Profiling of engineering hotspots identifies an allosteric CRISPR–Cas9 switch. Nat. Biotechnol. 34, 646–651 (2016)
Jinek, M. et al. Structures of Cas9 endonucleases reveal RNA-mediated conformational activation. Science 343, http://dx.doi.org/10.1126/science.1247997 (2014)
Acknowledgements
We thank N. Ma, K. Zhou and D. McGrath for technical assistance; C. Brown, M. Olm, M. O’Connell, J. Chen and S. Floor for reading the manuscript and discussions; and V. Yu for the S. cerevisiae expression strain. D.B. was supported by a long-term EMBO fellowship, L.B.H. by a US National Science Foundation Graduate Research Fellowship, and A.J.P. by a fellowship of the German Science Foundation (DFG PR 1603/1-1). J.A.D. is an Investigator of the Howard Hughes Medical Institute. This research was supported in part by the Allen Distinguished Investigator Program, through The Paul G. Allen Frontiers Group, the National Science Foundation (MCB-1244557 to J.A.D.) and the Lawrence Berkeley National Laboratory’s Sustainable Systems Scientific Focus Area funded by the US Department of Energy (DE-AC02-05CH11231 to J.F.B.). DNA sequencing was conducted at the DOE Joint Genome Institute, a DOE Office of Science User Facility, via the Community Science Program.
Author information
Authors and Affiliations
Contributions
D.B., L.B.H., S.C.S., J.A.D. and J.F.B. designed the study and wrote the manuscript. A.J.P., K.A., J.F.B., B.T.C. and D.B. assembled the data and reconstructed the genomes. D.B., L.B.H., S.C.S. and J.F.B. computationally analysed the CRISPR–Cas systems. L.B.H. and D.B. designed and executed experimental work with CRISPR–CasX and CRISPR–CasY. S.C.S. designed and executed the experimental work with ARMAN Cas9. The manuscript was read, edited and approved by all authors.
Corresponding authors
Ethics declarations
Competing interests
The Regents of the University of California have filed a provisional patent application related to the technology described in this work to the United States Patent and Trademark Office, in which D.B., L.B.H., S.C.S., J.A.D. and J.F.B. are listed as inventors.
Additional information
Reviewer Information Nature thanks E. Sontheimer, R. Sorek and M. White for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Multiple sequence alignment of newly described Cas9 proteins.
Alignment of Cas9 proteins from ARMAN-1 and ARMAN-4, as well as two closely related Cas9 proteins from uncultivated bacteria, to the Actinomyces naeslundii Cas9, whose structure has been solved67.
Extended Data Figure 2 Within-population variability of ARMAN-1 CRISPR arrays.
Variability of reconstructed CRISPR arrays, including the most well represented (and thus assembled) sequences (Fig. 2) and array segments representing locus variants that were reconstructed from the short DNA reads. Variability is due to spacers that were present in only a subset of archaeal cells in the population, as well as spacers whose context differed owing to spacer loss (indicated by black lines). White boxes indicate repeats and coloured arrows indicate CRISPR spacers (spacers with different colours have different sequences, except for unique spacers that are black). In CRISPR systems, spacers are typically added unidirectionally, so the high variety of spacers on the left side is attributed to recent acquisition.
Extended Data Figure 3 Novelty of the reported CRISPR–Cas systems.
a, Simplified phylogenetic tree of the universal Cas1 protein. CRISPR types of known systems are noted on the wedges and branches; the newly described systems are in bold. Detailed Cas1 phylogeny is provided in Supplementary Data 4. b, Proposed evolutionary scenario that gave rise to the archaeal type II system as a result of a recombination between type II-B and type II-C loci. c, Similarity of CasX and CasY to known proteins based on the following searches: (1) BLAST search against the non-redundant (NR) protein database of NCBI; (2) HMM search against an HMM database of known Cas proteins; and (3) distant homology search using HHpred49 (E, e value).
Extended Data Figure 4 Evolutionary tree of Cas9 homologues.
Maximum-likelihood phylogenic tree of Cas9 proteins, showing the previously described systems coloured based on their type. II-A, blue; II-B, green; II-C, purple. The archaeal Cas9 (red) cluster with type II-C CRISPR–Cas systems, together with two newly described bacterial Cas9 from uncultivated bacteria. A detailed tree is provided in Supplementary Data 5.
