Abstract
The human cannabinoid G-protein-coupled receptors (GPCRs) CB1 and CB2 mediate the functional responses to the endocannabinoids anandamide and 2-arachidonyl glycerol (2-AG) and to the widely consumed plant phytocannabinoid Δ9-tetrahydrocannabinol (THC)1. The cannabinoid receptors have been the targets of intensive drug discovery efforts, because modulation of these receptors has therapeutic potential to control pain2, epilepsy3, obesity4, and other disorders. Although much progress in understanding the biophysical properties of GPCRs has recently been made, investigations of the molecular mechanisms of the cannabinoids and their receptors have lacked high-resolution structural data. Here we report the use of GPCR engineering and lipidic cubic phase crystallization to determine the structure of the human CB1 receptor bound to the inhibitor taranabant at 2.6-Å resolution. We found that the extracellular surface of CB1, including the highly conserved membrane-proximal N-terminal region, is distinct from those of other lipid-activated GPCRs, forming a critical part of the ligand-binding pocket. Docking studies further demonstrate how this same pocket may accommodate the cannabinoid agonist THC. Our CB1 structure provides an atomic framework for studying cannabinoid receptor function and will aid the design and optimization of therapeutic modulators of the endocannabinoid system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Mechoulam, R. & Parker, L. A. The endocannabinoid system and the brain. Annu. Rev. Psychol. 64, 21–47 (2013)
Lynch, M. E. & Ware, M. A. Cannabinoids for the treatment of chronic non-cancer pain: an updated systematic review of randomized controlled trials. J. Neuroimmune Pharmacol. 10, 293–301 (2015)
Reddy, D. S. & Golub, V. M. The pharmacological basis of cannabis therapy for epilepsy. J. Pharmacol. Exp. Ther. 357, 45–55 (2016)
Kim, J., Li, Y. & Watkins, B. A. Endocannabinoid signaling and energy metabolism: a target for dietary intervention. Nutrition 27, 624–632 (2011)
Pertwee, R. G. The diverse CB1 and CB2 receptor pharmacology of three plant cannabinoids: Δ9-tetrahydrocannabinol, cannabidiol and Δ9-tetrahydrocannabivarin. Br. J. Pharmacol. 153, 199–215 (2008)
Vemuri, V. K. & Makriyannis, A. Medicinal chemistry of cannabinoids. Clin. Pharmacol. Ther. 97, 553–558 (2015)
Howlett, A. C. et al. Cannabinoid physiology and pharmacology: 30 years of progress. Neuropharmacology 47 (Suppl. 1), 345–358 (2004)
Wilson, R. I. & Nicoll, R. A. Endocannabinoid signaling in the brain. Science 296, 678–682 (2002)
Fowler, C. J. Transport of endocannabinoids across the plasma membrane and within the cell. FEBS J. 280, 1895–1904 (2013)
Tam, J. et al. Peripheral cannabinoid-1 receptor inverse agonism reduces obesity by reversing leptin resistance. Cell Metab. 16, 167–179 (2012)
Jourdan, T. et al. Activation of the Nlrp3 inflammasome in infiltrating macrophages by endocannabinoids mediates beta cell loss in type 2 diabetes. Nat. Med. 19, 1132–1140 (2013)
Gaoni, Y. & Mechoulam, R. Isolation, structure, and partial synthesis of an active constituent of hashish. J. Am. Chem. Soc. 86, 1646–1647 (1964)
Janero, D. R. & Makriyannis, A. Cannabinoid receptor antagonists: pharmacological opportunities, clinical experience, and translational prognosis. Expert Opin. Emerg. Drugs 14, 43–65 (2009)
Yin, J. et al. Structure and ligand-binding mechanism of the human OX1 and OX2 orexin receptors. Nat. Struct. Mol. Biol. 23, 293–299 (2016)
D’Antona, A. M., Ahn, K. H. & Kendall, D. A. Mutations of CB1 T210 produce active and inactive receptor forms: correlations with ligand affinity, receptor stability, and cellular localization. Biochemistry 45, 5606–5617 (2006)
González-Mariscal, I. et al. Human CB1 receptor isoforms, present in hepatocytes and β-cells, are involved in regulating metabolism. Sci. Rep. 6, 33302 (2016)
