Abstract
Quality control mechanisms intervene appropriately when defective translation events occur, in order to preserve the integrity of protein synthesis. Rescue of ribosomes translating on messenger RNAs that lack stop codons is one of the co-translational quality control pathways. In many bacteria, ArfA recognizes stalled ribosomes and recruits the release factor RF2, which catalyses the termination of protein synthesis1,2,3. Although an induced-fit mechanism of nonstop mRNA surveillance mediated by ArfA and RF2 has been reported4, the molecular interaction between ArfA and RF2 in the ribosome that is responsible for the mechanism is unknown. Here we report an electron cryo-microscopy structure of ArfA and RF2 in complex with the 70S ribosome bound to a nonstop mRNA. The structure, which is consistent with our kinetic and biochemical data, reveals the molecular interactions that enable ArfA to specifically recruit RF2, not RF1, into the ribosome and to enable RF2 to release the truncated protein product in this co-translational quality control pathway. The positively charged C-terminal domain of ArfA anchors in the mRNA entry channel of the ribosome. Furthermore, binding of ArfA and RF2 induces conformational changes in the ribosomal decoding centre that are similar to those seen in other protein-involved decoding processes. Specific interactions between residues in the N-terminal domain of ArfA and RF2 help RF2 to adopt a catalytically competent conformation for peptide release. Our findings provide a framework for understanding recognition of the translational state of the ribosome by new proteins, and expand our knowledge of the decoding potential of the ribosome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Shimizu, Y. . ArfA recruits RF2 into stalled ribosomes. J. Mol. Biol. 423, 624–631 (2012)
Chadani, Y., Ito, K., Kutsukake, K. & Abo, T. . ArfA recruits release factor 2 to rescue stalled ribosomes by peptidyl-tRNA hydrolysis in Escherichia coli. Mol. Microbiol. 86, 37–50 (2012)
Chadani, Y. et al. Ribosome rescue by Escherichia coli ArfA (YhdL) in the absence of trans-translation system. Mol. Microbiol. 78, 796–808 (2010)
Zeng, F. & Jin, H. Peptide release promoted by methylated RF2 and ArfA in nonstop translation is achieved by an induced-fit mechanism. RNA 22, 49–60 (2016)
Rodnina, M. V. Quality control of mRNA decoding on the bacterial ribosome. Adv. Protein Chem. Struct. Biol. 86, 95–128 (2012)
Keiler, K. C. Mechanisms of ribosome rescue in bacteria. Nat. Rev. Microbiol. 13, 285–297 (2015)
Giudice, E. & Gillet, R. The task force that rescues stalled ribosomes in bacteria. Trends Biochem. Sci. 38, 403–411 (2013)
Ito, K., Uno, M. & Nakamura, Y. A tripeptide ‘anticodon’ deciphers stop codons in messenger RNA. Nature 403, 680–684 (2000)
Zaher, H. S. & Green, R. Quality control by the ribosome following peptide bond formation. Nature 457, 161–166 (2009)
Frolova, L. Y. et al. Mutations in the highly conserved GGQ motif of class 1 polypeptide release factors abolish ability of human eRF1 to trigger peptidyl-tRNA hydrolysis. RNA 5, 1014–1020 (1999)
Mora, L. et al. The essential role of the invariant GGQ motif in the function and stability in vivo of bacterial release factors RF1 and RF2. Mol. Microbiol. 47, 267–275 (2003)
Dinçbas-Renqvist, V. et al. A post-translational modification in the GGQ motif of RF2 from Escherichia coli stimulates termination of translation. EMBO J. 19, 6900–6907 (2000)
Kurita, D., Chadani, Y., Muto, A., Abo, T. & Himeno, H. . ArfA recognizes the lack of mRNA in the mRNA channel after RF2 binding for ribosome rescue. Nucleic Acids Res. 42, 13339–13352 (2014)
Rodnina, M. V., Beringer, M. & Wintermeyer, W. Mechanism of peptide bond formation on the ribosome. Q. Rev. Biophys. 39, 203–225 (2006)
Youngman, E. M., He, S. L., Nikstad, L. J. & Green, R. Stop codon recognition by release factors induces structural rearrangement of the ribosomal decoding center that is productive for peptide release. Mol. Cell 28, 533–543 (2007)
Garza-Sánchez, F., Schaub, R. E., Janssen, B. D. & Hayes, C. S. tmRNA regulates synthesis of the ArfA ribosome rescue factor. Mol. Microbiol. 80, 1204–1219 (2011)