Extended Data Figure 5 ARMAN-1 spacers map to genomes of archaeal community members.
a, Protospacers from ARMAN-1 map to the genome of ARMAN-2, a nanoarchaeon from the same environment. Six protospacers (red arrowheads) map uniquely to a portion of the genome flanked by two long-terminal repeats (LTRs), and two additional protospacers match perfectly within the LTRs (blue and green arrowheads). This region is likely to be a transposon, suggesting that the CRISPR–Cas system of ARMAN-1 plays a role in suppressing mobilization of this element. b, Protospacers also map to a Thermoplasmatales archaeon (I-plasma), another member of the Richmond Mine ecosystem that is found in the same samples as ARMAN organisms. The protospacers cluster within a region of the genome encoding short, hypothetical proteins, suggesting this might also represent a mobile element. NCBI accession codes are provided in parentheses.
Extended Data Figure 6 Archaeal Cas9 from ARMAN-4 with a degenerate CRISPR array is found on numerous contigs.
Cas9 from ARMAN-4 is highlighted in dark red on 16 nearly identical contigs from different samples. Proteins with putative domains or functions are labelled, whereas hypothetical proteins are unlabelled. Fifteen of the contigs contain two degenerate direct repeats (36 nucleotides long with one mismatch) and a single conserved spacer of 36 nucleotides. The remaining contig contains only one direct repeat. Unlike ARMAN-1, no additional Cas proteins are found adjacent to Cas9 in ARMAN-4.
Extended Data Figure 7 Predicted structures of guide RNA and purification schema for in vitro biochemistry studies.
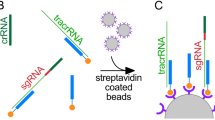
a, The CRISPR repeat and tracrRNA anti-repeat are depicted in black whereas the spacer-derived sequence is shown as a series of green Ns. No clear termination signal can be predicted from the locus, so three different tracrRNA lengths were tested based on their secondary structure: 69, 104, and 179 nucleotides in red, blue, and pink, respectively. b, Engineered single-guide RNA corresponding to dual-guide in a. c, Dual-guide RNA for ARMAN-4 Cas9 with two different hairpins on 3′ end of tracrRNA (75 and 122 nucleotides). d, Engineered single-guide RNA corresponding to dual-guide in c. e, Conditions tested in E. coli in vivo targeting assay. f, ARMAN-1 (AR1) and ARMAN-4 (AR4) Cas9 were expressed and purified under a variety of conditions as outlined in the Methods section. Proteins outlined in blue boxes were tested for cleavage activity in vitro. g, Fractions of AR1-Cas9 and AR4-Cas9 purifications were separated on a 10% SDS–PAGE gel.
Extended Data Figure 8 Programmed DNA interference by CasX.
a, Plasmid interference assays for CasX.1 (Deltaproteobacteria) and CasX.2 (Planctomycetes), continued from Fig. 3c (sX1, CasX spacer 1; sX2, CasX spacer 2; NT, non-target). Experiments were conducted in triplicate and mean ± s.d. is shown. b, Serial dilution of E. coli expressing a CasX locus and transformed with the specified target, continued from Fig. 3b. c, PAM depletion assays for the Deltaproteobacteria CasX and d, Planctomycetes CasX expressed in E. coli. PAM sequences depleted greater than the indicated PAM depletion value threshold (PDVT) compared to a control library were used to generate the sequence logo. e, Diagram depicting the location of northern blot probes for CasX.1. f, Northern blots for CasX.1 tracrRNA in total RNA extracted from E. coli expressing the CasX.1 locus. The sequences of the probes used are provided in Supplementary Table 2.
Supplementary information
Supplementary Table 1
This file contains Supplementary Table 1, reconstructed spacer and protospacers of the ARMAN-1 Type II CRISPR-Cas system. (XLSX 31 kb)
Supplementary Table 2
This file contains Supplementary Table 2, a list of primers and plasmids used in the study. (XLSX 38 kb)
Supplementary Data
This zipped file contains Supplementary Data sets 1-6. (ZIP 10208 kb)
Rights and permissions
About this article
Cite this article
Burstein, D., Harrington, L., Strutt, S. et al. New CRISPR–Cas systems from uncultivated microbes. Nature 542, 237–241 (2017). https://doi.org/10.1038/nature21059
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21059
This article is cited by
-
Molecular basis and engineering of miniature Cas12f with C-rich PAM specificity
Nature Chemical Biology (2024)
-
Mechanistic understanding on the uptake of micro-nano plastics by plants and its phytoremediation
Environmental Science and Pollution Research (2024)
-
Soil microbial ecology through the lens of metatranscriptomics
Soil Ecology Letters (2024)
-
Nucleic acid drug vectors for diagnosis and treatment of brain diseases
Signal Transduction and Targeted Therapy (2023)
-
Prevalence and transmission risk of colistin and multidrug resistance in long-distance coastal aquaculture
ISME Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.