Abstract
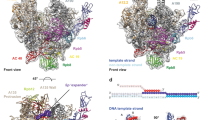
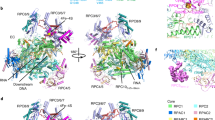
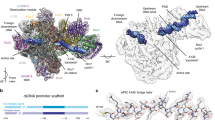
RNA polymerase I (Pol I) is a highly processive enzyme that transcribes ribosomal DNA (rDNA) and regulates growth of eukaryotic cells1,2,3,4. Crystal structures of free Pol I from the yeast Saccharomyces cerevisiae have revealed dimers of the enzyme stabilized by a ‘connector’ element and an expanded cleft containing the active centre in an inactive conformation5,6,7. The central bridge helix was unfolded and a Pol-I-specific ‘expander’ element occupied the DNA-template-binding site. The structure of Pol I in its active transcribing conformation has yet to be determined, whereas structures of Pol II and Pol III have been solved with bound DNA template and RNA transcript8,9,10. Here we report structures of active transcribing Pol I from yeast solved by two different cryo-electron microscopy approaches. A single-particle structure at 3.8 Å resolution reveals a contracted active centre cleft with bound DNA and RNA, and a narrowed pore beneath the active site that no longer holds the RNA-cleavage-stimulating domain of subunit A12.2. A structure at 29 Å resolution that was determined from cryo-electron tomograms of Pol I enzymes transcribing cellular rDNA confirms contraction of the cleft and reveals that incoming and exiting rDNA enclose an angle of around 150°. The structures suggest a model for the regulation of transcription elongation in which contracted and expanded polymerase conformations are associated with active and inactive states, respectively.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Roeder, R. G. & Rutter, W. J. Multiple forms of DNA-dependent RNA polymerase in eukaryotic organisms. Nature 224, 234–237 (1969)
Novello, F. & Stirpe, F. Simultaneous assay of RNA polymerase I and II in nuclei isolated from resting and growing rat liver with the use of alpha-amanitin. FEBS Lett . 8, 57–60 (1970)
Grummt, I. The nucleolus—guardian of cellular homeostasis and genome integrity. Chromosoma 122, 487–497 (2013)
Goodfellow, S. J. & Zomerdijk, J. C. Basic mechanisms in RNA polymerase I transcription of the ribosomal RNA genes. Subcell. Biochem . 61, 211–236 (2013)
Engel, C., Sainsbury, S., Cheung, A. C., Kostrewa, D. & Cramer, P. RNA polymerase I structure and transcription regulation. Nature 502, 650–655 (2013)
Fernández-Tornero, C. et al. Crystal structure of the 14-subunit RNA polymerase I. Nature 502, 644–649 (2013)
Kostrewa, D., Kuhn, C. D., Engel, C. & Cramer, P. An alternative RNA polymerase I structure reveals a dimer hinge. Acta Crystallogr. D Biol. Crystallogr . 71, 1850–1855 (2015)
Gnatt, A. L., Cramer, P., Fu, J., Bushnell, D. A. & Kornberg, R. D. Structural basis of transcription: an RNA polymerase II elongation complex at 3.3 A resolution. Science 292, 1876–1882 (2001)
Kettenberger, H., Armache, K. J. & Cramer, P. Complete RNA polymerase II elongation complex structure and its interactions with NTP and TFIIS. Mol. Cell 16, 955–965 (2004)
Hoffmann, N. A. et al. Molecular structures of unbound and transcribing RNA polymerase III. Nature 528, 231–236 (2015)
Bernecky, C., Herzog, F., Baumeister, W., Plitzko, J. M. & Cramer, P. Structure of transcribing mammalian RNA polymerase II. Nature 529, 551–554 (2016)
Jennebach, S., Herzog, F., Aebersold, R. & Cramer, P. Crosslinking-MS analysis reveals RNA polymerase I domain architecture and basis of rRNA cleavage. Nucleic Acids Res . 40, 5591–5601 (2012)
Pilsl, M. et al. Structure of the initiation-competent RNA polymerase I and its implication for transcription. Nat. Commun. 7, 12126 (2016)
Cramer, P., Bushnell, D. A. & Kornberg, R. D. Structural basis of transcription: RNA polymerase II at 2.8 angstrom resolution. Science 292, 1863–1876 (2001)