Schaub, R. E., Poole, S. J., Garza-Sánchez, F., Benbow, S. & Hayes, C. S. Proteobacterial ArfA peptides are synthesized from nonstop messenger RNAs. J. Biol. Chem. 287, 29765–29775 (2012)
Ogle, J. M., Murphy, F. V., Tarry, M. J. & Ramakrishnan, V. Selection of tRNA by the ribosome requires a transition from an open to a closed form. Cell 111, 721–732 (2002)
Klaholz, B. P. et al. Structure of the Escherichia coli ribosomal termination complex with release factor 2. Nature 421, 90–94 (2003)
Korostelev, A. et al. Crystal structure of a translation termination complex formed with release factor RF2. Proc. Natl Acad. Sci. USA 105, 19684–19689 (2008)
Rawat, U. B. S. et al. A cryo-electron microscopic study of ribosome-bound termination factor RF2. Nature 421, 87–90 (2003)
Weixlbaumer, A. et al. Insights into translational termination from the structure of RF2 bound to the ribosome. Science 322, 953–956 (2008)
Gagnon, M. G., Seetharaman, S. V., Bulkley, D. & Steitz, T. A. Structural basis for the rescue of stalled ribosomes: structure of YaeJ bound to the ribosome. Science 335, 1370–1372 (2012)
Neubauer, C., Gillet, R., Kelley, A. C. & Ramakrishnan, V. Decoding in the absence of a codon by tmRNA and SmpB in the ribosome. Science 335, 1366–1369 (2012)
Laurberg, M. et al. Structural basis for translation termination on the 70S ribosome. Nature 454, 852–857 (2008)
Jin, H., Kelley, A. C., Loakes, D. & Ramakrishnan, V. Structure of the 70S ribosome bound to release factor 2 and a substrate analog provides insights into catalysis of peptide release. Proc. Natl Acad. Sci. USA 107, 8593–8598 (2010)
Schmeing, T. M., Huang, K. S., Strobel, S. A. & Steitz, T. A. An induced-fit mechanism to promote peptide bond formation and exclude hydrolysis of peptidyl-tRNA. Nature 438, 520–524 (2005)
Cammack, K. A. & Wade, H. E. The sedimentation behaviour of ribonuclease-active and -inactive ribosomes from bacteria. Biochem. J. 96, 671–680 (1965)
Bommer, U. et al. in Subcellular Fractionation: A Practical Approach (eds Graham, J. & Rickwood, D. ) 271–301 (IRL, Washington, 1997)
Kurylo, C. M. et al. Genome sequence and analysis of Escherichia coli MRE600, a colicinogenic, nonmotile strain that lacks RNase I and the type I methyltransferase, EcoKI. Genome Biol. Evol. 8, 742–752 (2016)
Wolfrum, A., Brock, S., Mac, T. & Grillenbeck, N. Expression in E. coli and purification of Thermus thermophilus translation initiation factors IF1 and IF3. Protein Expr. Purif. 29, 15–23 (2003)
Chadani, Y. et al. Trans-translation-mediated tight regulation of the expression of the alternative ribosome-rescue factor ArfA in Escherichia coli. Genes Genet. Syst. 86, 151–163 (2011)
Schmitt, E. et al. Crystallization and preliminary X-ray analysis of Escherichia coli methionyl-tRNA(fMet) formyltransferase. Proteins 25, 139–141 (1996)
Walker, S. E. & Fredrick, K. Preparation and evaluation of acylated tRNAs. Methods 44, 81–86 (2008)
Monteiro, R. A., Souza, E. M., Yates, M. G., Pedrosa, F. O. & Chubatsu, L. S. Use of lactose to induce expression of soluble NifA protein domains of Herbaspirillum seropedicae in Escherichia coli. Can. J. Microbiol. 46, 1087–1090 (2000)
Sprink, T. et al. Structures of ribosome-bound initiation factor 2 reveal the mechanism of subunit association. Sci. Adv. 2, e1501502 (2016)
Stark, H. & Chari, A. Sample preparation of biological macromolecular assemblies for the determination of high-resolution structures by cryo-electron microscopy. Microscopy 65, 23–34 (2015)
Grassucci, R. A., Taylor, D. J. & Frank, J. Preparation of macromolecular complexes for cryo-electron microscopy. Nat. Protocols 2, 3239–3246 (2007)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013)
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015)
Scheres, S. H. A Bayesian view on cryo-EM structure determination. J. Mol. Biol. 415, 406–418 (2012)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Kimanius, D., Forsberg, B. O., Scheres, S. H. W. & Lindahl, E. Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. eLife 5, e18722 (2016)
Scheres, S. H. Semi-automated selection of cryo-EM particles in RELION-1.3. J. Struct. Biol. 189, 114–122 (2015)
