Abstract
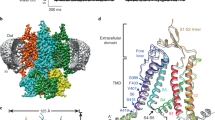
Cyclic-nucleotide-gated channels are essential for vision and olfaction. They belong to the voltage-gated ion channel superfamily but their activities are controlled by intracellular cyclic nucleotides instead of transmembrane voltage. Here we report a 3.5-Å-resolution single-particle electron cryo-microscopy structure of a cyclic-nucleotide-gated channel from Caenorhabditis elegans in the cyclic guanosine monophosphate (cGMP)-bound open state. The channel has an unusual voltage-sensor-like domain, accounting for its deficient voltage dependence. A carboxy-terminal linker connecting S6 and the cyclic-nucleotide-binding domain interacts directly with both the voltage-sensor-like domain and the pore domain, forming a gating ring that couples conformational changes triggered by cyclic nucleotide binding to the gate. The selectivity filter is lined by the carboxylate side chains of a functionally important glutamate and three rings of backbone carbonyls. This structure provides a new framework for understanding mechanisms of ion permeation, gating and channelopathy of cyclic-nucleotide-gated channels and cyclic nucleotide modulation of related channels.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Kaupp, U. B. & Seifert, R. Cyclic nucleotide-gated ion channels. Physiol. Rev. 82, 769–824 (2002)
Zagotta, W. N. & Siegelbaum, S. A. Structure and function of cyclic nucleotide-gated channels. Annu. Rev. Neurosci. 19, 235–263 (1996)
Varnum, M. D. & Dai, G. in The Handbook of Ion Channels (eds Zheng, J. & Trudeau, M. C. ) 361–382 (CRC, 2015)
Fesenko, E. E., Kolesnikov, S. S. & Lyubarsky, A. L. Induction by cyclic GMP of cationic conductance in plasma membrane of retinal rod outer segment. Nature 313, 310–313 (1985)
Yau, K. W. & Baylor, D. A. Cyclic GMP-activated conductance of retinal photoreceptor cells. Annu. Rev. Neurosci. 12, 289–327 (1989)
Nakamura, T. & Gold, G. H. A cyclic nucleotide-gated conductance in olfactory receptor cilia. Nature 325, 442–444 (1987)
Kaupp, U. B. et al. Primary structure and functional expression from complementary DNA of the rod photoreceptor cyclic GMP-gated channel. Nature 342, 762–766 (1989)
Dhallan, R. S., Yau, K. W., Schrader, K. A. & Reed, R. R. Primary structure and functional expression of a cyclic nucleotide-activated channel from olfactory neurons. Nature 347, 184–187 (1990)
Gordon, S. E. & Zagotta, W. N. A histidine residue associated with the gate of the cyclic nucleotide-activated channels in rod photoreceptors. Neuron 14, 177–183 (1995)
Gordon, S. E. & Zagotta, W. N. Localization of regions affecting an allosteric transition in cyclic nucleotide-activated channels. Neuron 14, 857–864 (1995)
Brown, R. L., Snow, S. D. & Haley, T. L. Movement of gating machinery during the activation of rod cyclic nucleotide-gated channels. Biophys. J. 75, 825–833 (1998)
Zong, X., Zucker, H., Hofmann, F. & Biel, M. Three amino acids in the C-linker are major determinants of gating in cyclic nucleotide-gated channels. EMBO J. 17, 353–362 (1998)
Zhou, L., Olivier, N. B., Yao, H., Young, E. C. & Siegelbaum, S. A. A conserved tripeptide in CNG and HCN channels regulates ligand gating by controlling C-terminal oligomerization. Neuron 44, 823–834 (2004)
Paoletti, P., Young, E. C. & Siegelbaum, S. A. C-Linker of cyclic nucleotide-gated channels controls coupling of ligand binding to channel gating. J. Gen. Physiol. 113, 17–34 (1999)
Long, S. B., Campbell, E. B. & Mackinnon, R. Crystal structure of a mammalian voltage-dependent Shaker family K+ channel. Science 309, 897–903 (2005)
