Abstract
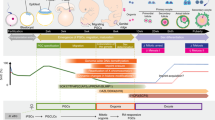
The female germ line undergoes a unique sequence of differentiation processes that confers totipotency to the egg1,2. The reconstitution of these events in vitro using pluripotent stem cells is a key achievement in reproductive biology and regenerative medicine. Here we report successful reconstitution in vitro of the entire process of oogenesis from mouse pluripotent stem cells. Fully potent mature oocytes were generated in culture from embryonic stem cells and from induced pluripotent stem cells derived from both embryonic fibroblasts and adult tail tip fibroblasts. Moreover, pluripotent stem cell lines were re-derived from the eggs that were generated in vitro, thereby reconstituting the full female germline cycle in a dish. This culture system will provide a platform for elucidating the molecular mechanisms underlying totipotency and the production of oocytes of other mammalian species in culture.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Surani, M. A., Hayashi, K. & Hajkova, P. Genetic and epigenetic regulators of pluripotency. Cell 128, 747–762 (2007)
Edson, M. A., Nagaraja, A. K. & Matzuk, M. M. The mammalian ovary from genesis to revelation. Endocr. Rev. 30, 624–712 (2009)
Hübner, K. et al. Derivation of oocytes from mouse embryonic stem cells. Science 300, 1251–1256 (2003)
Qing, T. et al. Induction of oocyte-like cells from mouse embryonic stem cells by co-culture with ovarian granulosa cells. Differentiation 75, 902–911 (2007)
Salvador, L. M., Silva, C. P., Kostetskii, I., Radice, G. L. & Strauss, J. F., III . The promoter of the oocyte-specific gene, Gdf9, is active in population of cultured mouse embryonic stem cells with an oocyte-like phenotype. Methods 45, 172–181 (2008)
Ohinata, Y. et al. Blimp1 is a critical determinant of the germ cell lineage in mice. Nature 436, 207–213 (2005)
McLaren, A. & Southee, D. Entry of mouse embryonic germ cells into meiosis. Dev. Biol. 187, 107–113 (1997)
Hayashi, K. et al. Offspring from oocytes derived from in vitro primordial germ cell-like cells in mice. Science 338, 971–975 (2012)
Morohaku, K. et al. Complete in vitro generation of fertile oocytes from mouse primordial germ cells. Proc. Natl Acad. Sci. USA 113, 9021–9026 (2016)
Ohinata, Y., Sano, M., Shigeta, M., Yamanaka, K. & Saitou, M. A comprehensive, non-invasive visualization of primordial germ cell development in mice by the Prdm1-mVenus and Dppa3-ECFP double transgenic reporter. Reproduction 136, 503–514 (2008)
Hayashi, K., Ohta, H., Kurimoto, K., Aramaki, S. & Saitou, M. Reconstitution of the mouse germ cell specification pathway in culture by pluripotent stem cells. Cell 146, 519–532 (2011)
McClellan, K. A., Gosden, R. & Taketo, T. Continuous loss of oocytes throughout meiotic prophase in the normal mouse ovary. Dev. Biol. 258, 334–348 (2003)
Lucifero, D., Mann, M. R., Bartolomei, M. S. & Trasler, J. M. Gene-specific timing and epigenetic memory in oocyte imprinting. Hum. Mol. Genet. 13, 839–849 (2004)
Peaston, A. E. et al. Retrotransposons regulate host genes in mouse oocytes and preimplantation embryos. Dev. Cell 7, 597–606 (2004)
Veselovska, L. et al. Deep sequencing and de novo assembly of the mouse oocyte transcriptome define the contribution of transcription to the DNA methylation landscape. Genome Biol. 16, 209 (2015)
Tanaka, S. et al. Placentomegaly in cloned mouse concepti caused by expansion of the spongiotrophoblast layer. Biol. Reprod. 65, 1813–1821 (2001)
Ying, Q. L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008)
Irie, N. et al. SOX17 is a critical specifier of human primordial germ cell fate. Cell 160, 253–268 (2015)
Sasaki, K. et al. Robust in vitro induction of human germ cell fate from pluripotent stem cells. Cell Stem Cell 17, 178–194 (2015)
Hayashi, K. & Saitou, M. Generation of eggs from mouse embryonic stem cells and induced pluripotent stem cells. Nat. Protocols 8, 1513–1524 (2013)
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006)
Nagamatsu, G. et al. A germ cell-specific gene, Prmt5, works in somatic cell reprogramming. J. Biol. Chem. 286, 10641–10648 (2011)
Hamazaki, N., Uesaka, M., Nakashima, K., Agata, K. & Imamura, T. Gene activation-associated long noncoding RNAs function in mouse preimplantation development. Development 142, 910–920 (2015)
Patel, R. K. & Jain, M. NGS QC Toolkit: a toolkit for quality control of next generation sequencing data. PLoS One 7, e30619 (2012)
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013)
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protocols 7, 562–578 (2012)
Gaidatzis, D., Lerch, A., Hahne, F. & Stadler, M. B. QuasR: quantification and annotation of short reads in R. Bioinformatics 31, 1130–1132 (2015)