Vassylyev, D. G., Vassylyeva, M. N., Perederina, A., Tahirov, T. H. & Artsimovitch, I. Structural basis for transcription elongation by bacterial RNA polymerase. Nature 448, 157–162 (2007)
Sekine, S., Murayama, Y., Svetlov, V., Nudler, E. & Yokoyama, S. The ratcheted and ratchetable structural states of RNA polymerase underlie multiple transcriptional functions. Mol. Cell 57, 408–421 (2015)
Engel, C., Plitzko, J. & Cramer, P. RNA polymerase I-Rrn3 complex at 4.8 Å resolution. Nat. Commun. 7, 12129 (2016)
Plaschka, C. et al. Transcription initiation complex structures elucidate DNA opening. Nature 533, 353–358 (2016)
Sosunov, V. et al. Unified two-metal mechanism of RNA synthesis and degradation by RNA polymerase. EMBO J . 22, 2234–2244 (2003)
Kuhn, C. D. et al. Functional architecture of RNA polymerase I. Cell 131, 1260–1272 (2007)
Kettenberger, H., Armache, K. J. & Cramer, P. Architecture of the RNA polymerase II-TFIIS complex and implications for mRNA cleavage. Cell 114, 347–357 (2003)
Ruan, W., Lehmann, E., Thomm, M., Kostrewa, D. & Cramer, P. Evolution of two modes of intrinsic RNA polymerase transcript cleavage. J. Biol. Chem. 286, 18701–18707 (2011)
Weixlbaumer, A., Leon, K., Landick, R. & Darst, S. A. Structural basis of transcriptional pausing in bacteria. Cell 152, 431–441 (2013)
Tagami, S. et al. Crystal structure of bacterial RNA polymerase bound with a transcription inhibitor protein. Nature 468, 978–982 (2010)
Miller, O. L., Jr & Beatty, B. R. Visualization of nucleolar genes. Science 164, 955–957 (1969)
Osheim, Y. N. et al. Pre-18S ribosomal RNA is structurally compacted into the SSU processome prior to being cleaved from nascent transcripts in Saccharomyces cerevisiae. Mol. Cell 16, 943–954 (2004)
Albert, B. et al. RNA polymerase I-specific subunits promote polymerase clustering to enhance the rRNA gene transcription cycle. J. Cell Biol. 192, 277–293 (2011)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013)
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Scheres, S. H. A Bayesian view on cryo-EM structure determination. J. Mol. Biol. 415, 406–418 (2012)
Plaschka, C. et al. Architecture of the RNA polymerase II-Mediator core initiation complex. Nature 518, 376–380 (2015)
Cardone, G., Heymann, J. B. & Steven, A. C. One number does not fit all: mapping local variations in resolution in cryo-EM reconstructions. J. Struct. Biol. 184, 226–236 (2013)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Barad, B. A. et al. EMRinger: side chain-directed model and map validation for 3D cryo-electron microscopy. Nat. Methods 12, 943–946 (2015)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Osheim, Y. N., French, S. L., Sikes, M. L. & Beyer, A. L. Electron microscope visualization of RNA transcription and processing in Saccharomyces cerevisiae by Miller chromatin spreading. Methods Mol. Biol . 464, 55–69 (2009)
Machín, F. et al. Transcription of ribosomal genes can cause nondisjunction. J. Cell Biol. 173, 893–903 (2006)
Castaño-Díez, D., Al-Amoudi, A., Glynn, A. M., Seybert, A. & Frangakis, A. S. Fiducial-less alignment of cryo-sections. J. Struct. Biol. 159, 413–423 (2007)
Kunz, M. & Frangakis, A. S. Super-sampling SART with ordered subsets. J. Struct. Biol. 188, 107–115 (2014)
Kunz, M. & Frangakis, A. S. Three-dimensional CTF correction improves the resolution of electron tomograms. J. Struct. Biol. http://dx.doi.org/10.1016/j.jsb.2016.06.016 (2016)
Pruggnaller, S., Mayr, M. & Frangakis, A. S. A visualization and segmentation toolbox for electron microscopy. J. Struct. Biol. 164, 161–165 (2008)
Korinek, A., Beck, F., Baumeister, W., Nickell, S. & Plitzko, J. M. Computer controlled cryo-electron microscopy—TOM2 a software package for high-throughput applications. J. Struct. Biol. 175, 394–405 (2011)
