Abstract
Autism spectrum disorder (ASD) involves substantial genetic contributions. These contributions are profoundly heterogeneous but may converge on common pathways that are not yet well understood1,2,3. Here, through post-mortem genome-wide transcriptome analysis of the largest cohort of samples analysed so far, to our knowledge4,5,6,7, we interrogate the noncoding transcriptome, alternative splicing, and upstream molecular regulators to broaden our understanding of molecular convergence in ASD. Our analysis reveals ASD-associated dysregulation of primate-specific long noncoding RNAs (lncRNAs), downregulation of the alternative splicing of activity-dependent neuron-specific exons, and attenuation of normal differences in gene expression between the frontal and temporal lobes. Our data suggest that SOX5, a transcription factor involved in neuron fate specification, contributes to this reduction in regional differences. We further demonstrate that a genetically defined subtype of ASD, chromosome 15q11.2-13.1 duplication syndrome (dup15q), shares the core transcriptomic signature observed in idiopathic ASD. Co-expression network analysis reveals that individuals with ASD show age-related changes in the trajectory of microglial and synaptic function over the first two decades, and suggests that genetic risk for ASD may influence changes in regional cortical gene expression. Our findings illustrate how diverse genetic perturbations can lead to phenotypic convergence at multiple biological levels in a complex neuropsychiatric disorder.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
11 July 2018
In this Letter, the labels for splicing events A3SS and A5SS were swapped in column D of Supplementary Table 3a and b. Supplementary Table 3 has been corrected online.
References
Gaugler, T. et al. Most genetic risk for autism resides with common variation. Nat. Genet. 46, 881–885 (2014)
Gratten, J., Visscher, P. M., Mowry, B. J. & Wray, N. R. Interpreting the role of de novo protein-coding mutations in neuropsychiatric disease. Nat. Genet. 45, 234–238 (2013)
de la Torre-Ubieta, L., Won, H., Stein, J. L. & Geschwind, D. H. Advancing the understanding of autism disease mechanisms through genetics. Nat. Med. 22, 345–361 (2016)
Gupta, S. et al. Transcriptome analysis reveals dysregulation of innate immune response genes and neuronal activity-dependent genes in autism. Nat. Commun. 5, 5748 (2014)
Garbett, K. et al. Immune transcriptome alterations in the temporal cortex of subjects with autism. Neurobiol. Dis. 30, 303–311 (2008)
Purcell, A. E., Jeon, O. H., Zimmerman, A. W., Blue, M. E. & Pevsner, J. Postmortem brain abnormalities of the glutamate neurotransmitter system in autism. Neurology 57, 1618–1628 (2001)
Chow, M. L. et al. Age-dependent brain gene expression and copy number anomalies in autism suggest distinct pathological processes at young versus mature ages. PLoS Genet. 8, e1002592 (2012)
Voineagu, I. et al. Transcriptomic analysis of autistic brain reveals convergent molecular pathology. Nature 474, 380–384 (2011)
Iossifov, I. et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 515, 216–221 (2014)
Parikshak, N. N. et al. Integrative functional genomic analyses implicate specific molecular pathways and circuits in autism. Cell 155, 1008–1021 (2013)
Necsulea, A. et al. The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature 505, 635–640 (2014)
Jaffe, A. E. et al. Developmental regulation of human cortex transcription and its clinical relevance at single base resolution. Nat. Neurosci. 18, 154–161 (2015)
Weyn-Vanhentenryck, S. M. et al. HITS-CLIP and integrative modeling define the Rbfox splicing-regulatory network linked to brain development and autism. Cell Reports 6, 1139–1152 (2014)
Irimia, M. et al. A highly conserved program of neuronal microexons is misregulated in autistic brains. Cell 159, 1511–1523 (2014)