Li, X. et al. A maternal-zygotic effect gene, Zfp57, maintains both maternal and paternal imprints. Dev. Cell 15, 547–557 (2008)
Lee, J. et al. Erasing genomic imprinting memory in mouse clone embryos produced from day 11.5 primordial germ cells. Development 129, 1807–1817 (2002)
Kumaki, Y., Oda, M. & Okano, M. QUMA: quantification tool for methylation analysis. Nucleic Acids Res. 36, W170–W175 (2008)
Takada, Y. et al. HP1γ links histone methylation marks to meiotic synapsis in mice. Development 138, 4207–4217 (2011)
Hodges, C. A. & Hunt, P. A. Simultaneous analysis of chromosomes and chromosome-associated proteins in mammalian oocytes and embryos. Chromosoma 111, 165–169 (2002)
Acknowledgements
We thank Y. Takada for technical support, H. Ohta for technical advice on the iPSCs and embryo transfer, Y. Ohkawa for technical assistance on the RNA-seq, K. Kitajima and C. Meno for providing microscopes, F. Arai for providing FACSAriaII, and H. Leitch for proofreading the manuscript. We also thank the Research Support Center, Research Center for Human Disease Modeling, Kyushu University Graduate School of Medical Sciences for technical assistance. N.H. was a JSPS Research Fellow. This study was supported in part by a Grant-in-Aid from the Ministry of Education, Culture, Sports, Science, and Technology of Japan (KAKENHI #25114006); by JST-PRESTO; by the Uehara Memorial Foundation; and by the Takeda Science Foundation.
Author information
Authors and Affiliations
Contributions
O.H., N.H., S.S. and K.H. performed the culture, embryo transfer, immunofluorescence, PCR analysis and chimaera analysis; O.H., N.H., S.S. and K.H. performed the differentiation to mature oocytes and are able to replicate the entire process, independently. N.H., T.I., and K.N. performed the RNA-seq analysis. G.N. generated the iPSC lines. Y.O and Y.H. conducted the assessment of the culture conditions. M.S. contributed to the inception of the study. K.H. designed the experiments and wrote the manuscript.
Corresponding author
Additional information
Reviewer Information Nature thanks D. Egli, S. Mitalipov and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Evaluation of the basal culture conditions.
a, Culture of rOvaries. Representative images of rOvaries at 3 weeks of culture in the medium indicated on the top are shown. Note that a number of SC-positive oocytes were formed with firm structure of rOvary under the combined culture condition. BF, bright field; SC, stella-ECFP. b, Culture of rOvaries with ICI182780, an oestrogen inhibitor. Representative images of rOvaries under the culture condition indicated on the top are shown. The images on the bottom of the figure are high-magnification images of the region in the dashed box. Note that multiple oocyte follicles (white arrowheads) were frequently observed in the culture without ICI182780 but were rarely observed in the culture with the inhibitor. c, Immunofluorescence analysis of the follicle structure. Images are rOvaries at 3 weeks of culture with or without ICI182780. Immunostaining with laminin, a component of the basement membrane, was used to define the follicle structure that encloses oocyte(s). The bar graph on the right shows the percentage of follicles with single or multiple oocytes. The mean values ± s.d. from 3 biological replicates are shown. The total numbers of follicles counted are shown in the box. Scale bars, 100 μm.
Extended Data Figure 2 Meiotic progression and establishment of genomic imprints in culture.
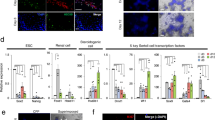
a, PGC-to-oocyte differentiation during IVDi. PGCLCs and oocytes were visualized by anti-GFP antibody (green) cross-reactive to BV and SC. At 21 days of culture, cells surrounding the oocyte expressed Foxl2 (red). Scale bars, 20 μm. b, Representative images of each stage of meiotic prophase I in culture. Meiotic cells from the culture were spread and stained with the antibodies indicated. The colours were converted by ZEN software (Zeiss). c, Meiotic progression in IVDi culture. Percentages of each stage in total meiotic cells are shown. The meiotic cells in the rOvaries were obtained from 3 independent experiments. d, Asynapsis of meiotic chromosome at the pachytene stage in culture. Representative immunofluorescence images of asynapsis at the pachytene stage are shown. Note that γH2AX is densely accumulated in the meiotic chromosome. The graph below the images shows the percentage of asynapsis in cells at the pachytene stages in three E17.5 female gonads, four rOvaries or three E12.5 gonads cultured for 5 days, from 3 independent experiments. Scale bars in b and d, 10 μm.
