Abstract
Innate lymphoid cells (ILCs) functionally resemble T lymphocytes in cytotoxicity and cytokine production but lack antigen-specific receptors, and they are important regulators of immune responses and tissue homeostasis1,2. ILCs are generated from common lymphoid progenitors, which are subsequently committed to innate lymphoid lineages in the α-lymphoid progenitor, early innate lymphoid progenitor, common helper innate lymphoid progenitor and innate lymphoid cell progenitor compartments3,4,5,6,7,8. ILCs consist of conventional natural killer cells and helper-like cells (ILC1, ILC2 and ILC3)9. Despite recent advances1,2,10, the cellular heterogeneity, developmental trajectory and signalling dependence of ILC progenitors are not fully understood. Here, using single-cell RNA-sequencing (scRNA-seq) of mouse bone marrow progenitors, we reveal ILC precursor subsets, delineate distinct ILC development stages and pathways, and report that high expression of programmed death 1 (PD-1hi) marked a committed ILC progenitor that was essentially identical to an innate lymphoid cell progenitor. Our data defined PD-1hiIL-25Rhi as an early checkpoint in ILC2 development, which was abolished by deficiency in the zinc-finger protein Bcl11b but restored by IL-25R overexpression. Similar to T lymphocytes, PD-1 was upregulated on activated ILCs. Administration of a PD-1 antibody depleted PD-1hi ILCs and reduced cytokine levels in an influenza infection model in mice, and blocked papain-induced acute lung inflammation. These results provide a perspective for exploring PD-1 and its ligand (PD-L1) in immunotherapy, and allow effective manipulation of the immune system for disease prevention and therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others

References
Eberl, G., Colonna, M., Di Santo, J. P. & McKenzie, A. N. Innate lymphoid cells. Innate lymphoid cells: a new paradigm in immunology. Science 348, aaa6566 (2015)
Artis, D. & Spits, H. The biology of innate lymphoid cells. Nature 517, 293–301 (2015)
Chea, S. et al. Single-cell gene expression analyses reveal heterogeneous responsiveness of fetal innate lymphoid progenitors to Notch signaling. Cell Reports 14, 1500–1516 (2016)
Constantinides, M. G., McDonald, B. D., Verhoef, P. A. & Bendelac, A. A committed precursor to innate lymphoid cells. Nature 508, 397–401 (2014)
Klose, C. S. et al. Differentiation of type 1 ILCs from a common progenitor to all helper-like innate lymphoid cell lineages. Cell 157, 340–356 (2014)
Yang, Q. et al. TCF-1 upregulation identifies early innate lymphoid progenitors in the bone marrow. Nat. Immunol. 16, 1044–1050 (2015)
Yu, X. et al. The basic leucine zipper transcription factor NFIL3 directs the development of a common innate lymphoid cell precursor. eLife 3, (2014)
Ishizuka, I. E. et al. Single-cell analysis defines the divergence between the innate lymphoid cell lineage and lymphoid tissue-inducer cell lineage. Nat. Immunol. 17, 269–276 (2016)
Spits, H. et al. Innate lymphoid cells—a proposal for uniform nomenclature. Nat. Rev. Immunol. 13, 145–149 (2013)
Serafini, N., Vosshenrich, C. A. & Di Santo, J. P. Transcriptional regulation of innate lymphoid cell fate. Nat. Rev. Immunol. 15, 415–428 (2015)
Brennecke, P. et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods 10, 1093–1095 (2013)
Yu, Y. et al. Bcl11a is essential for lymphoid development and negatively regulates p53. J. Exp. Med. 209, 2467–2483 (2012)
Xu, W. et al. NFIL3 orchestrates the emergence of common helper innate lymphoid cell precursors. Cell Reports 10, 2043–2054 (2015)
Seillet, C. et al. Nfil3 is required for the development of all innate lymphoid cell subsets. J. Exp. Med. 211, 1733–1740 (2014)
Seehus, C. R. et al. The development of innate lymphoid cells requires TOX-dependent generation of a common innate lymphoid cell progenitor. Nat. Immunol. 16, 599–608 (2015)
Zook, E. C. et al. The ETS1 transcription factor is required for the development and cytokine-induced expansion of ILC2. J. Exp. Med. 213, 687–696 (2016)