Wu, J., Anczuków, O., Krainer, A. R., Zhang, M. Q. & Zhang, C. OLego: fast and sensitive mapping of spliced mRNA-Seq reads using small seeds. Nucleic Acids Res . 41, 5149–5163 (2013)
Li, Y. I. et al. RNA splicing is a primary link between genetic variation and disease. Science 352, 600–604 (2016)
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014)
Lovci, M. T. et al. Rbfox proteins regulate alternative mRNA splicing through evolutionarily conserved RNA bridges. Nat. Struct. Mol. Biol. 20, 1434–1442 (2013)
Raj, B. et al. A global regulatory mechanism for activating an exon network required for neurogenesis. Mol. Cell 56, 90–103 (2014)
Gueroussov, S. et al. An alternative splicing event amplifies evolutionary differences between vertebrates. Science 349, 868–873 (2015)
Maze, I. et al. Critical role of histone turnover in neuronal transcription and plasticity. Neuron 87, 77–94 (2015)
Mullins, C., Fishell, G. & Tsien, R. W. Unifying views of autism spectrum disorders: a consideration of autoregulatory feedback loops. Neuron 89, 1131–1156 (2016)
Kwan, K. Y. et al. SOX5 postmitotically regulates migration, postmigratory differentiation, and projections of subplate and deep-layer neocortical neurons. Proc. Natl Acad. Sci. USA 105, 16021–16026 (2008)
Lamb, A. N. et al. Haploinsufficiency of SOX5 at 12p12.1 is associated with developmental delays with prominent language delay, behavior problems, and mild dysmorphic features. Hum. Mutat. 33, 728–740 (2012)
Willsey, A. J. et al. Coexpression networks implicate human midfetal deep cortical projection neurons in the pathogenesis of autism. Cell 155, 997–1007 (2013)
Sanders, S. J. et al. Insights into autism spectrum disorder genomic architecture and biology from 71 risk loci. Neuron 87, 1215–1233 (2015)
Blumenthal, I. et al. Transcriptional consequences of 16p11.2 deletion and duplication in mouse cortex and multiplex autism families. Am. J. Hum. Genet. 94, 870–883 (2014)
Huttenlocher, P. R. Morphometric study of human cerebral cortex development. Neuropsychologia 28, 517–527 (1990)
Khundrakpam, B. S., Lewis, J. D., Zhao, L., Chouinard-Decorte, F. & Evans, A. C. Brain connectivity in normally developing children and adolescents. Neuroimage 134, 192–203 (2016)
Zhang, Y. E., Landback, P., Vibranovski, M. D. & Long, M. Accelerated recruitment of new brain development genes into the human genome. PLoS Biol. 9, e1001179 (2011)
Harrow, J. et al. GENCODE: producing a reference annotation for ENCODE. Genome Biol . 7 (Suppl. 1), 1–9 (2006)
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat. Biotechnol. 31, 46–53 (2013)
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015)
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. 57, 289–300 (1995)
Shen, S. et al. MATS: a Bayesian framework for flexible detection of differential alternative splicing from RNA-Seq data. Nucleic Acids Res . 40, e61 (2012)
Scoles, H. A., Urraca, N., Chadwick, S. W., Reiter, L. T. & Lasalle, J. M. Increased copy number for methylated maternal 15q duplications leads to changes in gene and protein expression in human cortical samples. Mol. Autism 2, 19 (2011)
Zhang, B. & Horvath, S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol . 4, 17 (2005)
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008)
Langfelder, P. & Horvath, S. Eigengene networks for studying the relationships between co-expression modules. BMC Syst. Biol. 1, 54 (2007)
Zambon, A. C. et al. GO-Elite: a flexible solution for pathway and ontology over-representation. Bioinformatics 28, 2209–2210 (2012)
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015)
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015)
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protocols 7, 562–578 (2012)
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014)