Long, S. B., Campbell, E. B. & Mackinnon, R. Voltage sensor of Kv1.2: structural basis of electromechanical coupling. Science 309, 903–908 (2005)
Payandeh, J., Scheuer, T., Zheng, N. & Catterall, W. A. The crystal structure of a voltage-gated sodium channel. Nature 475, 353–358 (2011)
Wu, J. et al. Structure of the voltage-gated calcium channel Cav1.1 complex. Science 350, aad2395 (2015)
Liao, M., Cao, E., Julius, D. & Cheng, Y. Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 504, 107–112 (2013)
Paulsen, C. E., Armache, J. P., Gao, Y., Cheng, Y. & Julius, D. Structure of the TRPA1 ion channel suggests regulatory mechanisms. Nature 520, 511–517 (2015)
Komatsu, H., Mori, I., Rhee, J. S., Akaike, N. & Ohshima, Y. Mutations in a cyclic nucleotide-gated channel lead to abnormal thermosensation and chemosensation in C. elegans. Neuron 17, 707–718 (1996)
Komatsu, H. et al. Functional reconstitution of a heteromeric cyclic nucleotide-gated channel of Caenorhabditis elegans in cultured cells. Brain Res. 821, 160–168 (1999)
Long, S. B., Tao, X., Campbell, E. B. & MacKinnon, R. Atomic structure of a voltage-dependent K+ channel in a lipid membrane-like environment. Nature 450, 376–382 (2007)
Arcangeletti, M., Marchesi, A., Mazzolini, M. & Torre, V. Multiple mechanisms underlying rectification in retinal cyclic nucleotide-gated (CNGA1) channels. Physiol. Rep. 1, e00148 (2013)
Flynn, G. E. & Zagotta, W. N. Conformational changes in S6 coupled to the opening of cyclic nucleotide-gated channels. Neuron 30, 689–698 (2001)
Contreras, J. E. & Holmgren, M. Access of quaternary ammonium blockers to the internal pore of cyclic nucleotide-gated channels: implications for the location of the gate. J. Gen. Physiol. 127, 481–494 (2006)
Contreras, J. E., Srikumar, D. & Holmgren, M. Gating at the selectivity filter in cyclic nucleotide-gated channels. Proc. Natl Acad. Sci. USA 105, 3310–3314 (2008)
Goulding, E. H., Tibbs, G. R., Liu, D. & Siegelbaum, S. A. Role of H5 domain in determining pore diameter and ion permeation through cyclic nucleotide-gated channels. Nature 364, 61–64 (1993)
Eismann, E., Müller, F., Heinemann, S. H. & Kaupp, U. B. A single negative charge within the pore region of a cGMP-gated channel controls rectification, Ca2+ blockage, and ionic selectivity. Proc. Natl Acad. Sci. USA 91, 1109–1113 (1994)
Root, M. J. & MacKinnon, R. Identification of an external divalent cation-binding site in the pore of a cGMP-activated channel. Neuron 11, 459–466 (1993)
Morrill, J. A. & MacKinnon, R. Isolation of a single carboxyl-carboxylate proton binding site in the pore of a cyclic nucleotide-gated channel. J. Gen. Physiol. 114, 71–83 (1999)
Derebe, M. G. et al. Tuning the ion selectivity of tetrameric cation channels by changing the number of ion binding sites. Proc. Natl Acad. Sci. USA 108, 598–602 (2011)
Derebe, M. G., Zeng, W., Li, Y., Alam, A. & Jiang, Y. Structural studies of ion permeation and Ca2+ blockage of a bacterial channel mimicking the cyclic nucleotide-gated channel pore. Proc. Natl Acad. Sci. USA 108, 592–597 (2011)
Napolitano, L. M. et al. A structural, functional, and computational analysis suggests pore flexibility as the base for the poor selectivity of CNG channels. Proc. Natl Acad. Sci. USA 112, E3619–E3628 (2015)
Doyle, D. A. et al. The structure of the potassium channel: molecular basis of K+ conduction and selectivity. Science 280, 69–77 (1998)
Zagotta, W. N. et al. Structural basis for modulation and agonist specificity of HCN pacemaker channels. Nature 425, 200–205 (2003)
Saponaro, A. et al. Structural basis for the mutual antagonism of cAMP and TRIP8b in regulating HCN channel function. Proc. Natl Acad. Sci. USA 111, 14577–14582 (2014)