Weber, B. N. et al. A critical role for TCF-1 in T-lineage specification and differentiation. Nature 476, 63–68 (2011)
McConnell, M. J. et al. Growth suppression by acute promyelocytic leukemia-associated protein PLZF is mediated by repression of c-myc expression. Mol. Cell. Biol. 23, 9375–9388 (2003)
Hoyler, T. et al. The transcription factor GATA-3 controls cell fate and maintenance of type 2 innate lymphoid cells. Immunity 37, 634–648 (2012)
Pardoll, D. M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 12, 252–264 (2012)
Yagi, R. et al. The transcription factor GATA3 is critical for the development of all IL-7Rα-expressing innate lymphoid cells. Immunity 40, 378–388 (2014)
Yu, Y. et al. The transcription factor Bcl11b is specifically expressed in group 2 innate lymphoid cells and is essential for their development. J. Exp. Med. 212, 865–874 (2015)
Walker, J. A. et al. Bcl11b is essential for group 2 innate lymphoid cell development. J. Exp. Med. 212, 875–882 (2015)
Huang, Y. et al. IL-25-responsive, lineage-negative KLRG1hi cells are multipotential ‘inflammatory’ type 2 innate lymphoid cells. Nat. Immunol. 16, 161–169 (2015)
Sharma, P. & Allison, J. P. Immune checkpoint targeting in cancer therapy: toward combination strategies with curative potential. Cell 161, 205–214 (2015)
Agata, Y. et al. Expression of the PD-1 antigen on the surface of stimulated mouse T and B lymphocytes. Int. Immunol. 8, 765–772 (1996)
Barlow, J. L. et al. Innate IL-13-producing nuocytes arise during allergic lung inflammation and contribute to airways hyperreactivity. J. Allergy Clin. Immunol. 129, 191–198 (2012)
Kasagi, S. et al. Anti-programmed cell death 1 antibody reduces CD4+PD-1+ T cells and relieves the lupus-like nephritis of NZB/W F1 mice. J. Immunol. 184, 2337–2347 (2010)
Novey, H. S., Marchioli, L. E., Sokol, W. N. & Wells, I. D. Papain-induced asthma—physiological and immunological features. J. Allergy Clin. Immunol. 63, 98–103 (1979)
Halim, T. Y., Krauss, R. H., Sun, A. C. & Takei, F. Lung natural helper cells are a critical source of Th2 cell-type cytokines in protease allergen-induced airway inflammation. Immunity 36, 451–463 (2012)
Jackson, J. T. et al. Id2 expression delineates differential checkpoints in the genetic program of CD8α+ and CD103+ dendritic cell lineages. EMBO J. 30, 2690–2704 (2011)
Serafini, N. et al. Gata3 drives development of RORγt+ group 3 innate lymphoid cells. J. Exp. Med. 211, 199–208 (2014)
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protocols 9, 171–181 (2014)
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013)
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015)
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014)
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 4, 1–48 (2008)
Robinette, M. L. et al. Transcriptional programs define molecular characteristics of innate lymphoid cell classes and subsets. Nat. Immunol. 16, 306–317 (2015)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Acknowledgements
We thank F. Colucci and J. Di Santo for providing Rag2−/−Il2rg−/− mice. We thank the Sanger Institute RSF (J. Bussell, D. Key, A. Kirton, L. Bulman, S. Kemp, P. Green, P. Zielezinski, R. Lacey, C. Rogerson, A. Logan and G. Notley), Flow Cytometry Core Facility (B. L. Ng, J. Graham and C. Hall), Single Cell Genomic Core Facility (S. Loren and I. Bronner) and DNA sequencing pipeline (N. Smerdon) for technical assistances. We thank K. Chen, J. Pramanik and R. Miragaia for technical help. C.W. is supported by the Plan of Youth Growth from Shanghai Municipal Agricultural Committee (Hunongqingzi (2015. No. A-35)). L.L. is funded by National Natural Science Foundation of China (31370904, 81671579). G.T.B. is supported by the Australian Research Council (Future Fellowship FT110100283) and the National Health and Medical Research Council (Fellowship 10402092). A.N.J.M. is supported by the Medical Research Council (U105178805) and Wellcome Trust (100963/Z/13/Z). This work is supported by Wellcome Trust (grant number 098051) (P.L).