Leek, J. T. & Storey, J. D. Capturing heterogeneity in gene expression studies by surrogate variable analysis. PLoS Genet. 3, 1724–1735 (2007)
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014)
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010)
Dittmar, K. A. et al. Genome-wide determination of a broad ESRP-regulated posttranscriptional network by high-throughput sequencing. Mol. Cell. Biol. 32, 1468–1482 (2012)
Kang, H. J. et al. Spatio-temporal transcriptome of the human brain. Nature 478, 483–489 (2011)
Sunkin, S. M. et al. Allen Brain Atlas: an integrated spatio-temporal portal for exploring the central nervous system. Nucleic Acids Res . 41, D996–D1008 (2013)
Langfelder, P., Zhang, B. & Horvath, S. Defining clusters from a hierarchical cluster tree: the Dynamic Tree Cut package for R. Bioinformatics 24, 719–720 (2008)
Langfelder, P. & Horvath, S. Fast R functions for robust correlations and hierarchical clustering. J. Stat. Softw. 46, i11 (2012)
Winden, K. D. et al. The organization of the transcriptional network in specific neuronal classes. Mol. Syst. Biol. 5, 291 (2009)
Robinson, E. B. et al. Genetic risk for autism spectrum disorders and neuropsychiatric variation in the general population. Nat. Genet. 48, 552–555 (2016)
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014)
Liu, J. Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015)
Morris, A. P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012)
Global Lipids Genetics Consortium Discovery and refinement of loci associated with lipid levels. Nat. Genet. 45, 1274–1283 (2013)
Acknowledgements
Tissue, biological specimens or data used in this research were obtained from the Autism BrainNet (formerly the Autism Tissue Program), which is sponsored by the Simons Foundation, and the University of Maryland Brain and Tissue Bank, which is a component of the NIH NeuroBioBank. We are grateful to the patients and families who participate in the tissue donation programs. The authors acknowledge R. Zielke, J. Cottrell and R. Johnson, who assisted with sample acquisition from the latter brain bank. Funding for this work was provided by grants to D.H.G. (NIMH 5R37 MH060233, 5R01 MH09714 and 5R01 MH100027), N.N.P. (NRSA F30 MH099886, UCLA Medical Scientist Training Program), V.L. (Sigrid Juselius Fellowship) and T.G.B. (training grant 5T32 MH073526). Additional grants supporting this work include those to B.J.B. (CIHR, Alzheimer’s Research Foundation and University of Toronto McLaughlin Centre) and M.I. (ERC-StG-LS2-637591). We also thank D. Polioudakis for assistance with data management and V. Chandran for discussion of transcription factor binding site analysis and providing software.
Author information
Authors and Affiliations
Contributions
N.N.P. and D.H.G. planned and directed experiments, guided analyses, and wrote the manuscript with assistance from all authors. N.N.P., V.S. and T.G.B. performed dissections, RNA-seq analysis, and differential gene expression analysis. N.N.P. and V.S. performed splicing analysis. M.I. and B.J.B. provided splicing validation data and assisted with splicing analysis. N.N.P., V.S., S.H., G.R., M.J.G. and C.H. performed co-expression network analysis. N.N.P., T.G.B., V.L. and J.K.L. performed analysis of duplication 15q syndrome samples. V.S. performed RT–PCR validation experiments and V.S., L.d.l.T.U. and J.H. performed SOX5 validation experiments.
Corresponding author
Ethics declarations
Competing interests
D.H.G. is a paid consultant to Ovid Therapeutics. The authors declare no other competing financial interests related to this work.
Additional information
Reviewer Information
Nature thanks K. Mirnics and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Methodology, quality control, and differential expression replication analysis.