Extended Data Figure 3 Isolation of individual 2FLs for IVG culture and MII oocytes from IVM culture.
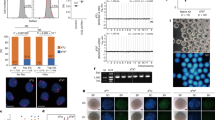
a, Attenuated maturation in an intact rOvary. Cell proliferation in the granulosa cell layer is observed only in 2FLs located at the edge of the rOvary (arrows). 2FLs in the middle seldom show cell proliferation in the granulosa cell layer (arrowheads). b, Representative images of isolation of individual 2FLs. A rOvary (left, upper) was mechanically dissociated (right upper). The high-magnification image of the boxed region shows individual 2FLs after dissociation. c, The number of germinal vesicle and MII oocytes derived from BVSCH18 ESCs. The number of oocytes obtained from each experiment and the total number from all experiments were shown. d, A large image of the MII oocytes shown in Fig. 1d. e, Bisulfite sequence analysis of differentially methylated regions (DMRs) of the imprinted genes (H19 and Igf2r) in MII oocytes generated in vitro and in vivo. White and black circles represent unmethylated and methylated CpG sequences, respectively. Scale bars, 100 μm (a, d), 500 μm (b).
Extended Data Figure 4 Expression of a specific class of LTR.
a, Box plots showing the expression of repetitive elements. Reads per million mapped reads (RPMs) of each product are shown. Note that LTR products increased sharply in oocytes at the secondary follicle stage. b, c, d, Heat map of the normalized expression profile of the LTR family (b), ERVK (c) and ERVL-MaLR (d). e, The gene ontology enrichments of genes categorized by their gene expression dynamics. All genes listed are shown in Supplementary Table 1. All expression data are based on the mean values from 3 biological replicates except for PGCLC(d6) that is from 2 biological replicates.
Extended Data Figure 5 Growth and fertility of offspring from in-vitro-generated oocytes.
a, The number of 2-cell embryos and pups derived from BVSCH18 ESCs. b, Summary of eye colour and transgene of the pups (or adult mouse) derived from in-vitro-generated oocytes from BVSCH18 ESCs. c, Weights of placentas (left), newborn pups (middle) and development of the body weights (right) of offspring from the MII oocytes from BVSCH18 (closed circles) and the genetically matched wild-type mice (129X1/svjC57BL/6F1 × ICR) (closed circles) in 3 independent experiments (t-test, **P < 0.01, *P < 0.05) d, Combined bisulfite restriction analysis (COBRA) of DMRs of the imprinted genes (H19, Igf2r, Peg3 and Snrpn) in the 10 mice (1–10) derived from in-vitro-generated MII oocytes and the two wild-type mice (WT1 and WT2). PCR products were either digested (D) or undigested (U) with the respective enzyme. The digested and undigested fragments are indicated by black and white triangles, respectively. e, Bisulfite sequence analysis of DMRs of the imprinted genes in the two mice (1 and 2) from in-vitro-generated oocytes and the wild-type mouse (WT1). White and black circles represent unmethylated and methylated CpG sequences, respectively. f, A female (left) and a male (right) mouse from in-vitro-generated oocytes with full fertility. The table below the images shows the number of pups in a litter. g, The 6 mice derived from in-vitro-generated MII oocytes at 11 months after birth. For gel source data see Supplementary Fig. 1.
Extended Data Figure 6 Developmental potential of in-vitro-generated oocytes.
a, Percentages of pups from 2-cell embryos transferred. The numbers of pups obtained from 3 independent embryo transfer experiments are shown. b, Fertilization and preimplantation development. Eggs with pronucleus/pronuclei (PN) (left), the number of PN (middle) and early embryonic development in culture (right). The mean values ± s.d. from 5 biological replicates are shown. The total numbers of oocytes/embryos are shown in each graph. c, A uterus at 10 days after transfer of embryos from ESC-derived oocytes. White arrowheads indicate degenerating conceptuses that had decidua without any apparent embryo, and therefore embryo loss at early gestation. The table below the image summarizes the numbers of remaining embryos and decidua without embryo observed at 10 days of gestation. On average, 3.7 embryos remained in the uterus. Scale bar, 5 mm. d, Embryos collected from a day 10 uterus after transfer of embryos from ESC-derived oocytes. The developmental stage of the embryos was varied. Scale bar, 2 mm. e, Implantation marks in the uterus on Caesarean section. White arrowheads indicate the implantation marks without embryos and the asterisk shows the uterus wall after delivering newborn pups by Caesarean section. The table below the image summarizes the numbers of pups and implantation marks without embryos on Caesarean section. On average, 2 pups were obtained per uterus, indicating there was some amount of embryo loss during late gestation.