Acknowledgements
We thank O. Gadal and I. Lèger-Silvestre for technical assistance with the yeast Miller tree spreading technique, and H. Schwalbe for initial discussions. We thank T. Gubbey for initial experiments on the Pol I elongation complex and C. Bernecky, C. Plaschka and D. Tegunov for support with the single-particle data analysis. We thank T. Schulz for yeast fermentation. S.N. was supported by a PhD student fellowship from the Boehringer Ingelheim Fonds. P.C. was supported by the Deutsche Forschungsgemeinschaft (SFB860, SPP1935), the Advanced Grant ‘TRANSREGULON’ from the European Research Council (grant agreement No 693023), and the Volkswagen Foundation. A.S.F. was supported by the Deutsche Forschungsgemeinschaft (SFB 902) and the Starting Grant ‘JTOMO’ from the European Research Council.
Author information
Authors and Affiliations
Contributions
S.N. planned and carried out the single particle sample preparation, data collection and data analysis. M.K. planned and carried out the tomographic data analysis. C.G. carried out the sample preparation for tomography. M.H. advised on structure determination procedures. V.V.H. advised on and carried out sample preparation for tomography. A.S. advised on sample preparation for tomography. C.E. advised on biochemical procedures. M.P.S. advised on tomographic data analysis. P.C. designed and supervised research, and supervised single particle structure determination. A.S.F. designed and supervised research, supervised single particle data collection and performed tomographic data collection and analysis. S.N., P.C. and A.S.F. prepared the manuscript, with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information: Nature thanks R. Ebright, E. Nogales and E. Nudler for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Preparation of Pol I elongation complex (EC) for single-particle cryo-EM.
a, Size-exclusion chromatogram (Superose 6 Increase 3.2/300; GE Healthcare) of reconstituted Pol I EC. Higher absorbance at 260 nm (red line) than at 280 nm (blue line) indicates presence of nucleic acids. Coomassie-stained SDS–PAGE analysis of pooled peak fractions shows the presence of all 14 Pol I subunits. b, Coomassie-stained SDS–PAGE analysis of titration with BS3 cross-linker. Gel is cropped to large subunits A190 and A135. A shift to higher molecular weight is observed with increasing BS3 concentration indicating successful crosslinking. Based on interpolation, we chose 0.9 mM BS3 (not shown) as the appropriate concentration for final sample preparation.
Extended Data Figure 2 Single-particle cryo-EM particle sorting pipeline.
Annotated arrows indicate the direction of processing and provide information regarding the number of particles used and the classification masks applied. A representative micrograph of the Pol I EC under cryo conditions showed particles of the expected size. A set of 1,500 particles was picked manually with EMAN2 (ref. 30) and used to generate initial 2D classes for template based auto-picking in Relion31. After cleaning by manual inspection and in 2D classification, per frame B-factor weighting and translational movie alignment was applied to the remaining 282,000 particles. The colouring of the surfaces is according to the standard polymerase subunit colouring: A190, grey; A135, wheat; A49, light blue; A43, slate; AC40, red; A34.5, pink; Rpb5, magenta; Rpb6, silver-blue; AC19, yellow; Rpb8, green; A14, hot pink; A12.2, orange; Rpb10, blue; Rpb12, lemon. Template DNA, non-template DNA and RNA are depicted in medium blue, sky blue and red, respectively. The structures against greyed background indicate final EC and Pol I monomer structures.
