Abstract
In the dividing eukaryotic cell, the spindle assembly checkpoint (SAC) ensures that each daughter cell inherits an identical set of chromosomes. The SAC coordinates the correct attachment of sister chromatid kinetochores to the mitotic spindle with activation of the anaphase-promoting complex (APC/C), the E3 ubiquitin ligase responsible for initiating chromosome separation. In response to unattached kinetochores, the SAC generates the mitotic checkpoint complex (MCC), which inhibits the APC/C and delays chromosome segregation. By cryo-electron microscopy, here we determine the near-atomic resolution structure of a human APC/C–MCC complex (APC/CMCC). Degron-like sequences of the MCC subunit BubR1 block degron recognition sites on Cdc20, the APC/C coactivator subunit responsible for substrate interactions. BubR1 also obstructs binding of the initiating E2 enzyme UbcH10 to repress APC/C ubiquitination activity. Conformational variability of the complex enables UbcH10 association, and structural analysis shows how the Cdc20 subunit intrinsic to the MCC (Cdc20MCC) is ubiquitinated, a process that results in APC/C reactivation when the SAC is silenced.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Meyer, H. J. & Rape, M. Processive ubiquitin chain formation by the anaphase-promoting complex. Semin. Cell Dev. Biol. 22, 544–550 (2011)
Primorac, I. & Musacchio, A. Panta rhei: the APC/C at steady state. J. Cell Biol. 201, 177–189 (2013)
Lara-Gonzalez, P., Westhorpe, F. G. & Taylor, S. S. The spindle assembly checkpoint. Curr. Biol. 22, R966–R980 (2012)
Musacchio, A. The molecular biology of spindle assembly checkpoint signaling dynamics. Curr. Biol. 25, R1002–R1018 (2015)
Hoyt, M. A., Totis, L. & Roberts, B. T. S. cerevisiae genes required for cell cycle arrest in response to loss of microtubule function. Cell 66, 507–517 (1991)
Li, R. & Murray, A. W. Feedback control of mitosis in budding yeast. Cell 66, 519–531 (1991)
De Antoni, A. et al. The Mad1/Mad2 complex as a template for Mad2 activation in the spindle assembly checkpoint. Curr. Biol. 15, 214–225 (2005)
Mapelli, M., Massimiliano, L., Santaguida, S. & Musacchio, A. The Mad2 conformational dimer: structure and implications for the spindle assembly checkpoint. Cell 131, 730–743 (2007)
Kulukian, A., Han, J. S. & Cleveland, D. W. Unattached kinetochores catalyze production of an anaphase inhibitor that requires a Mad2 template to prime Cdc20 for BubR1 binding. Dev. Cell 16, 105–117 (2009)
Luo, X., Tang, Z., Rizo, J. & Yu, H. The Mad2 spindle checkpoint protein undergoes similar major conformational changes upon binding to either Mad1 or Cdc20. Mol. Cell 9, 59–71 (2002)
Sironi, L. et al. Crystal structure of the tetrameric Mad1-Mad2 core complex: implications of a ‘safety belt’ binding mechanism for the spindle checkpoint. EMBO J. 21, 2496–2506 (2002)
Sudakin, V., Chan, G. K. & Yen, T. J. Checkpoint inhibition of the APC/C in HeLa cells is mediated by a complex of BUBR1, BUB3, CDC20, and MAD2. J. Cell Biol. 154, 925–936 (2001)
Hardwick, K. G., Johnston, R. C., Smith, D. L. & Murray, A. W. MAD3 encodes a novel component of the spindle checkpoint which interacts with Bub3p, Cdc20p, and Mad2p. J. Cell Biol. 148, 871–882 (2000)
Fang, G. Checkpoint protein BubR1 acts synergistically with Mad2 to inhibit anaphase-promoting complex. Mol. Biol. Cell 13, 755–766 (2002)
Davenport, J., Harris, L. D. & Goorha, R. Spindle checkpoint function requires Mad2-dependent Cdc20 binding to the Mad3 homology domain of BubR1. Exp. Cell Res. 312, 1831–1842 (2006)