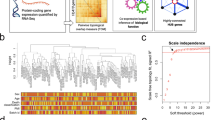
a, RNA-seq workflow (see Supplementary Information for details). b, RNA-seq quality and alignment statistics from this study, including RNA integrity number (RIN), sequencing depth (aligned reads), proportion of reads mapping to different genomic regions, and bias in coverage from the 5′ to the 3′ ends of transcripts. c, RNA-seq read coverage relative to normalized gene length across transcript length across samples. d, Dependence between coverage and RIN across gene body. e–g, Correlation of transcript model quantifications comparing the union exon model (used throughout this study), the whole gene model (which includes introns), and the Cufflinks approach43 to estimating FPKM. h, Summary table describing the characteristics of the matched covariate data used in the DGE and differential alternative splicing (DS) analysis of ASD in cortex and cerebellum. This includes the number of samples overlapping with our previous work8, the age and RIN distributions, and the dependence between diagnosis and age and RIN (summarized from Supplementary Table 1). i, Independent replication of ASD versus control DGE fold changes between previously evaluated and new ASD samples in cortex by RNA-seq using samples from ref. 8 (similar to Fig. 1a, but with RNA-seq in all samples). j, Correlation of P value rankings with Spearman’s correlation across different DGE methods for DGE analysis in cortex, comparing the ‘full model’ (LME P value) described in the Supplementary Information with other methods. Methods include removal of three additional principal components of sequencing surrogate variables(SVs) (LME with 5 SVs, top left), application of a permutation analysis for DGE P value computation (LME P, permuted, top right), application of variance-weighted linear regression for DGE44 (limma voom, middle left), application of surrogate variable analysis for DGE45 (full model + 17 SVs, middle right), and application of DESeq2 with the full model46, which uses a negative binomial distribution (bottom left). k. Comparison of fold changes between frontal cortex (FC) and temporal cortex (TC) for all samples, demonstrating similar changes in both regions. l, Average linkage hierarchical clustering of samples in ASD cortex using the top 100 upregulated and top 100 downregulated protein coding genes, demonstrating that confounders do not drive clustering of about two-thirds of samples. m, The first principal component of the cortex DGE set is primarily associated with diagnosis, and not with other factors. The red line marks a Bonferroni-corrected P = 0.05.
Extended Data Figure 2 Transcriptome-wide DGE analysis.
a, We applied a classification method robust to overfitting (elastic net model47) by training on the RNA-seq data from samples previously analysed in ref. 8 (Extended Data Fig. 1h, similar to the comparison in Extended Data Fig. 1i) and classifying ASD versus control status in independent samples. Results are shown as a comparison of classification scores (left) and area under the receiver operator characteristic curve (AUROC, right). Approximately 85% of ASD samples are classified successfully around a false positive rate of 20%. b, Summary table describing the subset of representative, covariate matched samples used for qRT–PCR validations. Supplementary Table 2 contains the underlying values. c, Fold changes from RNA-seq compared against fold changes from qRT–PCR (see Supplementary Table 2 for data). d, GO term enrichment analysis of genes that are upregulated or downregulated in individuals with ASD. e, Enrichment analysis of cell-type specific gene sets (defined as genes with fivefold higher expression in the cell type than in other cell types) with genes that are decreased or increased in ASD. f, g, Independent replication analysis of ASD versus control DGE fold changes between previously evaluated and new ASD samples from cerebellum by microarray and RNA-seq using samples from ref. 8 (similar to Fig. 1a and Extended Data Fig. 1i). The RNA-seq data show a replication signal between previously evaluated and new samples from this study. h, Comparison of fold changes that were significant at FDR < 0.05 in the ASD versus control DGE analysis from cortex compared with fold changes observed in cerebellum, revealing strong concordance but a lower average fold change in the cerebellum. i, Sample summary and quality control (QC) statistics for ref. 4. Compare to Extended Data Fig. 1b and see Supplementary Information for additional discussion. Compared to this study, samples from ref. 4 were prepared by poly(A) selection RNA-seq, exhibit lower RNA integrity number (RIN, median 4.8 versus 7.3), have lower median sequencing depth (11 million versus 40 million), exhibit greater 5′-3′ bias, and have generally greater variability across all QC metrics. j, Comparison of fold-changes for the top significant genes from ref. 4 (P < 0.01 as provided in their Supplementary Information) with the fold changes for the same genes in this study. Co-expression network analysis demonstrated that the moderate agreement is largely driven by concordance in upregulation of microglial genes in both studies (Extended Data Fig. 8e). k, Average linkage hierarchical clustering of lncRNAs in the DGE set. l, Boxplots of expression values of DGE lncRNAs across multiple tissue types from the Illumina Body Map (expression data from ref. 12). Lines above the plot indicate pairwise significance with a one-sided Wilcoxon rank-sum test between brain and the other tissues. m, Similar to l, except for embryonic stem cells and stem-cell-derived cell types. n, RT–PCR validation of the two lncRNAs shown in Fig. 1c, d; P values computed by two-sided Wilcoxon rank-sum test.
