Abstract
Three-dimensional physical interactions within chromosomes dynamically regulate gene expression in a tissue-specific manner1,2,3. However, the 3D organization of chromosomes during human brain development and its role in regulating gene networks dysregulated in neurodevelopmental disorders, such as autism or schizophrenia4,5,6, are unknown. Here we generate high-resolution 3D maps of chromatin contacts during human corticogenesis, permitting large-scale annotation of previously uncharacterized regulatory relationships relevant to the evolution of human cognition and disease. Our analyses identify hundreds of genes that physically interact with enhancers gained on the human lineage, many of which are under purifying selection and associated with human cognitive function. We integrate chromatin contacts with non-coding variants identified in schizophrenia genome-wide association studies (GWAS), highlighting multiple candidate schizophrenia risk genes and pathways, including transcription factors involved in neurogenesis, and cholinergic signalling molecules, several of which are supported by independent expression quantitative trait loci and gene expression analyses. Genome editing in human neural progenitors suggests that one of these distal schizophrenia GWAS loci regulates FOXG1 expression, supporting its potential role as a schizophrenia risk gene. This work provides a framework for understanding the effect of non-coding regulatory elements on human brain development and the evolution of cognition, and highlights novel mechanisms underlying neuropsychiatric disorders.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009)
Jin, F. et al. A high-resolution map of the three-dimensional chromatin interactome in human cells. Nature 503, 290–294 (2013)
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014)
Parikshak, N. N. et al. Integrative functional genomic analyses implicate specific molecular pathways and circuits in autism. Cell 155, 1008–1021 (2013)
McCarthy, S. E. et al. De novo mutations in schizophrenia implicate chromatin remodeling and support a genetic overlap with autism and intellectual disability. Mol. Psychiatry 19, 652–658 (2014)
De Rubeis, S. et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515, 209–215 (2014)
Geschwind, D. H. & Rakic, P. Cortical evolution: judge the brain by its cover. Neuron 80, 633–647 (2013)
Reilly, S. K. et al. Evolutionary changes in promoter and enhancer activity during human corticogenesis. Science 347, 1155–1159 (2015)
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012)
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012)
Sanyal, A., Lajoie, B. R., Jain, G. & Dekker, J. The long-range interaction landscape of gene promoters. Nature 489, 109–113 (2012)
Mifsud, B. et al. Mapping long-range promoter contacts in human cells with high-resolution capture Hi-C. Nat. Genet. 47, 598–606 (2015)
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012)
Cookson, W., Liang, L., Abecasis, G., Moffatt, M. & Lathrop, M. Mapping complex disease traits with global gene expression. Nat. Rev. Genet. 10, 184–194 (2009)
Duggal, G., Wang, H. & Kingsford, C. Higher-order chromatin domains link eQTLs with the expression of far-away genes. Nucleic Acids Res. 42, 87–96 (2014)
Ramasamy, A. et al. Genetic variability in the regulation of gene expression in ten regions of the human brain. Nat. Neurosci. 17, 1418–1428 (2014)
Kim, T. K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010)
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014)
Florio, M. et al. Human-specific gene ARHGAP11B promotes basal progenitor amplification and neocortex expansion. Science 347, 1465–1470 (2015)
Bond, J. et al. ASPM is a major determinant of cerebral cortical size. Nat. Genet . 32, 316–320 (2002)
Necsulea, A. et al. The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature 505, 635–640 (2014)
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014)
Gulsuner, S. et al. Spatial and temporal mapping of de novo mutations in schizophrenia to a fetal prefrontal cortical network. Cell 154, 518–529 (2013)
Hormozdiari, F., Kostem, E., Kang, E. Y., Pasaniuc, B. & Eskin, E. Identifying causal variants at loci with multiple signals of association. Genetics 198, 497–508 (2014)
Fromer, M. et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 506, 179–184 (2014)
Network and Pathway Analysis Subgroup of Psychiatric Genomics Consortium. Psychiatric genome-wide association study analyses implicate neuronal, immune and histone pathways. Nat. Neurosci. 18, 199–209 (2015)