Schünke, S. & Stoldt, M. Structural snapshot of cyclic nucleotide binding domains from cyclic nucleotide-sensitive ion channels. Biol. Chem. 394, 1439–1451 (2013)
Xu, X., Vysotskaya, Z. V., Liu, Q. & Zhou, L. Structural basis for the cAMP-dependent gating in the human HCN4 channel. J. Biol. Chem. 285, 37082–37091 (2010)
Lolicato, M. et al. Tetramerization dynamics of C-terminal domain underlies isoform-specific cAMP gating in hyperpolarization-activated cyclic nucleotide-gated channels. J. Biol. Chem. 286, 44811–44820 (2011)
Taraska, J. W., Puljung, M. C., Olivier, N. B., Flynn, G. E. & Zagotta, W. N. Mapping the structure and conformational movements of proteins with transition metal ion FRET. Nature Methods 6, 532–537 (2009)
Altieri, S. L. et al. Structural and energetic analysis of activation by a cyclic nucleotide binding domain. J. Mol. Biol. 381, 655–669 (2008)
Puljung, M. C. & Zagotta, W. N. A secondary structural transition in the C-helix promotes gating of cyclic nucleotide-regulated ion channels. J. Biol. Chem. 288, 12944–12956 (2013)
Puljung, M. C., DeBerg, H. A., Zagotta, W. N. & Stoll, S. Double electron-electron resonance reveals cAMP-induced conformational change in HCN channels. Proc. Natl Acad. Sci. USA 111, 9816–9821 (2014)
Zhou, L. & Siegelbaum, S. A. Gating of HCN channels by cyclic nucleotides: residue contacts that underlie ligand binding, selectivity, and efficacy. Structure 15, 655–670 (2007)
Nair, A. V., Nguyen, C. H. & Mazzolini, M. Conformational rearrangements in the S6 domain and C-linker during gating in CNGA1 channels. Eur. Biophys. J. 38, 993–1002 (2009)
Saotome, K., Singh, A. K., Yelshanskaya, M. V. & Sobolevsky, A. I. Crystal structure of the epithelial calcium channel TRPV6. Nature 534, 506–511 (2016)
Whicher, J. R. & MacKinnon, R. Structure of the voltage-gated K+ channel Eag1 reveals an alternative voltage sensing mechanism. Science 353, 664–669 (2016)
Li, X., Zheng, S., Agard, D. A. & Cheng, Y. Asynchronous data acquisition and on-the-fly analysis of dose fractionated cryoEM images by UCSFImage. J. Struct. Biol. 192, 174–178 (2015)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nature Methods 10, 584–590 (2013)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nature Methods 11, 63–65 (2014)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014)
Acknowledgements
We thank Y. Chen and M. Chalfie (Columbia University) for providing the C. elegans cDNA library, the China National Center for Protein Sciences (Beijing) for providing facility support, and J. Cui, S. Siegelbaum, M. Zhou and Yang laboratory members for reading and commenting on the manuscript. This work was supported by grants to J.Y. from the National Key Basic Research Program of China (2014CB910301), the National Institutes of Health (R01GM085234 and RO1NS053494), the National Natural Science Foundation of China (31370821), the Top Talents Program of Yunnan Province (2011HA012) and the High-level Overseas Talents of Yunnan Province; to X.L. from the China Youth 1000-Talent Program of the State Council of China, Beijing Advanced Innovation Center for Structural Biology, Tsinghua-Peking Joint Center for Life Sciences and the National Natural Science Foundation of China (31570730); and to S.W. from the Key Research Program of the Chinese Academy of Sciences (KJZD-EW-L03), the National Natural Science Foundation of China (81302865), West Light Foundation of the Chinese Academy of Sciences, Yunnan Applied Basic Research Projects (2013FB074) and the Youth Innovation Promotion Association of the Chinese Academy of Sciences.