Author information
Authors and Affiliations
Contributions
Y.Y. designed research, performed experiments and analysed data. J.C.H.T. performed all of the bioinformatics analyses, C.W. performed experiments. S.C. and C.B. performed influenza infection. J.W. generated the Bcl11btdTomato conditional knockout reporter mice. X.C. performed ChIP–PCR. L.K. did the histologic section staining. L.S.C. analysed the histologic data. L.L. contributed intellectually to the PD-1 experiments. G.T.B. and A.N.J.M. provided Id2GFP and Il13+/tdTomato reporter mice, respectively. S.A.T. and G.D. provided intellectual input for the experiments performed in their laboratories. Y.Y., J.C.H.T. and P. L. wrote the paper. Y. Y. and P. L. conceived the PD-1 as an ILC marker concept. P.L. supervised the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information
Nature thanks I. Amit and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Quality control of scRNA-seq data of ILC-progenitor-enriched bone marrow cells.
a, FACS sorting strategies of the adult bone marrow cells from wild-type or Vav-Cre-Bcl11bfl/fl mice. Flt3lo or IL-7Rαlo cells were included to detect more ILC progenitors. Lin−Flt3lo/−IL-7Rαlo/+α4β7+ cells were further divided into three populations (CD244+CD25−, CD244−CD25− and CD244−CD25+). We sorted two 96-well plates of CD244+CD25−, one plate of CD244−CD25− and one plate of ILC2 progenitors CD244−CD25+ to include most ILC progenitors for scRNA-seq. Two 96-well plates of Lin−Flt3−IL-7Rα+α4β7+ bone marrow cells from Vav-Cre-Bcl11bfl/fl mice were purified to investigate early ILC2 development defects. Lin: CD19, CD3, CD4, CD5, CD8, TCRβ, TCRγδ, NK1.1, CD11b, Gr-1, CD11c and Ter119. b, Column charts show the fraction of cells passing specific quality control criteria in each plate: unique count mapped to annotated genes >500,000 (top panel); count mapped to mitochondrial-encoded genes <10% (middle panel); and number of annotated gene detected >2,500 (bottom panel). c, Percentage of cells passing all criteria. d, The percentages of ERCC RNA spike-ins in each plate. The black dots represent the mean of the dataset. e, The total number of unique counts mapped to annotated genes in different plates. The black dots represent the mean of the dataset. f, The fractions of the 92 external ERCC RNA spike-ins in different plates. g, Kolmogorov–Smirnov test of individual ERCC spike-ins between the two plates did not detect any ERCC spike-ins showing significantly different (log2 fold change > 1, and adjusted P value < 0.05) levels. The two vertical lines mark the log2 fold change levels of −1 and 1, the horizontal line marks the adjusted P value threshold of 0.05. h, Identification of highly variable genes. Brown points represent annotated mouse genes. Blue points represent external ERCC RNA spike-ins. The magenta points represent the mouse genes that show significantly higher variability (false discovery rate < 0.1). The solid line represents the fit of the technical noise, the dashed line represents the 50% biological CV (coefficient of variation). i, Biaxial t-SNE clustering of the sequenced wild-type cells. j, Column chart comparing the percentage of cells which show detectable mRNA expression of lineage markers in the wild-type bone marrow cells.
Extended Data Figure 2 t-SNE plots showing expression and distribution of genes described in the manuscript.
The colour key shows the expression level.
Extended Data Figure 3 Violin plots of genes highly enriched in specific clusters.
The y axis indicates the log2 (normalized count + 1) expression levels. The black point indicates the mean of expression level. The x axis indicates different clusters.
Extended Data Figure 5 PD-1hi expression marks ILC progenitors.
a, Correlation of expression levels of Pdcd1 and Zbtb16 in C6 cells. Correlation was calculated by Pearson’s method. The fit represents the line of linear regression. b, Violin plot showing the selected gene expression in PD-1hi cells of C6. The y axis indicates the log2 (normalized count + 1) expression levels. The black point indicates the mean of expression level. c, Expression of ILC markers in PD-1hi cells was analysed by FACS. d, The in vivo developmental potential of PD-1hi cells. CD45.1 Rag2−/−Il2rg−/− recipients were injected with the equal numbers of CD45.1−CD45.2+ PD-1hi cells and CD45.1+CD45.2+ (F1 of CD45.1 and CD45.2 parents) common lymphoid progenitors (200–800 cells). The progenies of these donor cells were analysed by FACS after 5–7 weeks (n = 3 per donor cell type). e, Clonal analysis of PD-1hi cells in vitro. The PD-1hi, PD-1hiBcl11b+ and PD-1hiBcl11b− bone marrow cells were FACS-purified and cultured on stromal cells and analysed by FACS. ILC1 was defined as CD45+NK1.1+Bcl11b−, ILC2 as CD45+NK1.1−Bcl11b+ and ILC3 as CD45+NK1.1−Bcl11b−RORγt+. Data are representatives of two (c) or three (d) independent experiments.