Jones, C. K., Byun, N. & Bubser, M. Muscarinic and nicotinic acetylcholine receptor agonists and allosteric modulators for the treatment of schizophrenia. Neuropsychopharmacology 37, 16–42 (2012)
Graham, V., Khudyakov, J., Ellis, P. & Pevny, L. SOX2 functions to maintain neural progenitor identity. Neuron 39, 749–765 (2003)
Roussos, P. et al. A role for noncoding variation in schizophrenia. Cell Reports 9, 1417–1429 (2014)
Kortüm, F. et al. The core FOXG1 syndrome phenotype consists of postnatal microcephaly, severe mental retardation, absent language, dyskinesia, and corpus callosum hypogenesis. J. Med. Genet. 48, 396–406 (2011)
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. http://dx.doi.org/10.1038/nn.4399 (2016)
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015)
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015)
Miller, J. A. et al. Transcriptional landscape of the prenatal human brain. Nature 508, 199–206 (2014)
Miller, J. A., Horvath, S. & Geschwind, D. H. Divergence of human and mouse brain transcriptome highlights Alzheimer disease pathways. Proc. Natl Acad. Sci. USA 107, 12698–12703 (2010)
Ernst, J. & Kellis, M. Large-scale imputation of epigenomic datasets for systematic annotation of diverse human tissues. Nat. Biotechnol. 33, 364–376 (2015)
van de Leemput, J. et al. CORTECON: a temporal transcriptome analysis of in vitro human cerebral cortex development from human embryonic stem cells. Neuron 83, 51–68 (2014)
Yao, P. et al. Coexpression networks identify brain region-specific enhancer RNAs in the human brain. Nat. Neurosci. 18, 1168–1174 (2015)
Forrest, A. R. et al. A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014)
Zhang, B. & Horvath, S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol. 4, e17 (2005)
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010)
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015)
Stein, J. L. et al. A quantitative framework to evaluate modeling of cortical development by neural stem cells. Neuron 83, 69–86 (2014)
Hansen, K. D., Irizarry, R. A. & Wu, Z. Removing technical variability in RNA-seq data using conditional quantile normalization. Biostatistics 13, 204–216 (2012)
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007)
Oldham, M. C., Langfelder, P. & Horvath, S. Network methods for describing sample relationships in genomic datasets: application to Huntington’s disease. BMC Syst. Biol. 6, 63 (2012)
Acknowledgements
This work is a component of the psychENCODE project and was supported by NIH grants to D.H.G. (5R01MH060233; 5R01MH100027; 3U01MH103339; 1R01MH110927; 1R01MH094714), F.H. and E.E. (R01MH101782; R01ES022282; T32MH073526), J.L.S. (K99MH102357), and J.E. (R01ES024995), NSF CAREER Award (#1254200) to J.E., Glenn/AFAR Postdoctoral Fellowship Program (20145357) and Basic Science Research Program through the National Research Foundation of Korea (2013024227) to H.W., CIRM- BSCRC Training Grant (TG2-01169) to L.T.U., NRSA Training Grant to N.N.P. (F30MH099886; UCLA MSTP), NHMRC project grant (APP1062510) and ARC DECRA fellowship (DE140101033) to I.V. The Hi-C library was sequenced by the BSCRC, and fetal tissue was collected from the UCLA Center for Aids Research (CFAR, 5P30 AI028697). Schizophrenia RNA-seq data were generated as part of the CommonMind Consortium (see Methods and Supplementary Information). eQTL data was provided by M. Ryten and A. Ramasamy. We thank S. Feng, Y. Tian, V. Swarup, and P. S. Mischel for helpful discussions and critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
H.W. designed and performed the Hi-C experiments, implemented the analysis pipeline, interpreted results of Hi-C and performed integration with other expression and epigenetic data, and co-wrote the manuscript. L.T.U., C.O., D.L. and J.H. performed sample collection, brain dissection, and genome editing experiments. J.L.S. and N.N.P. contributed to integrative and statistical analysis. M.J.G. analysed the schizophrenia transcriptomic data. G.J.S. and I.V. performed eRNA–mRNA correlation analyses. F.H. and E.E. aided in the implementation of CAVIAR for credible SNP selection and interpretation of the data. C.L. helped establish the initial Hi-C protocol. J.E. aided in the integration and interpretation of epigenetic data, and edited the manuscript. D.H.G. supervised the experimental design and analysis, interpreted results, provided funding, and co-wrote the manuscript.