Author information
Authors and Affiliations
Contributions
M.L. and J.Y. conceived the project. M.L., X.Z., S.W., I.M., X.L. and J. Y. designed experiments, analysed results and wrote the manuscript. M.L. performed all molecular biological and biochemical experiments and built the atomic model. X.Z. and X.L. performed all cryo-EM experiments, including data acquisition and processing, and checked the model. S.W., Y.G., D.S. and H. L. performed electrophysiology experiments. I.M. performed confocal imaging experiments. All authors contributed to manuscript discussion, preparation and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks M. Biel, U. B. Kaupp and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Functional characterization of wild-type and mutant TAX-4 channels.
a, Family of macroscopic currents recorded in an inside-out patch from a HEK 293T cell expressing wild-type TAX-4 channels. Currents were elicited by 10 μM cGMP, in response to voltage steps from −150 mV to +150 mV in 10-mV increments from a holding potential of 0 mV. b, Current–voltage (I–V) relationship from the same patch as in a. Currents were recorded first in the control bath solution (bath), then in the presence of 10 μM cGMP, and then again in the control bath solution (washout). c, Averaged I–V relationship of macroscopic currents from seven independent inside-out patch recordings from HEK 293T cells expressing wild-type TAX-4 channels. Currents were obtained as in a and b. For each patch, currents at different voltages were normalized by the absolute current value at −150 mV. In this and subsequent figures, error bars represent s.e.m. and are smaller than the symbols at some voltages. Notice the lack of voltage-dependent gating. d, Dose–response relationship of cGMP activation of wild-type TAX-4 channels at –100 mV. Currents were recorded in inside-out patches obtained from HEK 293T cell expressing wild-type TAX-4 channels. Data points represent mean ± s.e.m. of the indicated number of measurements. Solid curve represents fit to the Hill equation in the form of I(X) = Xn/(Xn + EC50n), where I(X) is the normalized current, X the cGMP concentration, n the Hill coefficient, and EC50 the cGMP concentration producing half maximal current. The fit yields an EC50 of 0.16 μM and n of 1.7, which are very close to the values of 0.34 μM and 1.6 reported previously. e, Family of currents recorded in the whole-cell configuration from a HEK 293T cell expressing wild-type TAX-4 channels. Currents were elicited by 100 μM cGMP added in the recording pipette, in response to voltage steps from −100 mV to +150 mV in 10-mV increments from a holding potential of 0 mV. f–j, Averaged I–V relationships of whole-cell currents recorded from the indicated number of HEK 293T cell expressing wild-type TAX-4 channels and the indicated mutant TAX-4 channels. The recording pipette contained either no cGMP or 100 μM cGMP. The nonlinear I–V relationship exhibited by cGMP-activated wild-type TAX-4 channels (f) is probably caused by block by an endogenous cytoplasmic molecule. In TAX-4_5A, R421, Q425, D429, K432 and D453 in helices A′B′ are mutated to alanine. In TAX-4_1G, TAX-4_2G and TAX-4_3G, one, two or three glycine residues are inserted between M417 and S418. k, Surface expression of wild-type and mutant TAX-4 channels in HEK 293T cells. The channels contain a GFP tag on its N terminus and an HA tag in the extracellular linker between S1 and S2, except for GFP–TAX-4_WT, which does not contain the HA tag. Red fluorescence represents surface expression. Similar observations were made in more than ten cells for each channel type (including GFP–TAX-4_1G-HA and GFP–TAX-4_2G-HA) in blind experiments.
Extended Data Figure 2 Amino-acid sequence alignment of TAX-4, human CNGA1, bovine CNGA1, human HCN2 and the Kv1.2–2.1 chimaera.
Sequence alignment begins at D115 of TAX-4 and ends at L677. Green and yellow highlight identical and similar amino acids, respectively. Secondary structures are marked for TAX-4. The TAX-4 channel selectivity filter is boxed in blue. Amino acids in red in TAX-4 participate in intersubunit interactions between helices A′B′ and C′D′. Red triangles mark the amino acids involved in cGMP binding in TAX-4 and HCN2. Red dots mark the amino acids involved in forming the charge transfer centre in the Kv1.2–2.1 chimaera. Positions of S4 positive charges are boxed in red in the Kv1.2–2.1 chimaera.