Extended Data Figure 3 RNA-seq gene expression on genome browser tracks for selected primate-specific lncRNAs in human, macaque and mouse.
For each lncRNA, expression for representative samples for ASD versus control (top) in human, macaque (middle), and mouse (bottom) are shown. The genome location for macaque and mouse displayed is syntenic to the human region, with the expected location of the lncRNA highlighted. a–g, Examples of specific lncRNA transcripts that show primate-specific (in human and macaque, or only in human, but not in mouse) expression. h, Example of a strongly conserved lncRNA, which shows robust expression in all three species.
Extended Data Figure 4 Splicing analyses and validation in ASD.
a, Schematic of the PSI metric used for differential alternative splicing35. b, Distribution of LME model P values for changes in the PSI between ASD and control in cortex for all events and event subtypes. c, Distribution of LME model P values for changes in the PSI between ASD and control in cerebellum. d, Average linkage hierarchical clustering in ASD and control cortex samples using top 100 differentially included and top 100 differentially excluded exons from the differential splicing set. e, The first principal component of the cortex differential splicing set is strongly associated with diagnosis, but not other factors. Red line marks Bonferroni-corrected P = 0.05. f, Comparison of the cortex differential splicing with the pipeline used here (TopHat2 (ref. 43) followed by multivariate analysis of transcript splicing, MATS35) with PSI values obtained via another method (read alignment by OLego followed by PSI quantification with Quantas15). g, Comparison of ΔPSI values between RT–PCR and RNA-seq for nine splicing events (Supplementary Table 3). h, Differential splicing analysis identifies events independent of DGE signal. Top,difference between ASD and control in the differential splicing set based on PC1 of the differential splicing set at the PSI level, and PC1 of the gene expression levels of genes in the differential splicing set. Bottom, same comparison after removing nominally differentially expressed genes (P < 0.05). P values computed by two-sided Wilcoxon rank-sum test. i, GO term enrichment analysis of genes with differential splicing events in ASD. j, Clustering dendrogram and heat map for neuronal splicing factor gene expression levels across samples demonstrating three major clusters and the known positive correlation between SRRM4 and RBFOX1 and anticorrelation between PTBP1 and SRRM4 (refs 14,19).
Extended Data Figure 5 Additional splicing analyses in ASD.
a, PCR validation and sashimi plots for nine splicing events delineated in Extended Data Fig. 4d, from the indicated samples (see Extended Data Fig. 2b for details of these samples). Notably, these genes are not in the DGE set, but are detected in the differential alternative splicing set owing to altered transcript structure. b, Heat map as in Fig. 1h for the splicing regulator ESRP48. ESRP is not known to be involved in neuronal function, ESRP1 is not expressed in cortex, and ESRP2 is expressed but not significantly different between ASD and control cortex. Therefore, we show ESRP enrichment analysis in differential splicing events as a control for Fig. 1h. Enrichment P values are computed as described in Methods.
Extended Data Figure 6 Attenuation of cortical patterning in ASD.