Extended Data Figure 6 Direct comparison of PD-1hi and PLZFhi ILC progenitors and dissection of the heterogeneity in the ILC progenitor compartments.
a, FACS plots show PLZFhi and PD-1hi cells had the same development potential in vivo. The equal numbers of PD-1hi cells and PLZFhi cells were adoptively transferred into the same recipient and analysed 3–4 weeks later (n = 3 per donor cell type). b, Schematic diagram of the Bcl11btdTomato conditional knockout reporter allele, where the loxP-IRES-tdTomato cassette was inserted to the 3′UTR of the Bcl11b locus. The other loxP site was in intron 3. Cre-loxP recombination would delete the exon 4. The selection cassette for initial gene targeting was excised by Flpase-FRT recombination. c, Expression of Bcl11b in CHILPs were analysed in the Id2GFP;Bcl11btdTomato duel reporter mice (n = 6). d, Expression of Bcl11b in PD-1hi bone marrow cells was analysed by FACS (n = 6). e, FACS analysis of the in vivo developmental potential of PD-1hiBcl11b− cells (n = 3 per donor cell type). Common lymphoid progenitors were used as the donor cell control. PD-1hiBcl11b− cells predominantly generated ILC1, ILC2 and ILC3. Data are representatives of two (a, e) or three (c, d) independent experiments.
Extended Data Figure 7 scRNA-seq analysis of PD-1hi bone marrow cells.
a, t-SNE clustering analysis of sequenced PD-1hi cells detected two subpopulations. b, Heat map showing the hierarchical clustering result of PD-1hi cells based on selected ILC regulators. The expression levels are log2 transformed and ERCC-size factor normalized.
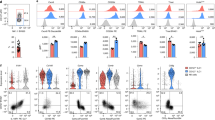
Extended Data Figure 8 scRNA-seq dissection of early ILC2 development.
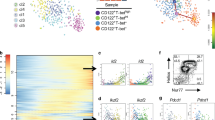
a, Analysis of scRNA-seq data identified genes showing expression changes in C6, C7a, C8 and C9 cells. Change and distribution of expression of selected genes are shown. Il17rb and Bcl11b are among the genes showing spike expression from C6 to C9, whereas Il1rl1 (IL-33R) showed steadily increased expression. The bottom t-SNE plots showing expression of representative genes. b, The expression of IL-25R in PD-1hi bone marrow cells in the Bcl11btdTomato mice (n = 3). c, Clonal differentiation assay of PD-1hiIL-25R+ and PD-1hiIL-25R− cells. Cells were cultured on OP9-DL1 stromal cells in the presence of IL-7 (20.0 ng ml−1) and SCF (50.0 ng ml−1) and were analysed 10 days later. ILC1 was defined as CD45+NK1.1+Bcl11b−, ILC2 as CD45+NK1.1−Bcl11b+ and ILC3 as CD45+NK1.1−Bcl11b−RORγt+. d, FACS analysis of the in vivo developmental potential of PD-1hiIL-25R+ cells. CD45.1 Rag2−/−Il2rg−/− recipients were injected with equal numbers of CD45.1−CD45.2+ PD-1hiIL-25R+ cells and CD45.1+CD45.2+ common lymphoid progenitors (100–500 of each) and the progenies of these populations were analysed 5–7 weeks later (n = 5 per group). e, Analysis of ILCs in Vav-Cre-Bcl11bfl/fl mice. Lin−IL-33R+IL-7Rα+ ILC2s, Lin−KLRG1+IL-7Rαlo ILC2 or Lin–NK1.1+NKp46+ ILCs from the bone marrow, lung or siLP were analysed (n = 4 per genotype), respectively. f, ‘Natural’ ILC2 (nILC2), ‘inflammatory’ ILC2 (iILC2) and BALF IL-5 were analysed in Vav-Cre-Bcl11bfl/fl and the control mice after administration of IL-25 (200 ng per mouse per day) for 3 consecutive days (n = 5 per treated group). Error bars denote s.e.m. g, FACS analysis of in vivo developmental potential of PD-1hi cells from Vav-Cre-Bcl11bfl/fl mice. CD45.1 Rag2−/−Il2rg−/− mice were injected with CD45.2 PD-1hi cells sorted from Bcl11bfl/fl or Vav-Cre-Bcl11bfl/fl mice. The progenies of these donor cells were analysed 4–7 weeks later by FACS (n = 3 per genotype). Data are representatives of three (b) or two (d–g) independent experiments. *P < 0.05, **P < 0.01 (two-tailed t-test).