Burton, J. L. & Solomon, M. J. Mad3p, a pseudosubstrate inhibitor of APCCdc20 in the spindle assembly checkpoint. Genes Dev. 21, 655–667 (2007)
Nilsson, J., Yekezare, M., Minshull, J. & Pines, J. The APC/C maintains the spindle assembly checkpoint by targeting Cdc20 for destruction. Nat. Cell Biol. 10, 1411–1420 (2008)
Chao, W. C., Kulkarni, K., Zhang, Z., Kong, E. H. & Barford, D. Structure of the mitotic checkpoint complex. Nature 484, 208–213 (2012)
Tang, Z., Bharadwaj, R., Li, B. & Yu, H. Mad2-Independent inhibition of APCCdc20 by the mitotic checkpoint protein BubR1. Dev. Cell 1, 227–237 (2001)
Izawa, D. & Pines, J. The mitotic checkpoint complex binds a second CDC20 to inhibit active APC/C. Nature 517, 631–634 (2015)
Chang, L., Zhang, Z., Yang, J., McLaughlin, S. H. & Barford, D. Molecular architecture and mechanism of the anaphase-promoting complex. Nature 513, 388–393 (2014)
Zhang, S. et al. Molecular mechanism of APC/C activation by mitotic phosphorylation. Nature 533, 260–264 (2016)
Reddy, S. K., Rape, M., Margansky, W. A. & Kirschner, M. W. Ubiquitination by the anaphase-promoting complex drives spindle checkpoint inactivation. Nature 446, 921–925 (2007)
Stegmeier, F. et al. Anaphase initiation is regulated by antagonistic ubiquitination and deubiquitination activities. Nature 446, 876–881 (2007)
Pan, J. & Chen, R. H. Spindle checkpoint regulates Cdc20p stability in Saccharomyces cerevisiae. Genes Dev. 18, 1439–1451 (2004)
King, E. M., van der Sar, S. J. & Hardwick, K. G. Mad3 KEN boxes mediate both Cdc20 and Mad3 turnover, and are critical for the spindle checkpoint. PLoS One 2, e342 (2007)
Ge, S., Skaar, J. R. & Pagano, M. APC/C- and Mad2-mediated degradation of Cdc20 during spindle checkpoint activation. Cell Cycle 8, 167–171 (2009)
Foster, S. A. & Morgan, D. O. The APC/C subunit Mnd2/Apc15 promotes Cdc20 autoubiquitination and spindle assembly checkpoint inactivation. Mol. Cell 47, 921–932 (2012)
Uzunova, K. et al. APC15 mediates CDC20 autoubiquitylation by APC/C(MCC) and disassembly of the mitotic checkpoint complex. Nat. Struct. Mol. Biol. 19, 1116–1123 (2012)
Mansfeld, J., Collin, P., Collins, M. O., Choudhary, J. S. & Pines, J. APC15 drives the turnover of MCC-CDC20 to make the spindle assembly checkpoint responsive to kinetochore attachment. Nat. Cell Biol. 13, 1234–1243 (2011)
Herzog, F. et al. Structure of the anaphase-promoting complex/cyclosome interacting with a mitotic checkpoint complex. Science 323, 1477–1481 (2009)
Chang, L., Zhang, Z., Yang, J., McLaughlin, S. H. & Barford, D. Atomic structure of the APC/C and its mechanism of protein ubiquitination. Nature 522, 450–454 (2015)
Malureanu, L. A. et al. BubR1 N terminus acts as a soluble inhibitor of cyclin B degradation by APC/C(Cdc20) in interphase. Dev. Cell 16, 118–131 (2009)
Suijkerbuijk, S. J. et al. The vertebrate mitotic checkpoint protein BUBR1 is an unusual pseudokinase. Dev. Cell 22, 1321–1329 (2012)
Han, J. S. et al. Catalytic assembly of the mitotic checkpoint inhibitor BubR1-Cdc20 by a Mad2-induced functional switch in Cdc20. Mol. Cell 51, 92–104 (2013)
Lara-Gonzalez, P., Scott, M. I., Diez, M., Sen, O. & Taylor, S. S. BubR1 blocks substrate recruitment to the APC/C in a KEN-box-dependent manner. J. Cell Sci. 124, 4332–4345 (2011)
Izawa, D. & Pines, J. How APC/C-Cdc20 changes its substrate specificity in mitosis. Nat. Cell Biol. 13, 223–233 (2011)