Extended Data Figure 7 Oogenesis in vitro using other ESC and iPSC lines.
a, Representative images of a rOvary in IVDi, MII oocytes and pups obtained from BVSCH14 ESCs and MEF-derived iPSC lines (MEF4FRC9 and MEF4FC14). b, Weights of placentas (left) and newborns (right) from the MII oocytes from ESC/iPSC lines indicated on the top. The values are from 2 independent experiments (see also Extended Data Table 1). The values of the control are same as those shown in Extended Data Fig. 5c. c, d, Genotyping of the pups from BVSCH14 ESCs (c) and iPSCs (d). PCR products from each gene indicated are shown. P, a positive control; N, a negative control. Details of the positive control and negative control are described in Fig. 3 legend. e, Representative images of a rOvary in IVDi using TTF-derived iPSCs (TTF4FRC3), MII oocytes and pups obtained from TTF4FRC3. Scale bars, 100 μm. For gel source data see Supplementary Fig. 1.
Extended Data Figure 8 Summary of eye colour and transgene of the pups derived from in-vitro-generated oocytes.
a–c, A compiled list of eye colour and transgene of the pups derived from in-vitro-generated oocytes from BVSCH14 ESCs (a), MEF-derived iPSCs (MEF4FRC9 and MEF4FC14) (b), and TTF-derived iPSCs (TTF4FC6 and TTF4FRC3) (c). 3* and 5* pups from TTF4FC6 had albino eyes. This is because MEF-iPSCs and TTF-iPSCs (129X1/svj (chinchilla) × C57BL/6) used in this study have Cc or Ccch genotype that effects eye pigmentation (see also Method). As ICR strain has cc genotype, half of the offspring would have albino eyes. Note that these pups had BV and the retroviral transgenes. d, Fertility of adult mice derived from in-vitro-generated oocytes from iPSCs. Both a female (left) and a male (middle) from TTF-derived iPSCs were fertile. The table (right) shows the number of pups in a litter.
Extended Data Figure 9 Chimaera analysis of rESCs.
a, Bright field (BF) and fluorescent image (BV or SC) of a representative chimaera and wild-type E12.5 embryos (left images), and their gonads with mesonephros (right images). BV expression was detectable in the sensory vibrissae (arrow), the mesenchyme of the forelimb and the hind limb (asterisks) and the myotome (arrowheads), as reported previously10. BV and SC expressions were detectable in PGCs in the gonads (double arrowheads). Scale bars, 1 mm. b, BV transgene in chimaera embryos. PCR products from each tissue (NT, neural tube; H, heart; Lu, lung; Li, liver; I, intestine; T, tail; NC, a negative control from the genomic DNA of a wild-type embryo) are shown. c, MII oocytes from rESCs. The image in Fig. 4d merged with SC is shown. Scale bars, 100 μm. d, A schematic drawing showing the in vitro reconstitution of the entire female germ line. For gel source data see Supplementary Fig. 1.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1 (gel source data) and Supplementary Tables 1-2. (PDF 1038 kb)
Rights and permissions
About this article
Cite this article
Hikabe, O., Hamazaki, N., Nagamatsu, G. et al. Reconstitution in vitro of the entire cycle of the mouse female germ line. Nature 539, 299–303 (2016). https://doi.org/10.1038/nature20104
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20104
This article is cited by
-
Enabling regulatory policy globally will promote realization of the potential of animal biotechnology
CABI Agriculture and Bioscience (2024)
-
Mesenchymal stem cells promote ovarian reconstruction in mice
Stem Cell Research & Therapy (2024)
-
Integration-free induced pluripotent stem cells from three endangered Southeast Asian non-human primate species
Scientific Reports (2024)
-
The impact of induced pluripotent stem cells in animal conservation
Veterinary Research Communications (2024)
-
Divergent Roles of KLF4 During Primordial Germ Cell Fate Induction from Human Embryonic Stem Cells
Reproductive Sciences (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.