Extended Data Figure 3 Quality of single-particle cryo-EM reconstructions.
a, Top and bottom view of local resolution surface maps. b, Representative areas of the single-particle cryo-EM density for Pol I EC (left panel) and Pol I monomer (right panel). The A190 helix α19 (upper panel) and the A135 strand β40 (lower panel) are depicted together with the refined model superimposed. c, Angular distribution of particle images. Red dots indicate views with at least one particle assigned within 1°. Black shading represents the number of particles. The orientation occupancy is similar for both structures and covers most of the angles. d, FSC curves. Blue lines indicate the FSC between half maps of the respective reconstruction and red lines indicate FSC between the derived model against the single-particle cryo-EM map.
Extended Data Figure 5 Additional details on Pol I EC.
a, Cleft loops. Ribbon model of ECs of all three S. cerevisiae RNA polymerases superimposed on the bridge helix9,10. Bridge helix (green) and downstream DNA together with DNA-RNA hybrid (blue and red) are given for Pol I. b, Ribbon model of free Pol I (PDB code 4C2M (ref. 5), black and orange) superimposed on the shown inner A190 funnel helix α21 with Pol I EC (grey, green and pink). As a consequence of cleft contraction, parts of the shelf module move in and reduce the width of the pore to impair binding of the C-terminal domain of A12.2. c, Modelling the A12.2 C-terminal domain into the pore of the contracted Pol I EC results in a clash. In the upper part, a surface representation of domains in free Pol I shows that the C-terminal domain of A12.2 fills the pore that is lined by the A190 funnel helix α21 and loop 1572-1579 of the A190 cleft domain in the shelf module. In the lower part, cleft contraction observed in the EC reduces the width of the pore, causing a steric clash in the model.
Extended Data Figure 6 Free monomeric Pol I single-particle cryo-EM structure.
a, Ribbon model of free, monomeric Pol I solved by single-particle cryo-EM. The views correspond to the ‘front’ and ‘top’ views with the incoming downstream DNA pointing towards the reader. Colour code as in Extended Data Fig. 2. b, Free Pol I (PDB 4C2M, black), Pol I Monomer (pink) and Pol I EC (orange) were superimposed onto A135. For clarity, only subunits A190 and A135 are shown. c, Scheme of the observed conformations as displayed in b. The cleft width was measured at two positions: (i) between residue K434 from chain A135 to residue E414 from chain A190 and (ii) between residue K1331 from chain A190 to residue G231 from chain A190. The distance bars above and below the polymerase cartoon indicate the distances between the protrusion and the clamp core helices (above) and at the entry site of downstream DNA (below). The difference (Δ) between the free Pol I (ref. 5) and the Pol I monomer, the Pol-I–Rrn3 (ref. 17) and Pol I EC, respectively, is given in brackets. For the measurement of the relative movement of the clamp core helices shown in the magnified inset, all Pol I structures were aligned on the A135 subunit and the distance between residue E414 of subunit A190 of the free Pol I5 to the same residue of the Pol I EC, Pol I – Rrn317 and Pol I monomer, respectively, was measured. d, Electron density of the bridge helix in the free Pol I Monomer. e, Comparison of bridge helices in the free Pol I monomer (orange) and free Pol I (black) from the crystal structure. f, Electron density (semi-transparent grey) is shown together with models for the bridge helix, trigger loop (both grey) and the C-terminal domain of A12.2. The expander (red) is not present in this structure but modelled here based on the crystal structure of the free Pol I dimer, revealing a clash. g, Inhibited24 and paused23 bacterial polymerase superimposed on Rpb1 of the free Pol I monomer. For clarity only A190 and A135 of Pol I are shown and the β′-NCD of the bacterial polymerases is excluded from the visualization.
