Abstract
CD4+ T cells are central mediators of autoimmune pathology; however, defining their key effector functions in specific autoimmune diseases remains challenging. Pathogenic CD4+ T cells within affected tissues may be identified by expression of markers of recent activation1. Here we use mass cytometry to analyse activated T cells in joint tissue from patients with rheumatoid arthritis, a chronic immune-mediated arthritis that affects up to 1% of the population2. This approach revealed a markedly expanded population of PD-1hiCXCR5−CD4+ T cells in synovium of patients with rheumatoid arthritis. However, these cells are not exhausted, despite high PD-1 expression. Rather, using multidimensional cytometry, transcriptomics, and functional assays, we define a population of PD-1hiCXCR5− ‘peripheral helper’ T (TPH) cells that express factors enabling B-cell help, including IL-21, CXCL13, ICOS, and MAF. Like PD-1hiCXCR5+ T follicular helper cells, TPH cells induce plasma cell differentiation in vitro through IL-21 secretion and SLAMF5 interaction (refs 3, 4). However, global transcriptomics highlight differences between TPH cells and T follicular helper cells, including altered expression of BCL6 and BLIMP1 and unique expression of chemokine receptors that direct migration to inflamed sites, such as CCR2, CX3CR1, and CCR5, in TPH cells. TPH cells appear to be uniquely poised to promote B-cell responses and antibody production within pathologically inflamed non-lymphoid tissues.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Maecker, H. T., McCoy, J. P. & Nussenblatt, R. Standardizing immunophenotyping for the Human Immunology Project. Nat. Rev. Immunol. 12, 191–200 (2012)
McInnes, I. B. & Schett, G. The pathogenesis of rheumatoid arthritis. NEJM 365, 2205–2219 (2011)
Crotty, S. Follicular helper CD4 T cells (TFH). Annu. Rev. Immunol. 29, 621–663 (2011)
Cannons, J. L. et al. Optimal germinal center responses require a multistage T cell:B cell adhesion process involving integrins, SLAM-associated protein, and CD84. Immunity 32, 253–265 (2010)
Amir, A. D. et al. viSNE enables visualization of high dimensional single-cell data and reveals phenotypic heterogeneity of leukemia. Nat. Biotechnol. 31, 545–552 (2013)
Takemura, S. et al. Lymphoid neogenesis in rheumatoid synovitis. J. Immunol. 167, 1072–1080 (2001)
Humby, F. et al. Ectopic lymphoid structures support ongoing production of class-switched autoantibodies in rheumatoid synovium. PLoS Med. 6, e1 (2009)
Wherry, E. J. & Kurachi, M. Molecular and cellular insights into T cell exhaustion. Nat. Rev. Immunol. 15, 486–499 (2015)
Kamphorst, A. O. & Ahmed, R. Manipulating the PD-1 pathway to improve immunity. Curr. Opin. Immunol. 25, 381–388 (2013)
Johnston, R. J. et al. Bcl6 and Blimp-1 are reciprocal and antagonistic regulators of T follicular helper cell differentiation. Science 325, 1006–1010 (2009)
Kroenke, M. A. et al. Bcl6 and Maf cooperate to instruct human follicular helper CD4 T cell differentiation. J. Immunol. 188, 3734–3744 (2012)
DeGrendele, H. C., Estess, P. & Siegelman, M. H. Requirement for CD44 in activated T cell extravasation into an inflammatory site. Science 278, 672–675 (1997)
Förster, R. et al. CCR7 coordinates the primary immune response by establishing functional microenvironments in secondary lymphoid organs. Cell 99, 23–33 (1999)
Chtanova, T. et al. T follicular helper cells express a distinctive transcriptional profile, reflecting their role as non-TH1/TH2 effector cells that provide help for B cells. J. Immunol. 173, 68–78 (2004)