Andersson, H., D’Antona, A. M., Kendall, D. A., Von Heijne, G. & Chin, C. N. Membrane assembly of the cannabinoid receptor 1: impact of a long N-terminal tail. Mol. Pharmacol. 64, 570–577 (2003)
Rosenbaum, D. M. et al. GPCR engineering yields high-resolution structural insights into β2-adrenergic receptor function. Science 318, 1266–1273 (2007)
Hanson, M. A. et al. Crystal structure of a lipid G protein-coupled receptor. Science 335, 851–855 (2012)
Hurst, D. P. et al. A lipid pathway for ligand binding is necessary for a cannabinoid G protein-coupled receptor. J. Biol. Chem. 285, 17954–17964 (2010)
Fay, J. F. & Farrens, D. L. The membrane proximal region of the cannabinoid receptor CB1 N-terminus can allosterically modulate ligand affinity. Biochemistry 52, 8286–8294 (2013)
Chrencik, J. E. et al. Crystal structure of antagonist bound human lysophosphatidic acid receptor 1. Cell 161, 1633–1643 (2015)
Palczewski, K. et al. Crystal structure of rhodopsin: A G protein-coupled receptor. Science 289, 739–745 (2000)
Park, J. H., Scheerer, P., Hofmann, K. P., Choe, H.-W. & Ernst, O. P. Crystal structure of the ligand-free G-protein-coupled receptor opsin. Nature 454, 183–187 (2008)
Fong, T. M. et al. Antiobesity efficacy of a novel cannabinoid-1 receptor inverse agonist, N-[(1S,2S)-3-(4-chlorophenyl)-2-(3-cyanophenyl)-1-methylpropyl]-2-methyl-2-[[5-(trifluoromethyl)pyridin-2-yl]oxy]propanamide (MK-0364), in rodents. J. Pharmacol. Exp. Ther. 321, 1013–1022 (2007)
Kapur, A. et al. Mutation studies of Ser7.39 and Ser2.60 in the human CB1 cannabinoid receptor: evidence for a serine-induced bend in CB1 transmembrane helix 7. Mol. Pharmacol. 71, 1512–1524 (2007)
Lin, L. S. et al. Conformational analysis and receptor docking of N-[(1S,2S)-3-(4-chlorophenyl)-2-(3-cyanophenyl)-1-methylpropyl]-2-methyl-2-[5-(trifluoromethyl)pyridin-2-yl]oxypropanamide (taranabant, MK-0364), a novel, acyclic cannabinoid-1 receptor inverse agonist. J. Med. Chem. 51, 2108–2114 (2008)
Shim, J.-Y., Bertalovitz, A. C. & Kendall, D. A. Probing the interaction of SR141716A with the CB1 receptor. J. Biol. Chem. 287, 38741–38754 (2012)
Sitkoff, D. F. et al. Cannabinoid CB(1) receptor ligand binding and function examined through mutagenesis studies of F200 and S383. Eur. J. Pharmacol. 651, 9–17 (2011)
Hurst, D. P. et al. N-(piperidin-1-yl)-5-(4-chlorophenyl)-1-(2,4-dichlorophenyl)-4-methyl-1H-pyrazole-3-carboxamide (SR141716A) interaction with LYS 3.28(192) is crucial for its inverse agonism at the cannabinoid CB1 receptor. Mol. Pharmacol. 62, 1274–1287 (2002)
McAllister, S. D. et al. Structural mimicry in class A G protein-coupled receptor rotamer toggle switches: the importance of the F3.36(201)/W6.48(357) interaction in cannabinoid CB1 receptor activation. J. Biol. Chem. 279, 48024–48037 (2004)
Shim, J.-Y., Bertalovitz, A. C. & Kendall, D. A. Identification of essential cannabinoid-binding domains: structural insights into early dynamic events in receptor activation. J. Biol. Chem. 286, 33422–33435 (2011)
Picone, R. P. et al. (−)-7′-Isothiocyanato-11-hydroxy-1,1′-dimethylheptylhexahydrocannabinol (AM841), a high-affinity electrophilic ligand, interacts covalently with a cysteine in helix six and activates the CB1 cannabinoid receptor. Mol. Pharmacol. 68, 1623–1635 (2005)
Hua, T. et al. Crystal structure of the human cannabinoid receptor CB1. Cell 167, 750–762.e14 (2016)
Manglik, A. et al. Structural insights into the dynamic process of β2-adrenergic receptor signaling. Cell 161, 1101–1111 (2015)
Console-Bram, L., Marcu, J. & Abood, M. E. Cannabinoid receptors: nomenclature and pharmacological principles. Prog. Neuropsychopharmacol. Biol. Psychiatry 38, 4–15 (2012)
Price, M. R. et al. Allosteric modulation of the cannabinoid CB1 receptor. Mol. Pharmacol. 68, 1484–1495 (2005)