Extended Data Figure 9 Restoration of development of Bcl11b-deficient PD-1hi ILC progenitors to ILC2 by overexpressing IL-25R.
a, FACS plots showing the expression of TCF-1 and Gata3 in mutant PD-1hi bone marrow cells. Protein expression was measured by intracellular antibody staining. b, Expression patterns of Tox, Id2, Tcf7 and Gata3 in the sequenced Bcl11b-deficient bone marrow cells. c, Overexpressing Il17rb in Bcl11b-deficient PD-1hi bone marrow cells. The rescued cells were analysed by FACS for ILC2 surface markers. PD-1hi cells sorted from Vav-Cre-Bcl11bfl/fl mice were transduced with the Il17rb or control retrovirus. The infected cells were cultured on OP9-DL1 stromal cells with the helper CD45.1 ILC2 progenitors in the presence of IL-25 (20.0 ng ml−1), IL-7 (20.0 ng ml−1) and SCF (50.0 ng ml−1). The cells were collected and analysed after two weeks of culture. Data are representatives of two (a, c) independent experiments.
Extended Data Figure 10 PD-1hi marks effector ILCs.
a, FACS analysis of PD-1 expression on peripheral ILCs in steady-state mice (n = 3). b, Gating strategies of lung ILCs. Lung cNK cells were gated as Lin−Id2+IL-7Rα−NK1.1+NKp46+; lung ILC1s as Lin−Id2+IL-7Rα+NK1.1+Bcl11b−; lung ILC2s as Lin−Id2+IL-7Rα+NK1.1−Bcl11b+; and lung ILC3s as Lin−Id2+IL-7Rα+NK1.1−Bcl11b−. The data were from influenza-infected mice at 5 days after infection. cNKs count for at least half of the Lin− leukocytes in these mice (n = 3). c, FACS analysis of PD-1 expression on CD3+ T cells, CD19+ B cells and peripheral ILCs after J43 treatment (n = 3). The tissues were collected at day 14 after J43 treatment. d, FACS plot shows the recognition of different epitopes of PD-1 by PD-1 antibody clones RMP1-30 and J43. The majority of lung PD-1hi ILC2s were stained with both RMP1-30 and J43. e, FACS analysis of lung PD-1hi cNK, ILC1 and ILC3 at 7 days after infection (n = 3). f, BALF cytokines were quantitated as shown (n = 3 per group per time point). The four experimental groups were: Rag1−/− mice with mock infection; Rag1−/− mice infected with A/X-31 and treated with either an antibody isotype control or J43; Rag2−/−Il2rg−/− mice infected with A/X-31 as the ILC-deficient control. g, More PD-1hi cells were found after papain challenge (day 6) in Rag1−/− mice (n = 3). h, Rag1−/− mice were pretreated with PD-1 antibody J43 or the isotype control antibody for 3 days and then administrated with papain (intranasally) for 5 consecutive days. The lung tissue was collected at day 6 for analysis (n = 5 per treatment). Lung ILC2s were reduced in J43 treated mice. PD-1hi and IL-5-producing ILC2 were undetectable after J43 administration. Data are representatives of two (a–h) independent experiments. Error bars (f, h) denote s.e.m. *P < 0.05, **P < 0.01 (two-tailed t-test).
Supplementary information
Supplementary Table 1
This file contains clustering information of bone marrow progenitors. (XLSX 38 kb)
Supplementary Table 2
This file contains clustering information of PD-1hi cells. (XLSX 17 kb)
Supplementary Table 3
This file contains differential gene analysis of WT C6 vs C8. (XLSX 2880 kb)
Supplementary Table 4
This file contains ILC gene set in GESA (XLSX 21 kb)
Rights and permissions
About this article
Cite this article
Yu, Y., Tsang, J., Wang, C. et al. Single-cell RNA-seq identifies a PD-1hi ILC progenitor and defines its development pathway. Nature 539, 102–106 (2016). https://doi.org/10.1038/nature20105
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20105
This article is cited by
-
Combination of IL-33 with PD-1 blockade augment mILC2s-mediated anti-tumor immunity
Cancer Immunology, Immunotherapy (2024)
-
Recent advancements in the B7/CD28 immune checkpoint families: new biology and clinical therapeutic strategies
Cellular & Molecular Immunology (2023)
-
PD-1 receptor outside the main paradigm: tumour-intrinsic role and clinical implications for checkpoint blockade
British Journal of Cancer (2023)
-
Tracing immune cells around biomaterials with spatial anchors during large-scale wound regeneration
Nature Communications (2023)
-
Versatile roles of innate lymphoid cells at the mucosal barrier: from homeostasis to pathological inflammation
Experimental & Molecular Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.