Scheres, S. H. W. Beam-induced motion correction for sub-megadalton cryo-EM particles. eLife 3, e03665 (2014)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014)
Bai, X. C., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, e11182 (2015)
Scheres, S. H. W. Processing of structurally heterogeneous cryo-EM data in RELION. Methods Enzymol. 579, 125–157 (2016)
Nguyen, T. H. D. et al. Cryo-EM structure of the yeast U4/U6.U5 tri-snRNP at 3.7 Å resolution. Nature 530, 298–302 (2016)
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012)
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocols 10, 845–858 (2015)
Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9, 40 (2008)
Noeske, J. et al. High-resolution structure of the Escherichia coli ribosome. Nat. Struct. Mol. Biol. 22, 336–341 (2015)
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Biasini, M. et al. SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 42, W252–W258 (2014)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011)
Nicholls, R. A., Fischer, M., McNicholas, S. & Murshudov, G. N. Conformation-independent structural comparison of macromolecules with ProSMART. Acta Crystallogr. D 70, 2487–2499 (2014)
Brown, A. et al. Tools for macromolecular model building and refinement into electron cryo-microscopy reconstructions. Acta Crystallogr. D 71, 136–153 (2015)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Schrodinger, LLC. The PyMOL Molecular Graphics System, Version 1.7. (2015)
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011)
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001)
Dolinsky, T. J., Nielsen, J. E., McCammon, J. A. & Baker, N. A. PDB2PQR: an automated pipeline for the setup of Poisson-Boltzmann electrostatics calculations. Nucleic Acids Res. 32, W665–W667 (2004)
Jenner, L. B., Demeshkina, N., Yusupova, G. & Yusupov, M. Structural aspects of messenger RNA reading frame maintenance by the ribosome. Nat. Struct. Mol. Biol. 17, 555–560 (2010)
Acknowledgements
We thank J. Peng for scripting, W. Jiang, Y. He, J. Liu and T. H. D. Nguyen for suggestions on EM data collection and structural refinement, members of the Jin laboratory for discussions, the structural biology facility at Northwestern University for the use of the microscope and the Chicago Biomedical Consortium for the purchase of the Gatan K2 detector. H.J. acknowledges support from the National Institute of General Medical Sciences of the NIH (R01-GM120552). The computational part of the study was supported by the National Institute of General Medical Sciences (P41-GM104601 to J.C.P. and E.T.).
Author information
Authors and Affiliations
Contributions
F.Z. and H.J. designed the study. F.Z. purified ribosomes, proteins and tRNAs, processed EM data and built the atomic model. Y.C. did the mutagenesis and purified ArfA mutants. F.Z. and Y.C. performed the peptide release assay. F.Z., Y.C., H.J. and J.R. collected EM data. M.S., J.C.P. and E.T. provided computational support. F.Z., Y.C. and H.J. analysed the data and refined the structure. F.Z. and Y.C. helped with manuscript preparation. H.J. wrote the paper. All authors discussed the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks Y. Hashem, K. Keiler and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Structure determination of 70S ribosome with ArfA and RF2 on a nonstop mRNA by cryoEM.
a, A representative micrograph with corresponding FFT shown in an insert. The insert shows the CTF estimation of an average background-subtracted power spectrum and fitted CTF (top-left corner) using CTFFIND440. b, Representative 2D class averages from reference-free 2D classification. c, Particle classification and structural refinement procedures used to in this study. d, Conformational differences between the nonstop ribosomal complexes in the two classes obtained from the global classification and refinement. Rotation of 16S rRNA in the 30S subunit relative to 23S rRNA in the 50S subunit from class I (red) to class II (blue) showing a 1.7° 16S body domain rotation and an orthogonal 1.8° head domain rotation. Ribosomes in the two classes were aligned using the 23S rRNA of the 50S subunit. Small subunit ribosomal proteins are omitted for clarity.