a, Histograms of P values from paired Wilcoxon rank-sum test differential gene expression between 16 frontal cortex (FC) and 16 temporal cortex (TC) samples from control and ASD individuals. b, Histogram of Bartlett’s test P values for differences in gene expression variance between ASD and control samples for all genes (white) and genes in the ACP set (red). The Kolmogorov–Smirnov (K–S) test P value for a difference between these two distributions is shown. c, Histograms of P values from unpaired Wilcoxon rank-sum test DGE between 21 frontal cortex and 22 temporal cortex samples after removing those used in ref. 8. d, Histogram of Bartlett’s test P values for differences in gene expression variance between ASD and control samples for all genes (white) and genes in the ACP set (red). The Kolmogorov–Smirnov test P value for a difference between these two distributions is reported. e, Approach to training the elastic net model on BrainSpan49,50 frontal cortex and temporal cortex samples and application of the model to 123 cortical samples in this study. f–h, Results of learned cortical region classifications with different starting gene sets, with the BrainSpan training set (left), control samples (middle) and ASD samples (right) in each panel and the Wilcoxon rank-sum test P value of frontal versus temporal cortex difference for each comparison. A1C, primary auditory cortex; DFC, dorsolateral prefrontal cortex; MFC, medial prefrontal cortex; STC, superior temporal cortex. i, Cell-type enrichment analysis for genes in the ACP set. j, GO term enrichment analysis of the ACP set. Enrichment P values are computed as described in Methods. k, Enrichment statistics for transcription factor motifs found to be significantly enriched in the ACP set (see Supplementary Information for details of P value computation). l, Average linkage hierarchical clustering of the global gene expression profiles for samples with overexpression of SOX5 and green fluorescent protein (GFP) tag overexpression (controls). m, Density plots of fold changes for the subset of ACP genes that are predicted SOX5 targets (top, green) and non-targets (bottom, green) against background (grey). The median log2[fold change] is marked (red line) and P values are from a one-sided Wilcoxon rank-sum test.
Extended Data Figure 7 Duplication 15q syndrome analyses.
a, Copy number between breakpoints in the 15q region. Genome-wide copy number analysis allowed evaluation of copy number in additional regions from previous studies36. b, Sample characteristics for the dup15q analyses (additional details available in Supplementary Table 1). c, Similar to Fig. 3b, but focusing on the lncRNAs found to be significantly differentially expressed in idiopathic ASD compared to control subjects. d, Comparison of DGE fold changes demonstrating that using different control samples (control samples used in the idiopathic analysis, column 2 of Extended Data Fig. 7b) for the dup15q cortex analysis yields similar findings. e, Similar to d except for the differential alternative splicing analysis. f, Comparison of heterogeneity in the DGE signal using the first principal component of the ASD cortex DGE set across all cortical samples used in DGE analyses. Samples from individuals with diagnoses confirmed by dup15q mutations, confirmed by Autism Diagnostic Interview-Revised (ADI-R), and supported by clinical records are all significantly different from controls by two-sided pairwise Wilcoxon rank sum tests. g, Similar to Fig. 3d, but with the larger set of controls from the idiopathic ASD versus control analysis in Fig. 1. h, i, P value distributions for DGE changes outside the 15q region for cortex and cerebellum. j, Similar to Fig. 3a, but for the cerebellum analysis. k, Comparison of significant DGE changes in the duplicated region from cortex with changes in cerebellum. l, Comparison of significant DGE changes outside of the dup15q region in cortex with changes in cerebellum. Scatter plot P values correspond to the statistical significance of the Pearson correlation coefficient between fold changes (see Methods).
Extended Data Figure 8 Cortex co-expression network analyses.
a, Sample characteristics for the cortex network analyses; additional details available in Supplementary Table 1. b, Average linkage hierarchical clustering using the topological overlap metric for co-expression dissimilarity37. Modules are identified from this dendrogram, which was constructed from a consensus of 100 bootstrapped datasets51,52 (see Methods). Correlations for each gene to covariates are delineated below the dendrogram (blue, negative; red, positive). Modules are labelled with colours and numerical labels (see Supplementary Table 4 for additional details). CTX.M11 is a module of genes that are not co-expressed (grey module) and was not evaluated in further comparisons. c, Module-trait associations as computed by an LME model with all factors on the x axis used as covariates. Technical covariates were removed as part of adjusting the FPKM values. All P values are displayed where the association passed Bonferroni-corrected P < 0.05. d, Module enrichments for cell-type specific gene expression patterns. Asterisks indicate FDR < 0.05 across all comparisons. e, Enrichment of ASD-associated modules with that from ref. 4. *FDR < 0.05 (see Supplementary Table 4 for details).