Extended Data Figure 3 Single-particle cryo-EM analysis of TAX-4.
a, A representative motion-corrected micrograph of TAX-4 recorded using the K2 Summit camera. Typical particles are marked with yellow boxes. b, Fourier power spectrum of the micrograph shown in a with the Thon ring extending to 3 Å. c, Gallery of 2D class averages. d, Three enlarged views of representative 2D classes. e, Work flow of two-reference 3D classification. Structures produced by the final refinement with or without C4 imposed are given. Particle number is given below each reconstruction.
Extended Data Figure 4 Three-dimensional reconstruction and refinement of TAX-4.
a, Two isosurface levels (low is grey, high is blue) of the final density map filtered to 5 Å. The density contributed by amphipol is visible at the low isosurface level. b, Selected z slice views of the unfiltered map in a at the corresponding layers indicated by the arrows. c, FSC curve of the final 3D reconstruction with C4 imposed (black) marked with a resolution of 3.5 Å corresponding to the FSC = 0.143 cut-off criterion. FSC curve of the final 3D reconstruction without C4 imposed (red) marked with a resolution of 4.5 Å corresponding to the FSC = 0.143 cut-off criterion. The FSC curve between the final reconstruction and the map calculated from the atom model (blue) shows a resolution of 3.8 Å according to the FSC = 0.5 cut-off criterion. d, Euler angle distribution of all particles used in the final map reconstruction. Each orientation is represented by a cylinder, for which both the height and colour (from blue to red) are proportional to the number of particles for that specific direction. e–i, Colour-coded final 3D reconstruction of TAX-4 showing local resolutions. The tetramer (e) and four different orientations (f–i) of the protomer are shown, viewed parallel to the membrane. The orientation in f is the same as that in Fig. 1c.
Extended Data Figure 5 Validation of the TAX-4 model.
a, Three-dimensional reconstruction and model refinement statistics. b, FSC curves for cross-validation of the final model. Blue, model versus the summed map. Red, model versus half 1 map (called ‘work’, used for model refinement). Black, model versus half 2 map (called ‘free’, not used for model refinement).
Extended Data Figure 6 Cryo-EM density maps and atomic models of selected key TAX-4 regions.
Maps were low-pass filtered to 3.5 Å, amplified by a temperature factor of −160 Å2 and contoured at 3.0σ.
Extended Data Figure 7 Unique arrangement of the TAX-4 pore domain and VSLD.
a, Comparison of the arrangement of the pore domain (S5/P-loop/S6) and VSD (that is, S1–S4/S4–S5 linker) or VSLD of TAX-4 and the selected channels. The pore domain and VSD or VSLD of one subunit (for AtTPC1 and Cav1.1, one homologous repeat) is shown in green, and the pore domain of an adjacent subunit (or homologous repeat) is shown in red. In all comparison channels, the pore domain of the red subunit (or repeat) cross-interacts with the VSD or VSLD of the green subunit (or repeat). In TAX-4, however, the pore domain and VSLD of the green subunit form intrasubunit interactions. AtTPC1 is made up of two identical two-pore subunits. Each subunit contains two homologous six-transmembrane repeats, each of which has its own pore domain and VSD. AtTPC1(a) and AtTPC1(b) represent the two types of pore domain/VSD interaction present in AtTPC1. Cav1.1 contains four homologous six-transmembrane repeats, each of which has its own pore domain and VSD. Only one representative pore domain/VSD interaction is shown, with green and red representing repeats III and IV, respectively. The PDB accession numbers of the comparison channels are Kv1.2–2.1, 2R9R; NavAb, 3RVY; NavRh, 4DXW; Mlotik1, 3BEH; AtTPC1, 5E1J; TRPV1, 3J5P; TRPA1, 3J9P; Cav1.1, 3JBR; RyR1, 3J8H; InsP3R, 3JAV. b, Superposition of all structures in a, aligned by S5 and S6 of the green subunit. For contrast, green is changed to indigo for TAX-4. c, Comparison of the tertiary and quaternary structures of TAX-4 and the Kv1.2–2.1 chimaera, viewed from the extracellular side of the membrane. d, Interactions between the pore domain and VSLD of TAX-4, viewed parallel to the membrane (left) and from the extracellular side of the membrane (right).