Extended Data Figure 2 Map and model quality.
a, Gold-standard FSC curves for the electron microscopy map from 155,440 particles (black), and electron microscopy maps of class I (red, 82,077 particles) and class II (blue, 73,363 particles) from the global 3D classification and refinement. Resolution is demarcated using the FSC = 0.143 criterion. b, Fit of the model to the map. FSC curves calculated between the refined structural model and the final electron microscopy map (sum, black), with the self-validated (half1, red) and cross-validated (half2, blue) correlations shown. c, The unfiltered and unsharpened density map coloured by local resolution in surface and slice views for the entire nonstop ribosomal complex. d, Gold-standard FSC curves for the electron microscopy maps obtained from the focused refinements with a mask over the ArfA/RF2/30S body domain (black) and a mask over the ArfA/RF2 region (red). e, Same as c for the electron microscopy map obtained from the focused refinement with a mask over the regions of ArfA, RF2 and the 30S body domain. f, Gold-standard FSC curves for the electron microscopy maps obtained from the local refinement with partial signal subtraction using a mask over the ArfA/RF2/30S body domain (black) and a mask over the ArfA/RF2 region (red). g, Fit of the model to the map. FSC curves are calculated between the refined structural model and the final electron microscopy map (sum, black) for the ArfA and RF2 region, with the self-validated (half1, red) and cross-validated (half2, blue) correlations shown. h, Same as c for the electron microscopy map obtained from the local refinement with a mask over the region of ArfA and RF2. i, Representative electron microscopy maps showing the refined structures of ArfA (density), RF2 (firebrick), 23S rRNA (cyan) and 16S rRNA (orange).
Extended Data Figure 3 Conformations of RF2 in the canonical and nonstop termination complexes.
Superposition of structures of RF2 in the canonical termination complex from T. thermophilus (PDB code: 4V5J26, coloured in grey) and nonstop translation complex from E. coli (this study, coloured in red) based on an alignment of the 16S rRNAs of the two ribosomal complexes. In both structures, domains II and IV of RF2 bind to the decoding centre of the ribosome, domain III extends into the 50S subunit positioning the universally conserved GGQ motif into the PTC, and domain I interacts with the L11 stalk and the 30S shoulder region.
Extended Data Figure 4 Sequence alignment of ArfA from different bacterial species.
a, Pictorial representation of the consensus sequence showing the frequency of different amino acids in residues 1–55 of ArfA from 66 species. Alignment was generated by Weblogo (http://weblogo.berkeley.edu/logo.cgi), and amino acids are coloured according to their chemical properties. Polar and uncharged amino acids (G, S, T, Y, C) are coloured green, amino acids (Q, N) in purple, basic amino acids (K, R, H) in blue, acidic amino acids (D, E) in red, and hydrophobic amino acids (A, V, L, I, P, W, F, M) in black. Secondary structure features of ArfA from E. coli when it binds to the ribosome are shown according to the atomic structure obtained in this study. b, Multiple sequence alignment of ArfA proteins from different bacterial species showing that all ArfA proteins contain a positively charged C-terminal domain, as framed in the blue box. E. coli ArfA (NCBI GenInfo Identifier (GI) number 1450289), Proteus mirabilis ArfA (GI6802815), Haemophilus influenza ArfA (GI951152), Pasteurella multocida ArfA (GI29389120), and Vibrio fischeri ArfA (GI3280674) are used as examples. The red box indicates the C-terminal truncated region during ArfA maturation17. ArfA mutants used in this study are highlighted in yellow. Multiple sequence alignment was carried out in Clustal Omega62.
Extended Data Figure 5 Interactions of the C-terminal tail of ArfA and the ribosome.
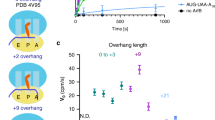
a, View of the mRNA entry channel showing the positively charged residues in the C-terminal domain of ArfA interacting with negatively charged phosphate backbones of rRNA. RF2 is omitted for clarity. Electrostatic potentials were calculated using APBS63 with pdb2pqr64, where k is Boltzmann’s constant, T is the system temperature of the calculation (310 K) and ec is the charge of an electron. b, c, Positively charged residues in ArfA interact with phosphate backbones of the rRNA in the ribosome. Densities for selected ArfA residues are shown. R41 of ArfA makes contact with nucleotides in the mRNA entry channel (b). R28 of ArfA interacts with A519 of 16S rRNA (c). ArfA and16S rRNA are coloured blue and orange, respectively. d, Flexibility of loop 3 (E30–G37) in ArfA allows accommodation of a few nucleotides downstream from the P-site in the ribosome. One to two codons downstream of the ribosomal P-site appear to be accommodated by ArfA, and three codons downstream from the P-site nearly abolishes the peptide release activity in the ribosome, consistent with the biochemical data1,4,13. 30S is shown in yellow. ArfA, nonstop mRNA and P-tRNA are coloured in blue, magenta and lemon, respectively. An mRNA (grey) taken from PDB 4V6F65 was used in the figure for the purpose of illustration.