Corresponding author
Additional information
Reviewer Information Nature thanks D. Goldstein, B. Ren and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Basic characterization of Hi-C libraries.
a, Hi-C library sequencing information. Percentage of double-stranded (DS) reads indicates percentage of DS reads to all reads, and percentage of valid pairs and filtered reads indicates percentage of valid pairs and filtered reads to DS reads. Cis ratio, ratio of cis (intra-chromosomal) reads to the total number of reads. b, Frequency distribution of Hi-C contacts in GZ (left) and CP (right). c, Pearson correlation between replicates at 100 kb resolution is >0.8, demonstrating a high degree of correlation between biological replicates from different individuals. d, Size distribution of TADs in GZ (left) and CP (right). e, f, Size distribution of genomic regions in between TADs that are less than (TAD boundaries, e) and bigger than (unorganized chromosome, f) 400 kb in GZ (left) and CP (right).
Extended Data Figure 2 Inter-chromosome conformation is associated with various genomic features.
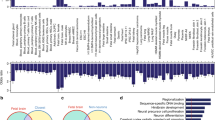
a, Representative heat map of the chromosome contact matrix of CP. Normalized contact frequency (contact enrichment) is colour-coded according to the legend on the right. b, Pearson correlation of the leading principle component (PC1) of inter-chromosomal contacts at 100 kb resolution between in vivo cortical layers and non-neuronal cell types (ES and IMR90 cells). PC1s from neuronal tissues (CP and GZ) have significantly higher correlation than the PC1s between non-neuronal cell types, consistent with the higher similarity between tissues from brain vs the two other cell lines, although batch effects are also likely to contribute. c, Spearman correlation of PC1 of chromatin interaction profile of fetal brain (GZ) with GC content, gene number, DHS of fetal brain, and gene expression level in fetal laminae. d, GO enrichment of genes located in the top 1,000 highly interacting inter-chromosomal regions specific to CP vs GZ (left), and CP vs ES (right), indicating that genes located on dynamic chromosomal regions are enriched for neuronal development.
Extended Data Figure 3 Intra-chromosomal conformation is associated with various genomic features.
a, GO enrichment of genes that change compartment status from A to B (left) and B to A (right) in GZ to CP. b, Heat map of PC1 values of the genome that change compartment status in different cell types. The faction of the genome with a compartment switch in different lineages is described below. c, Distribution of gene expression fold change (FC, left) and DHS FC (right) for genes/regions that change compartment status (‘A to B’ or ‘B to A’) or that remain the same (stable) in different cell/tissue types. B to A compartment shift is associated with increased DHS and gene expression, whereas the A to B shift is associated with decreased DHS and gene expression. P values from one-way ANOVA; whiskers, 1.5 × interquartile range (IQR); centre lines, median (black) and mean (grey). d, Percentage of epigenetic states for genomic regions that change compartment status between ES cells and GZ (left) and ES cells and CP (right). Note that B to A shift in ES cells to GZ/CP is associated with increased proportion of active promoter and transcribed regions (TssA and Tx) and enhancers (Enh, top), while A to B shift in ES cells to GZ/CP is associated with increased proportions of repressive marks (Het and ReprPCWk, bottom). *P < 0.05, **P < 0.01, ***P < 0.001. P values from Fisher’s test. Annotation for epigenetic marks described in a core 15-state model from ref. 33. e, Epigenetic changes in TADs mediate gene expression changes during neuronal differentiation. Genes were divided by expression FC between ES and differentiated neural cells, and epigenetic states in the TADs containing genes in each group were counted and compared between ES cells and CP. Upregulated genes in neurons reside in TADs with more active epigenetic marks in CP than in ES cells, while downregulated genes in neurons reside in TADs with more repressive marks in CP than in ES cells. Epigenetic states associated with activation and transcription of the genes were marked as red bars, while those associated with repression were marked as blue bars on the right. Annotation for epigenetic states described in ref. 33. f, Histone mark enrichment for adult cortical eQTL in fetal brain (FB, left) and adult frontal cortex (FCTX, right). g, Hi-C interaction frequency between eQTL and associated transcripts. LOESS smooth curve plotted with actual data points. Shaded area corresponds to 95% confidence intervals. GZ, chromatin contact frequency in GZ; ES, chromatin contact frequency in ES cells; Exp, expected interaction frequency given the distance between two regions; Opp, opposite interaction frequency: interaction frequency of SNPs and transcripts when the position of genes was mirrored relative to the eQTL. ***P < 0.001, P values from repeated measure of ANOVA.