Extended Data Figure 9 Additional figures for cortex co-expression network analyses.
a, Gene set enrichment analyses comparing the 24 cortex co-expression modules with multiple gene sets from this RNA-seq study, post-mortem ASD cortex microarray8, human cortical development10, the set of all brain-expressed lncRNAs, genes enriched for ASD-associated rare variants26, and genes with de novo variants associated with intellectual disability (ID)9. Boxes are filled if the odds ratio is greater than 0 and the enrichment P < 0.05. *FDR < 0.05 across all comparisons, controlling for gene length and expression level with logistic regression (Supplementary Information). b, Overlap of gene sets between firing-rate and mitochondrial associated modules from ref. 53 with ASD-associated modules in cortex. c–e, Module plot of ASD-associated modules not shown in Fig. 4 (CTX.M4, CTX.M9, CTX.M10) displaying the top hub genes along with the module’s GO term enrichment. f, Temporal trajectories for four module eigengenes (CTX.M4, CTX.M9, CTX.M10, CTX.M16) associated with ASD, similar to Fig. 4g. ASD samples are represented by red points and lines, control samples by black. g, Module plot and GO term enrichment for CTX.M24, which is enriched in ASD-associated rare variants and lncRNAs. h, Common variant enrichment across modules as calculated by GWAS enrichment with LD score regression41,42 (see Methods). Disease GWAS studies evaluated include ASD54, schizophrenia55, inflammatory bowel disease56, type 2 diabetes mellitus57 and serum lipid levels58. P values are FDR corrected across all GWAS studies and modules. i, Plot of the proportion of SNP heritability across diseases for ASD-associated modules. Error bars represent s.e.
Extended Data Figure 10 Cerebellum co-expression network analyses.
a, Sample characteristics for the cerebellum network analyses; additional details available in Supplementary Table 1. b, Modules identified from a dendrogram constructed from a consensus of 100 bootstrapped networks (see Methods). Correlations for each gene to each measured factor are delineated below the dendrogram (blue, negative; red, positive). Modules are labelled alphabetically instead of numerically to distinguish them from the cortex modules. Additional information is available in Supplementary Table 4. c, Signed association of module eigengenes with diagnosis; positive values indicate modules with increased expression in ASD samples. Grey bars with labels signify three ASD-associated modules. d, Cell-type enrichments for the three ASD-associated modules. e, Gene set enrichment analyses comparing the three ASD-associated cerebellum modules with post-mortem ASD cortex microarray, human brain development, six cortex ASD-associated modules from this RNA-seq study, and firing rate and mitochondrial associated modules from ref. 53. Boxes are filled if the odds ratio is greater than 0 and the enrichment P < 0.05. *FDR < 0.05 across all comparisons. f–h, Module plots of CB.ML, CB.MP, and CB.MT displaying the top hub genes along with the GO term enrichment. Additional details, including module preservation statistics for cerebellum in cortex and vice versa, are available in Supplementary Table 4.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, full legends for Supplementary Tables 1-4 and Supplementary References. (PDF 2016 kb)
Supplementary Table 1
Metadata for samples used in the study. (XLSX 147 kb)
Supplementary Table 2
Differential gene expression changes in cortex and cerebellum, cortical patterning results, and co-expression network module assignments. (XLSX 22921 kb)
Supplementary Table 3
Differential splicing changes in cortex and cerebellum. (XLSX 16396 kb)
Supplementary Table 4
Module preservation analyses and Gene Ontology term enrichment analyses. (XLSX 113 kb)
Rights and permissions
About this article
Cite this article
Parikshak, N., Swarup, V., Belgard, T. et al. Genome-wide changes in lncRNA, splicing, and regional gene expression patterns in autism. Nature 540, 423–427 (2016). https://doi.org/10.1038/nature20612
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20612
This article is cited by
-
Remodeling of the postsynaptic proteome in male mice and marmosets during synapse development
Nature Communications (2024)
-
Metabolic shift toward ketosis in asocial cavefish increases social-like affinity
BMC Biology (2023)
-
Exploratory analysis of L1 retrotransposons expression in autism
Molecular Autism (2023)
-
Epigenomic signatures reveal mechanistic clues and predictive markers for autism spectrum disorder
Molecular Psychiatry (2023)
-
Mapping the cortico-striatal transcriptome in attention deficit hyperactivity disorder
Molecular Psychiatry (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.