Locci, M. et al. Human circulating PD-1+CXCR3−CXCR5+ memory TFH cells are highly functional and correlate with broadly neutralizing HIV antibody responses. Immunity 39, 758–769 (2013)
Weinstein, J. S. et al. Global transcriptome analysis and enhancer landscape of human primary T follicular helper and T effector lymphocytes. Blood 124, 3719–3729 (2014)
Kenefeck, R. et al. Follicular helper T cell signature in type 1 diabetes. J. Clin. Invest. 125, 292–303 (2015)
Kuziel, W. A. et al. Severe reduction in leukocyte adhesion and monocyte extravasation in mice deficient in CC chemokine receptor 2. Proc. Natl Acad. Sci. USA 94, 12053–12058 (1997)
Rot, A. & von Andrian, U. H. Chemokines in innate and adaptive host defense: basic chemokinese grammar for immune cells. Annu. Rev. Immunol. 22, 891–928 (2004)
Vu Van, D. et al. Local T/B cooperation in inflamed tissues is supported by T follicular helper-like cells. Nat. Commun. 7, 10875 (2016)
Pitzalis, C., Jones, G. W., Bombardieri, M. & Jones, S. A. Ectopic lymphoid-like structures in infection, cancer and autoimmunity. Nat. Rev. Immunol. 14, 447–462 (2014)
Kobayashi, S. et al. A distinct human CD4+ T cell subset that secretes CXCL13 in rheumatoid synovium. Arthritis Rheum. 65, 3063–3072 (2013)
Manzo, A. et al. Mature antigen-experienced T helper cells synthesize and secrete the B cell chemoattractant CXCL13 in the inflammatory environment of the rheumatoid joint. Arthritis Rheum. 58, 3377–3387 (2008)
Shen, P. & Fillatreau, S. Antibody-independent functions of B cells: a focus on cytokines. Nat. Rev. Immunol. 15, 441–451 (2015)
Finck, R. et al. Normalization of mass cytometry data with bead standards. Cytometry A 83, 483–494 (2013)
Bray, N., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal RNA-Seq quantification. Preprint at https://arxiv.org/abs/1505.02710 (2015)
Acknowledgements
This work was supported by T32 AR007530-31 and the William Docken Inflammatory Autoimmune Disease Fund (to M.B.B.), Mallinckrodt Research Fellowship (to D.A.R.), R01 AR064850-03 (to Y.C.L.), NIH 5U01GM092691-05, 1U19 AI111224-01 and Doris Duke Charitable Foundation Grant #2013097 (to S.R.), Rheumatology Research Foundation Scientist Development Award (to L.A.H.), K01 AR066063 (to L.T.D.), Arthritis Research UK programme grant #19791 (to C.D.B.), and Arthritis Research UK Clinician Scientist Fellowship #18547 (to A.F.). J.L.M was supported by the FP7-HEALTH-F2-2012-305549 EuroTEAM. P.A.N. was supported by P30 AR070253 and the Fundación Bechara. We thank A. Chicoine and the BWH Human Immunology Center Flow Cytometry Core for assistance with cell sorting.
Author information
Authors and Affiliations
Contributions
D.A.R conceived the project, performed experiments, analysed data, and wrote the manuscript. M.F.G., Y.L., N.T., and F.M. performed experiments and analysed data. K.S. analysed transcriptomic data. C.F. analysed mass cytometry data. J.L.M. performed immunofluorescence microscopy. J.A.L. assisted with mass cytometry. K.W., L.A.H., P.A.N., M.E.W., Y.C.L., J.S.C., D.J.T., E.M.M., S.M.H., E.W.K., L.T.D., V.P.B., L.B.I., S.M.G., A.B.P., A.F. and C.D.B participated in study design, patient recruitment and sample acquisition. M.B.B. and S.R. conceived the project, supervised the work, analysed data, and co-wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks J. Craft, S. Fillatreau and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Detection of PD-1hiCD4+ T cells in RA synovial tissue and fluid by mass and flow cytometry.