Extended Data Figure 4 Interacting regions share co-expression and epigenetic states.
a, Top 2% (left), 5% (middle) and 10% (right) highest interacting regions both in GZ and CP (High) show positive correlation with gene expression, while the lowest interacting regions (Low) and variably interacting regions (Variant) have no skew in distribution. P values from Wilcoxon rank-sum test. b, Mean (top) and median (bottom) values for gene expression correlation for high, low and variant interacting regions with different cut-offs, indicating that higher the interaction, higher the correlation of gene expression. c, Top 2% highest interacting regions in fetal brain (FB) show more positive correlation in fetal brain gene expression compared with top 2% highest interacting regions in non-neuronal cells such as ES and IMR90 cells. d, Epigenetic state combination in inter-chromosomal interacting regions in GZ (left) and CP (right). Enhancers (TxEnh5′, TxEnh3′, TxEnhW, EnhA1), transcriptional regulatory regions (TxReg), and transcribed regions (Tx) interact highly with each other as marked in red. e, Epigenetic state combination in intra-chromosomal interacting regions in GZ (left) and CP (right). Enhancers (TxEnh5′, TxEnh3′, TxEnhW, EnhA1) and transcriptional regulatory regions (TxReg) interact highly to promoters (PromD1, PromD2) and transcribed regions (Tx5′, Tx) as marked in red. Inter- and intra-chromosomal contact frequency map is compared to epigenetic state combination matrix by Fisher’s test to calculate the enrichment of shared epigenetic combinations in interacting regions. Coloured bars on the left represent epigenetic marks associated with promoters and transcribed regions (orange), enhancers (red), and repressive marks (blue). Annotation for epigenetic marks is described in a 25-state model from ref. 36.
Extended Data Figure 5 Characterization of chromatin interactome of human-gained enhancers.
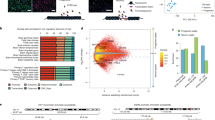
a, Distribution fitting of normalized chromatin interaction frequency between human-gained enhancers and 1 Mb (top) or 100 kb upstream (bottom) regions. The Weibull distribution (red line) fits Hi-C interaction frequency the best for every distance range. b, Distribution of the number of significant interacting loci with human-gained enhancers in GZ (top) and CP (bottom). c, The fraction of epigenetic states for loci interacting with human-gained enhancers in CP and GZ. d, The proportions of human-gained enhancers and interacting regions within the same TAD. e, GO enrichment for human-gained interacting genes in CP (left) and GZ (right).
Extended Data Figure 6 Human-gained enhancers interact with evolutionary lineage-specific lncRNAs.
a, GO enrichment for cell type-specific human-gained enhancer interacting genes. b, GO enrichment for human-gained enhancer interacting genes replicated in more than two individuals from CP (top) and GZ (bottom). Reg., regulation of. c, Protein-coding genes interacting with human-gained enhancers in CP and GZ have lower non-synonymous substitutions (dN) to synonymous substitutions (dS) ratio compared to protein-coding genes that do not interact with human-gained enhancers (All) in mammals (mouse), primates (rhesus macaque), and great apes (chimpanzee), indicative of purifying selection. P values from Wilcoxon rank-sum test. d, Number of lineage-specific lncRNAs interacting with human-gained enhancers (red vertical lines in the graph) in GZ (top) and CP (bottom). Null distribution was generated from 3,000 permutations, where the number of lncRNAs interacting with the same number of enhancers pooled from all fetal brain enhancers was counted.
Extended Data Figure 7 Defining schizophrenia risk genes based on functional annotation of credible SNPs.
a, b, Credible SNPs identified by CAVIAR (a) and defined in the original study (b) are categorized into functional SNPs, SNPs that fall onto gene promoters, and un-annotated SNPs. DHS and histone marks enrichment of credible SNPs was assessed in fetal brain (FB) and adult frontal cortex (FCTX). Functional SNPs and promoter SNPs were directly assigned to the target genes, while un-annotated SNPs were assigned to the target genes via Hi-C interactions in CP and GZ. GO enrichment for genes identified by each category is shown in the bottom. Note that two credible SNP lists overlap with each other; credible SNPs defined in the original study are not restricted to genome-wide significant loci, so they include a broader range (20,362 credible SNPs vs 7,547 CAVIAR SNPs) of SNPs than CAVIAR credible SNPs. NMD, nonsense-mediated decay.