Hofmann, L., Gulati, S., Sears, A., Stewart, P. L. & Palczewski, K. An effective thiol-reactive probe for differential scanning fluorimetry with a standard real-time polymerase chain reaction device. Anal. Biochem. 499, 63–65 (2016)
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nat. Protocols 4, 706–731 (2009)
Otwinowski, Z. & Minor, W. Processing of X-ray data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Horcajada, C., Guinovart, J. J., Fita, I. & Ferrer, J. C. Crystal structure of an archaeal glycogen synthase: insights into oligomerization and substrate binding of eukaryotic glycogen synthases. J. Biol. Chem. 281, 2923–2931 (2006)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Skubák, P., Murshudov, G. N. & Pannu, N. S. Direct incorporation of experimental phase information in model refinement. Acta Crystallogr. D 60, 2196–2201 (2004)
Schüttelkopf, A. W. & van Aalten, D. M. F. PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr. D 60, 1355–1363 (2004)
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001)
Case, D. A. et al. The Amber biomolecular simulation programs. J. Comput. Chem. 26, 1668–1688 (2005)
Izaguirre, J. A., Catarello, D. P., Wozniak, J. M. & Skeel, R. D. Langevin stabilization of molecular dynamics. J. Chem. Phys. 114, 2090–2098 (2001)
Friesner, R. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004)
Friesner, R. A. et al. Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J. Med. Chem. 49, 6177–6196 (2006)
Acknowledgements
We thank the staff of the GM/CA-CAT beamline 23ID at the Advanced Photon Source (APS) for support during data collection. This project was supported by Welch Foundation grant (I-1770 to D.M.R) and a Packard Foundation Fellowship (D.M.R.). APS is a US Department of Energy Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory (DE-AC02-06CH11357).
Author information
Authors and Affiliations
Contributions
Z.S. developed the CB1 construct and purification; expressed, purified and crystallized the receptor; collected diffraction data; and solved and refined the structure. J.Y. assisted with crystallographic refinement. K.C. performed ligand binding assays on CB1 constructs. M.G. carried out computational docking calculations. L.C. assisted with construct design and purification. J.W. performed and supervised computational docking calculations and molecular dynamics simulations. D.M.R supervised the overall project, assisted with collection of diffraction data, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks A. Christopoulos, G. Kunos and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Differential scanning fluorimetry on purified CB1–PGS.
a, Raw differential scanning fluorimetry traces of the receptor in the apo state or bound to each antagonist. b, First derivative analysis of data in a.
Extended Data Figure 2 Ligand-binding properties of CB1 constructs.
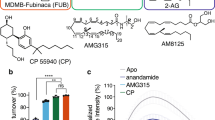
a, Saturation binding of the antagonist [3H]SR141716A (tritiated rimonabant radioligand) to wild-type CB1, CB1–PGS, and CB1(T210A)–PGS. Error bars represent s.d. for three separate experiments, each performed in duplicate. The fitted Kd values (± s.e.m.) for these three constructs are 4.8 ± 0.7 nM, 6.3 ± 0.6 nM, and 4.4 ± 0.5 nM, respectively. b, Competition binding of taranabant to the wild-type CB1 receptor, CB1–PGS, and CB1(T210A)–PGS. Error bars represent s.d. for three separate experiments, each performed in duplicate. The Ki values (± s.e.m.) of the three constructs for taranabant are 0.94 ± 0.17 nM, 1.10 ± 0.16 nM, and 0.91 ± 0.16 nM, respectively. c, Competition binding of the agonist CP55940 to the wild-type CB1 receptor, CB1–PGS, and CB1(T210A)–PGS. Error bars represent s.d. for three separate experiments, each performed in duplicate. The Ki values (± s.e.m.) of the three constructs for CP55940 are 53 ± 12 nM, 230 ± 43 nM, and 384 ± 62 nM, respectively.