a, viSNE plots of mass cytometry of RA synovial tissue CD4+ T cells as in Fig. 1a from 2 additional donors. b, Gating strategy to identify synovial tissue PD-1hi CD4+ T cell populations by mass cytometry. c, viSNE plots of flow cytometry of paired RA synovial fluid and blood memory CD4+ T cells. d, Gating strategy to identify synovial fluid PD-1hi memory CD4+ T cells by flow cytometry. e, Examples of gating used to sort memory CD4+ T cell populations from patient samples. f, Detection of CXCR5 mRNA by RT–PCR in sorted memory CD4+ T-cell populations from synovial tissue (n = 3 donors, 2 of which provided sufficient PD-1hi CXCR5+ cells for analysis), synovial fluid (n = 3 donors, 1 of which provided sufficient PD-1hi CXCR5+ cells for analysis), and blood (n = 2 donors). Purple boxes indicate PD-1− and PD-1hi CXCR5+ cells sorted from human tonsil as controls. Lines in f indicate mean for synovial or blood samples.
Extended Data Figure 2 PD-1hiCXCR5−CD4+ T cells are expanded in circulation of patients with active, seropositive RA and decrease with response to therapy.
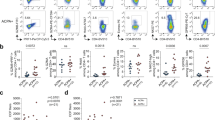
a, Mean expression of MHC II and ICOS in memory CD4+ T cell populations from synovial tissue (n = 10), synovial fluid (n = 9), and blood (n = 42) from patients with seropositive RA. Mean ± s.d. shown. b, Flow cytometric detection of PD-1 and CXCR5 expression on blood memory CD4+ T cells. c–e, Frequency of PD-1hi cells that co-express MHC II or ICOS (c), PD-1hi CXCR5+ cells (d) or cells with intermediate PD-1 expression (e) within memory CD4+ T cells from blood of patients with seropositive RA (n = 42), seronegative RA (n = 16), spondyloarthropathy (SpA, n = 11), and non-inflammatory controls (n = 35). f, Correlation between age or disease duration and blood PD-1hiCXCR5− cell frequency in seropositive RA patients (n = 38). g, PD-1hiCXCR5− cell frequencies in seropositive RA patients segregated based on sex or medication usage (n = 38). h, Correlation between serum anti-CCP antibody titer and blood PD-1hiCXCR5− cell frequency in all RA patients (n = 53, black line, P = 0.0049) or in only anti-CCP antibody+ patients (n = 29, green line, P = 0.48). i, Correlation between fold change in CDAI and fold change in PD-1hiCXCR5− cell frequency in patients 3 months after addition of a new RA medication (n = 23; methotrexate, 11; anti-TNF, 4; abatacept, 4; tocilizumab, 2; tofacitinib, 2). j, Frequency of PD-1hi T-cell subpopulations in blood before and after RA treatment escalation in 18 patients with reduced disease activity after therapy. Median ± interquartile range (c–e); mean ± s.d. (a, g) shown. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001 by Mann–Whitney (c, g), Kruskal–Wallis (d, e), Wilcoxon test (j). In f, h, i, P values calculated by Spearman correlation.
Extended Data Figure 3 Blood PD-1hiCXCR5−CD4+ T cells express factors associated with B-cell help.
a, RT–PCR for intracellular regulators in memory CD4+ T cell populations from RA synovial fluid (n = 5 or 6 donors). b, RT–PCR for cytokines (n = 10 donors, 6 RA patients (black), 4 controls (grey)) and intracellular regulators (n = 4 or 5 donors) in memory CD4+ T cell populations from blood. c, Cytokine and transcription factor mRNA expression in blood PD-1hi memory CD4+ T cell populations divided according to CXCR5 expression, relative to PD-1− cells (n = 6 donors). d, Flow cytometric quantification of BCL6 and BLIMP1 in blood PD-1hi memory CD4+ T cell subpopulations sorted according to chemokine receptor expression, then stimulated in vitro for 2 days with anti-CD3/CD28 beads. Representative data from 1 of 3 experiments using cells from different donors. Median ± interquartile range (a, b); mean ± s.d. (c, d) shown. *P < 0.05, **P < 0.01, ***P < 0.001 by Friedman’s test, compared to PD-1–MHC-II− group (a, b).
Extended Data Figure 4 Identification and characterization of circulating PD-1hiCXCR5− and PD-1hiCXCR5+ in mass cytometry and RNA-seq analyses.