Extended Data Figure 8 Structural and functional annotation of TAX-4.
a, Amino-acid sequence alignment of TAX-4 with human CNG channel subunits and TAX-2, a CNGB subunit in C. elegans. Secondary structures and selected functionally important amino acids are annotated for TAX-4. The selectivity filter is boxed in blue, and S4 positive charges are coloured in blue. Residues boxed in purple are involved in ion-pair interactions between S4 and S2–S3. Residues boxed in red in S6 and the pore helix interact with residues in or immediately adjacent to the selectivity filter. Residues boxed in green participate in intersubunit interactions between helices A′B′ and C′D′. Residues boxed in orange are engaged in interactions between helices A′B′ of one subunit and S4/S4–S5 linker/S5 of an adjacent subunit. Positions of single-amino-acid missense mutations that cause retinitis pigmentosa are highlighted in red in hCNGA1 and orange in hCNGB1, and those that cause achromatopsia are highlighted in cyan in hCNGA3 and green in hCNGB3. The disease-causing mutations are listed and colour-coded on both sides of the sequences. TAX-2 does not form functional homomeric channels but associates with TAX-4 to form functional heteromeric channels, both in heterologous expression systems and native cells. b, c, Mapping the disease-causing mutations listed in a on the TAX-4 protomer structure, shown in the same orientation here as in Fig. 1c.
Extended Data Figure 9 The TAX-4 channel selectivity filter.
a, Electrostatic surface representation of TAX-4, viewed from the extracellular side of the membrane, showing a highly electronegative external entrance to the selectivity filter in the centre. b, Comparison of the selectivity filter of TAX-4 and the selected Ca2+-conducting channels. For clarity, only two diagonally opposed subunits are shown. Cav1.1(a) and Cav1.1(b) represent repeats I/III and II/IV of Cav1.1, respectively. Note the reoccurring utilization of a combination of negative side chains and backbone carbonyls to line the selectivity filter. The PDB accession numbers of the comparison channels are Cav1.1, 3JBR; RyR1, 3J8H; TRPV1, 3J5P; TRPA1, 3J9P. c, Superposition of the selectivity filter of TAX-4 and NaK2CNG-E (PDB accession number 3K0G). Only two diagonally opposed subunits are shown. Purple spheres mark presumed ion-binding sites in TAX-4.
Extended Data Figure 10 The TAX-4 channel gating ring.
a, The gating ring, formed by helices A′B′C′D′ of the C-linker, is depicted in cylinder form and shown in the TAX-4 structure, viewed parallel to the membrane. b, The gating ring shown in isolation, viewed from the extracellular side of the membrane. c, A composite figure demonstrating that helix D′ of the gating ring must change its conformation in the unliganded state. The figure was generated in two steps: (1) the liganded structure of the TAX-4 C-linker/CNBD and the unliganded structure of the HCN2 C-linker/CNBD (PDB accession number 2MPF) were aligned by the β-strands of the CNBD, and only the indicated α-helices are shown for comparison; (2) helices A, B, C and E′ of TAX-4 were aligned with those of HCN2, producing a hypothetical unliganded TAX-4 structure. The resulting helix D′, however, clashes with helix A, as shown in e. d, Space-filling model of the interface between helices A and D′ in the liganded TAX-4 structure, showing a snug fit between these helices. e, Space-filling model of the interface between helices A and D′ in the hypothetical unliganded TAX-4 structure generated in c, showing a clash between these helices, indicating that helix D′ must adopt a different conformation.
Rights and permissions
About this article
Cite this article
Li, M., Zhou, X., Wang, S. et al. Structure of a eukaryotic cyclic-nucleotide-gated channel. Nature 542, 60–65 (2017). https://doi.org/10.1038/nature20819
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20819
This article is cited by
-
Fast functional mapping of ligand-gated ion channels
Communications Biology (2023)
-
Conformational trajectory of allosteric gating of the human cone photoreceptor cyclic nucleotide-gated channel
Nature Communications (2023)
-
Discrimination between cyclic nucleotides in a cyclic nucleotide-gated ion channel
Nature Structural & Molecular Biology (2023)
-
Gating intermediates reveal inhibitory role of the voltage sensor in a cyclic nucleotide-modulated ion channel
Nature Communications (2022)
-
Structural and functional characterization of an achromatopsia-associated mutation in a phototransduction channel
Communications Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.