Extended Data Figure 8 Chromatin interactions identify genes that are neither the closest nor in the LD with index SNPs.
a, Number of closest genes and LD genes that interact with credible SNPs (Hi-C identified) vs not (Hi-C non-identified). b, Number of credible SNP interacting genes that are closest to or in LD with index SNPs (Hi-C genes that are also) vs not (Hi-C alone). Hi-C genes here contain only physically interacting genes, but not genes identified by functional SNPs. c, GO enrichment for the closest genes (top) and genes in LD with index SNPs (bottom) that are identified by the schizophrenia risk gene assessment pipeline in Extended Data Fig. 7 (right) vs not (left). d, GO enrichment for schizophrenia risk genes that are neither the closest genes nor in LD to index SNPs. Intersection (left) and union (right) of genes identified by chromatin contacts in CP and GZ are indicated. Venn diagrams are marked in orange to depict the gene list assessed for GO enrichment. e, Representative interaction map of a 10 kb bin, in which credible SNPs reside, to the corresponding 1 Mb flanking regions. Credible SNPs, genomic coordinates for credible SNPs that interact with the target gene; GWAS locus, LD region for the index SNP.
Extended Data Figure 9 Cell-type specificity and reproducibility of Hi-C interactions with schizophrenia GWAS hits.
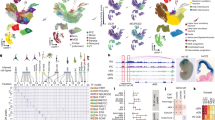
a, GO enrichment for schizophrenia risk genes replicated in more than two individuals in CP (left) and GZ (right). b, Overlap between genes that interact with schizophrenia credible SNPs in CP and GZ vs ES (left) and IMR90 (right) cells. c, GO enrichment for genes that interact with schizophrenia credible SNPs in cell-type specific manner. d, Schematic showing the incorporation of sequence flanking rs1191551 into a reporter (Luc) vector with a minimal promoter (mP). e, PCR amplification of targeted genomic region demonstrates deletion of the SNP-containing region. Expected band size, 587 bp (CRISPR1) and 813 bp (CRISPR2). f, CRISPR/Cas9-mediated deletion of rs1191551 flanking region does not affect the closest protein-coding gene PRKD1 expression. Normalized expression levels of PRKD1 relative to control (Ctrl) (mean ± standard error, n = 6 (Ctrl), 4 (CRISPR1 and CRISPR2)). P values, one-way ANOVA and post hoc Tukey test.
Extended Data Figure 10 High probability schizophrenia risk loci predicted by Hi-C interactions and cortical eQTL.
Hi-C interactions and eQTL association target the same gene (marked in red). Risk alleles lead to target gene dysregulation in the same direction as in schizophrenia brains. Chromosome ideogram and genomic axis (top); gene model based on Gencode v.19 and target genes identified by both Hi-C and eQTL are marked in red; Genomic coordinates for the 10 kb bin containing credible SNPs (schizophrenia GWAS) and eQTL; −log10[P value], P value for the significance of the interaction between schizophrenia credible SNPs and each 10 kb bin, grey dashed line denotes FDR = 0.01; TAD borders in CP and GZ are indicated. Protocadherins (PCDH) gene family is marked as A (PCDHA) and B (PCDHB) except target genes, PCDHA2 and PCDHA7. Whiskers and centre lines correspond to 1.5 × IQR and median, respectively.
Supplementary information
Supplementary Information
This file contains the legends for Supplementary Tables 1-27 (see separate excel file) and Supplementary Notes. (PDF 160 kb)
Supplementary Tables
This file contains Supplementary Tables 1-27. (XLSX 25356 kb)
Rights and permissions
About this article
Cite this article
Won, H., de la Torre-Ubieta, L., Stein, J. et al. Chromosome conformation elucidates regulatory relationships in developing human brain. Nature 538, 523–527 (2016). https://doi.org/10.1038/nature19847
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19847
This article is cited by
-
Cellular reprogramming is driven by widespread rewiring of promoter-enhancer interactions
BMC Biology (2023)
-
Prioritizing genes associated with brain disorders by leveraging enhancer-promoter interactions in diverse neural cells and tissues
Genome Medicine (2023)
-
Regulation of CTCF loop formation during pancreatic cell differentiation
Nature Communications (2023)
-
Annotating genetic variants to target genes using H-MAGMA
Nature Protocols (2023)
-
Integrative analyses highlight functional regulatory variants associated with neuropsychiatric diseases
Nature Genetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.