He, J. et al. Insights into degron recognition by APC/C coactivators from the structure of an Acm1-Cdh1 complex. Mol. Cell 50, 649–660 (2013)
Elowe, S. et al. Uncoupling of the spindle-checkpoint and chromosome-congression functions of BubR1. J. Cell Sci. 123, 84–94 (2010)
Lischetti, T., Zhang, G., Sedgwick, G. G., Bolanos-Garcia, V. M. & Nilsson, J. The internal Cdc20 binding site in BubR1 facilitates both spindle assembly checkpoint signalling and silencing. Nat. Commun. 5, 5563 (2014)
Di Fiore, B. et al. The ABBA motif binds APC/C activators and is shared by APC/C substrates and regulators. Dev. Cell 32, 358–372 (2015)
Diaz-Martinez, L. A. et al. The Cdc20-binding Phe box of the spindle checkpoint protein BubR1 maintains the mitotic checkpoint complex during mitosis. J. Biol. Chem. 290, 2431–2443 (2015)
Kaisari, S., Sitry-Shevah, D., Miniowitz-Shemtov, S. & Hershko, A. Intermediates in the assembly of mitotic checkpoint complexes and their role in the regulation of the anaphase-promoting complex. Proc. Natl Acad. Sci. USA 113, 966–971 (2016)
Brown, N. G. et al. RING E3 mechanism for ubiquitin ligation to a disordered substrate visualized for human anaphase-promoting complex. Proc. Natl Acad. Sci. USA 112, 5272–5279 (2015)
Varetti, G., Guida, C., Santaguida, S., Chiroli, E. & Musacchio, A. Homeostatic control of mitotic arrest. Mol. Cell 44, 710–720 (2011)
Garnett, M. J. et al. UBE2S elongates ubiquitin chains on APC/C substrates to promote mitotic exit. Nat. Cell Biol. 11, 1363–1369 (2009)
Williamson, A. et al. Identification of a physiological E2 module for the human anaphase-promoting complex. Proc. Natl Acad. Sci. USA 106, 18213–18218 (2009)
Kelly, A., Wickliffe, K. E., Song, L., Fedrigo, I. & Rape, M. Ubiquitin chain elongation requires E3-dependent tracking of the emerging conjugate. Mol. Cell 56, 232–245 (2014)
Reis, A., Levasseur, M., Chang, H. Y., Elliott, D. J. & Jones, K. T. The CRY box: a second APCcdh1-dependent degron in mammalian cdc20. EMBO Rep. 7, 1040–1045 (2006)
Plechanovová, A., Jaffray, E. G., Tatham, M. H., Naismith, J. H. & Hay, R. T. Structure of a RING E3 ligase and ubiquitin-loaded E2 primed for catalysis. Nature 489, 115–120 (2012)
Zhang, Z. et al. Recombinant expression, reconstitution and structure of human anaphase-promoting complex (APC/C). Biochem. J. 449, 365–371 (2013)
Zhang, Z., Yang, J. & Barford, D. Recombinant expression and reconstitution of multiprotein complexes by the USER cloning method in the insect cell-baculovirus expression system. Methods 95, 13–25 (2016)
van den Ent, F. & Löwe, J. RF cloning: a restriction-free method for inserting target genes into plasmids. J. Biochem. Biophys. Methods 67, 67–74 (2006)
Yudkovsky, Y., Shteinberg, M., Listovsky, T., Brandeis, M. & Hershko, A. Phosphorylation of Cdc20/fizzy negatively regulates the mammalian cyclosome/APC in the mitotic checkpoint. Biochem. Biophys. Res. Commun. 271, 299–304 (2000)
Labit, H. et al. Dephosphorylation of Cdc20 is required for its C-box-dependent activation of the APC/C. EMBO J. 31, 3351–3362 (2012)
Kraft, C. et al. Mitotic regulation of the human anaphase-promoting complex by phosphorylation. EMBO J. 22, 6598–6609 (2003)
Hegemann, B. et al. Systematic phosphorylation analysis of human mitotic protein complexes. Sci. Signal. 4, rs12 (2011)
Steen, J. A. et al. Different phosphorylation states of the anaphase promoting complex in response to antimitotic drugs: a quantitative proteomic analysis. Proc. Natl Acad. Sci. USA 105, 6069–6074 (2008)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Bai, X. C., Fernandez, I. S., McMullan, G. & Scheres, S. H. Ribosome structures to near-atomic resolution from thirty thousand cryo-EM particles. eLife 2, e00461 (2013)