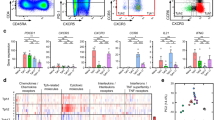
a, Gating of blood PD-1hi memory CD4+ T cells in mass cytometry analyses. b, Flow cytometric detection of FOXP3 and PD-1 in blood memory CD4+ T cells from patients with RA (black, n = 5) and controls (grey, n = 3). c, Flow cytometric detection of inhibitory receptors on blood CXCR5− memory CD4+ T cells. Data from 1 of 3 patients with RA with similar results. d, Sorting strategy for PD-1hiCXCR5− and PD-1hiCXCR5+ cell populations for RNA-seq. e, Hierarchical clustering T-cell subpopulations sorted as in d, with clustering based on expression of TFH-associated genes measured by RNA-seq. f, Chemokine receptor expression on blood memory CD4+ T cells from patients with RA (black) or controls (grey) by flow cytometry. Mean ± s.d. shown. *P < 0.05, **P < 0.001, ***P < 0.0001 by Kruskal–Wallis test compared to PD-1− cells (b) or Wilcoxon test (f).
Extended Data Figure 5 Limited interconversion of PD-1hiCCR2+ and PD-1hiCXCR5+ T cells in vitro.
a, Flow cytometry of CXCR5 and CCR2 on gated PD-1hi memory CD4+ T cells from blood. b, Expression of CXCR5 and CCR2 on indicated sorted PD-1hi T-cell populations 7 days after in vitro stimulation with anti-CD3/CD28 beads. c, d, Percentage of cells from each sorted PD-1hi population that expressed CXCR5 or CCR2 on day 2 (c) or day 7 (d) after in vitro stimulation. Naive CD4+ T cells are shown as control. Mean ± s.d. shown (n = 3 donors from 3 separate experiments).
Extended Data Figure 6 SLAMF5 is required for B-cell-helper function of PD-1hiCXCR5−CD4+ T cells.
a, Flow cytometric quantification of SLAM, SLAMF5, and SLAMF6 expression on memory CD4+ T cells (n = 10 donors; 5 patients with RA, 5 controls). b, Quantification of frequency of memory B cells with plasma cell markers after co-culture with PD-1hiCXCR5+CD4+ T cells with addition of blocking antibodies against SLAMF5 and/or SLAMF6. c, IgG quantification by ELISA in co-cultures of memory B cells with PD-1hiCXCR5− or PD-1hiCXCR5+ T cells with addition of blocking antibodies against SLAMF5 and/or SLAMF6. For b, c, 1 of 3 experiments with similar results (n = 3 replicates shown). Mean ± s.d. shown. *P < 0.05, **P < 0.01, ***P < 0.001 by Kruskal–Wallis compared to PD-1−CXCR5− (a) or isotype control (b, c). d, Immunofluorescence microscopy of CD20 (green), CXCR5 (red), and PD-1 (blue), in seropositive RA synovial tissue. Arrows point to PD-1hiCXCR5− cells adjacent to B cells. Scale bar, 50 μm.
Rights and permissions
About this article
Cite this article
Rao, D., Gurish, M., Marshall, J. et al. Pathologically expanded peripheral T helper cell subset drives B cells in rheumatoid arthritis. Nature 542, 110–114 (2017). https://doi.org/10.1038/nature20810
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20810
This article is cited by
-
Single-cell characterisation of tissue homing CD4 + and CD8 + T cell clones in immune-mediated refractory arthritis
Molecular Medicine (2024)
-
The response of CD27+CD38+ plasmablasts, CD24hiCD38hi transitional B cells, CXCR5−ICOS+PD-1+ Tph, Tph2 and Tfh2 subtypes to allergens in children with allergic asthma
BMC Pediatrics (2024)
-
Single-cell insights into immune dysregulation in rheumatoid arthritis flare versus drug-free remission
Nature Communications (2024)
-
The regulation and differentiation of regulatory T cells and their dysfunction in autoimmune diseases
Nature Reviews Immunology (2024)
-
Cytotoxic Tph subset with low B-cell helper functions and its involvement in systemic lupus erythematosus
Communications Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.