Extended Data Figure 7 Yeast cells, lysed to leak their nucleoplasm, prepared with negative stain and visualized under cryo conditions.
a, Electron micrograph of a negatively stained lysed yeast cell, with the nucleoplasm spread on the carbon support film. The upper left of the micrograph is occupied by the grid bar. The yeast cell has released the nuclear context on the grid, which appears as an electron-lucent leakage. b, Electron micrograph of the spread nucleoplasm of a plunge-frozen yeast cell at close-to-native conditions. In the lower left corner, the remains of a yeast cell can be seen as an electron-dense patch. The nucleoplasm is embedded in an ice layer and the asterisks indicate three Miller trees found in the vicinity of this cell. The Miller tree indicated with the red asterisk was used for recording of the tilt-series in Fig. 3.
Extended Data Figure 8 Relative positions of polymerases towards each other and of protruding nucleic acids.
a, Histogram of centre–centre distance d of two consecutive Pols as depicted in (Fig. 3c). The reason for measuring distances smaller than 12 nm is that the enzyme is not completely spherical. Thus at certain rotational arrangements the centre-to-centre distance can be smaller than the average diameter of Pol I. b, Histogram of in-plane angle φ spanned by three consecutive Pols as depicted in Fig. 3c. c, Focused sub-tomogram averaging around the RNA. The RNA exits Pol I as an ~10 Å thick density, both in the slice and in the isosurface representation. d, Sub-tomogram average with the alignment focused on the downstream DNA. The downstream DNA is a long, straight 2 nm density, both in the slice and in the isosurface representation. In both c and d, the Pol I molecule is a globular ~12 nm featureless density. e, Stereo pair of the sub-tomogram average shows the positions of the nascent RNA chain as green balls. The positions that were manually identified by three independent users, without previous knowledge of the positions of the sub-tomogram average, correspond closely to the position of the RNA exit site that was postulated by the X-ray crystallography structure5.
Extended Data Figure 9 Comparisons between cryo-electron tomography and single-particle cryo-EM structures.
a, FSC of the cryo-electron tomography structure with a resolution of 29 Å (purple line) and FSC between the cryo-electron tomography structure and single-particle cryo-EM structure with estimated resolutions of 44 Å and 31 Å, measured at FSC 0.5 and 0.143 criteria, respectively (green line). b, Poor fit of the expanded, free Pol I crystal structure5 (PDB: 4C2M) to the cryo-electron tomography density (grey). In the expanded state a significant part of the clamp core helices are outside the cryo-electron tomography density (26% outside, 74% inside), while in the contracted state they almost completely enclosed (4% outside, 96% inside). In addition, in the expanded state 54% of the cryo-electron tomography density remains unoccupied compared to 19% of the contracted state (also compare Fig. 3d, reproduced here for comparison purposes).
Supplementary information
Conformational changes between free Pol I and Pol I EC
The video illustrates the transition from free monomeric Pol I to elongating Pol I. The four major polymerase modules core, jaw-lobe, clamp and shelf are colored in grey, blue, yellow and pink, respectively. The movement of the rigid clamp-shelf module is highlighted together with the refolding of the bridge helix (green). The adjacent dimer of model of the crystal structure (PDB 4C2M) is shown briefly before the focus is set on the domain movements. Furthermore, the expander (light green), downstream DNA (blue), the DNA-RNA hybrid (blue-red) and the C-terminal domain of A12.2 are visualized. (MP4 10596 kb)
Rights and permissions
About this article
Cite this article
Neyer, S., Kunz, M., Geiss, C. et al. Structure of RNA polymerase I transcribing ribosomal DNA genes. Nature 540, 607–610 (2016). https://doi.org/10.1038/nature20561
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20561
This article is cited by
-
Structural insights into nuclear transcription by eukaryotic DNA-dependent RNA polymerases
Nature Reviews Molecular Cell Biology (2022)
-
Conserved strategies of RNA polymerase I hibernation and activation
Nature Communications (2021)
-
In situ cryo-electron tomography reveals gradient organization of ribosome biogenesis in intact nucleoli
Nature Communications (2021)
-
Organization and regulation of gene transcription
Nature (2019)
-
Ribosome assembly coming into focus
Nature Reviews Molecular Cell Biology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.