Extended Data Figure 6 Peptide release by RF2 and ArfA mutants in the nonstop stalled ribosome.
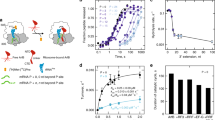
a, Representative time courses of peptide release for ArfA mutants with point mutations on residues interacting with the ribosomal decoding centre including P23A, R28A and E30A. b, Representative time courses of peptide release for ArfA mutants with point mutations in the C-terminal tail including K34A, K36A and R41A. Data on wild-type ArfA are shown. Ribosomal complexes (25 nM) with nonstop mRNA and P-site fMet-tRNAfMet were incubated with 62.5 nM ArfA and 5 μM RF2 at 37 °C and the released peptides were measured at different time points after adding ArfA and RF2. c. Observed rate versus RF2 concentrations showing fits for kcat and K1/2 calculations for wild-type ArfA and K34A, K36A and R41A mutants. d, Observed rate versus RF2 concentrations showing fits for kcat and K1/2 calculations for wild-type ArfA and A18T mutant. The kcat and K1/2 values on A18T, kcat = 0.0038 ± 0.0002 s−1 and K1/2 = 1.14 ± 0.28 × 10−6 M, were reported but the fitting curve was not shown4. kobs was determined as described in Methods. Catalytic rate constants and values of K1/2 were obtained by fitting the observed rates against the corresponding RF2 concentrations to the Michaelis–Menten equation. An average of three independent measurements is reported for each reaction and errors are calculated by standard error propagation.
Extended Data Figure 7 Multiple sequence alignment of domain II–IV of RF2 and RF1 from different bacterial species.
E. coli RF1 (NCBI GenInfo Identifier number 949002) and RF2 (GI947369), T. thermophilus RF1 (GI3169506) and RF2 (GI3168831), P. mirabilis RF1 (GI6801441) and RF2 (GI23391224), P. multocida RF1 (GI29388454) and RF2 (GI29389590), and V. fischeri RF1 (GI3277422) and RF2 (GI3277319) were submitted to Clustal Omega for alignment. Except for the genome of T. thermophiles, which does not contain a gene for ArfA, the genomes of the other species all contain the ArfA gene17. The sequence alignment is validated by structural alignment of RF1 and RF222,25. Domain I of RF1 and RF2 are omitted for clarity. The SPF and GGQ motifs are highlighted in cyan, and the four functionally important residues discussed in this study (V198, F217, F221 and W319) are highlighted in magenta. Sequences of RF1 and RF2 from E. coli and T. thermophilus are highlighted in yellow.
Extended Data Figure 8 A heterologous ribosomal nonstop complex can be formed in vitro but is biologically inactive.
a, RF2 from T. thermophilus binds to E. coli nonstop ribosomal complex in vitro. The heterologous nonstop ribosomal complex consisting of RF2 from T. thermophilus and ribosomes from E. coli was formed as described in Methods. SDS–PAGE shows T. thermophilus RF2 (lane 1), apo ribosome from E. coli (lane 2) and the formation of heterologous nonstop ribosomal complex after gel filtration (lane 3). **Ribosomal protein S1. b, The heterologous ribosomal complex fails to catalyse peptide release. Representative time courses of peptide release by 5 μM E. coli RF2 and T. thermophilus RF2 at 25 nM nonstop stalled ribosome and 62.5 nM ArfA from E. coli showing that T. thermophilus RF2 fails to catalyse peptide release in the ribosome.
Rights and permissions
About this article
Cite this article
Zeng, F., Chen, Y., Remis, J. et al. Structural basis of co-translational quality control by ArfA and RF2 bound to ribosome. Nature 541, 554–557 (2017). https://doi.org/10.1038/nature21053
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21053
This article is cited by
-
Mechanism of ribosome rescue by alternative ribosome-rescue factor B
Nature Communications (2020)
-
ArfB can displace mRNA to rescue stalled ribosomes
Nature Communications (2020)
-
The structural basis for release-factor activation during translation termination revealed by time-resolved cryogenic electron microscopy
Nature Communications (2019)
-
Release factor-dependent ribosome rescue by BrfA in the Gram-positive bacterium Bacillus subtilis
Nature Communications (2019)
-
Conformation of methylated GGQ in the Peptidyl Transferase Center during Translation Termination
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.