Extended Data Figure 3 Purification and crystallization of CB1(T210A)–PGS.
a, Superdex 200 gel-filtration trace of receptor after Ni immobilized metal-affinity chromatography (IMAC) and M1 anti-Flag chromatography (see Methods). b, SDS–PAGE analysis of samples at different stages of purification. The five lanes from left to right are: markers (molecular mass in kDa at left); IMAC/Flag-purified receptor; same sample after treatment with PNGaseF; receptor after TEV protease cleavage (removing 89 N-terminal amino acids); final sample after Superdex 200 gel filtration. c, Light microscopy image showing examples of LCP microcrystals of CB1(T210A)–PGS used to collect diffraction data.
Extended Data Figure 4 Packing and electron density in the CB1(T210A)–PGS crystals.
a, Lattice packing interactions in the monoclinic crystals of CB1(T210A)–PGS. Protomers are shown as ribbons, with the receptor component of the fusion protein coluored teal and the PGS domain coloured grey. b, 2Fo − Fc electron density map (contoured at 1.2σ) of taranabant and the surrounding ligand-binding residues. Protein and ligand are represented as yellow sticks. c, Stereo view of 2Fo − Fc electron density (contoured at 1.5σ) for only the ligand taranabant (magenta sticks).
Extended Data Figure 5 Residues lining the putative lipid access channel of CB1.
The receptor is shown as a teal transparent surface, and taranabant is in magenta spheres. The three residues lining the channel are shown as orange sticks and their solvent-accessible surfaces are coloured orange.
Extended Data Figure 6 Sequence alignment of the membrane-proximal N-terminal region of CB1 from different vertebrate species.
‘Frog’ is Xenopus laevis. The red bar (top) indicates the part of this region that is structured and visible in the electron density of the CB1 crystals. The blue box denotes positions that make contact with taranabant. Alignment was performed using Clustal Omega (https://www.ebi.ac.uk/Tools/msa/clustalo/).
Extended Data Figure 7 Molecular dynamics simulation of the CB1 structure.
a, A 60-ns molecular dynamics simulation of the CB1 receptor (after removing the PGS fusion protein) with taranabant present. Black trace is for the entire receptor, red trace is for only the structured membrane-proximal N-terminal region. b, 60-ns molecular dynamics simulation of the CB1 receptor without a ligand present. Black trace is for the entire receptor, red trace is for only the structured membrane-proximal N-terminal region.
Extended Data Figure 8 Sequence alignment of the entire sequence of CB1 from several different species, along with human CB2.
The blue boxes denote positions that make contact with taranabant within a 4 Å cut-off. The alignment was performed using Clustal Omega (https://www.ebi.ac.uk/Tools/msa/clustalo/).
Extended Data Figure 9 Comparison of the structures of CB1 bound to taranabant and CB1 bound to AM6538 (ref. 34; PDB accession 5TGZ).
a, Superposition of the two CB1 structures viewed from the extracellular space. The taranabant-bound structure is shown as a teal cartoon (ligand as magenta sticks), while the AM6538-bound structure is shown as a gold cartoon (ligand as green sticks). b, Comparison of 2Fo − Fc electron density (contoured at 1.5σ) for the ligands in each structure. On the left is taranabant from the current structure, on the right is AM6538 from ref. 34. c, Comparison of the membrane-proximal N-terminal regions in each structure. On the left is a side view of CB1 from the current structure, with 2Fo − Fc electron density (contoured at 1.0σ) shown for the N-terminal region, TM1, and taranabant. On the right is the analogous side view of CB1 from ref. 34 (gold cartoon), with 2Fo − Fc electron density (contoured at 1.0σ) shown for the N-terminal region, TM1 and AM6538.
Rights and permissions
About this article
Cite this article
Shao, Z., Yin, J., Chapman, K. et al. High-resolution crystal structure of the human CB1 cannabinoid receptor. Nature 540, 602–606 (2016). https://doi.org/10.1038/nature20613
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20613
This article is cited by
-
Distinct activation mechanisms regulate subtype selectivity of Cannabinoid receptors
Communications Biology (2023)
-
Orthosteric ligand selectivity and allosteric probe dependence at Hydroxycarboxylic acid receptor HCAR2
Signal Transduction and Targeted Therapy (2023)
-
A fluorescent sensor for spatiotemporally resolved imaging of endocannabinoid dynamics in vivo
Nature Biotechnology (2022)
-
Structural insights into sphingosine-1-phosphate recognition and ligand selectivity of S1PR3–Gi signaling complexes
Cell Research (2022)
-
Activation and signaling mechanism revealed by GPR119-Gs complex structures
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.