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014)
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010)
Tian, W. et al. Structural analysis of human Cdc20 supports multisite degron recognition by APC/C. Proc. Natl Acad. Sci. USA 109, 18419–18424 (2012)
Bolanos-Garcia, V. M. et al. Structure of a Blinkin-BUBR1 complex reveals an interaction crucial for kinetochore-mitotic checkpoint regulation via an unanticipated binding Site. Structure 19, 1691–1700 (2011)
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011)
Fernández, I. S., Bai, X. C., Murshudov, G., Scheres, S. H. & Ramakrishnan, V. Initiation of translation by cricket paralysis virus IRES requires its translocation in the ribosome. Cell 157, 823–831 (2014)
Yang, Z. et al. UCSF Chimera, MODELLER, and IMP: an integrated modeling system. J. Struct. Biol. 179, 269–278 (2012)
Landau, M. et al. ConSurf 2005: the projection of evolutionary conservation scores of residues on protein structures. Nucleic Acids Res. 33, W299–W302 (2005)
Perkins, D. N., Pappin, D. J., Creasy, D. M. & Cottrell, J. S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999)
Keller, A., Nesvizhskii, A. I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002)
Waterhouse, A. M., Procter, J. B., Martin, D. M., Clamp, M. & Barton, G. J. Jalview Version 2–a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009)
Acknowledgements
This work was supported by a Cancer Research UK grant (C576/A14109) and the Medical Research Council (MRC_UP_1201/6) to D.B. and a Long Term EMBO Fellowship to C.A. We thank members of the Barford group for discussions, X. Bai and S. Scheres for their help with RELION; C. Savva and S. Chen for EM facilities; J. Grimmett and T. Darling for computing and W. Zachariae and J. Pines for their invaluable advice and J. Pines for communicating data before publication.
Author information
Authors and Affiliations
Contributions
Z.Z. cloned an initial mutant MCC construct used by C.A. to generate other MCC constructs. C.A. cloned Cdc20, purified proteins, performed the protein complex reconstitutions and biochemical analysis. C.A. and L.C. prepared grids, collected and analysed EM data and determined the 3D reconstructions. C.A. fitted coordinates, built models and made the figures with help of L.C. Z.Z. together with J.Y. generated APC/C viruses. J.Y. provided E1, E2 and ubiquitin for ubiquitination assays. S.M. and M.S. performed mass spectrometry. D.B. directed the project and designed experiments with C.A. C.A. and D.B. wrote the manuscript.
Corresponding author
Additional information
Reviewer Information Nature thanks A. Leschziner and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Biochemical characterization of recombinant APC/CMCC complex and preparations of wild-type and mutant complexes.
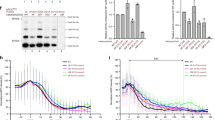
a–e, SDS–PAGE gels (stained with Coomassie) of gel-filtration peak fraction from wild-type and mutant APC/CMCC complex preparations used in this study. Western blot against Strep tag in e confirms the presence of Apc15ΔNTH-Strep construct. f, Top, western blot performed with anti-Apc3 antibody to monitor the time-dependent phosphorylation of this subunit induced by okadaic acid (OA) treatment in the APC/C-expressing insect cells. Bottom, western blot against Apc6 was used as a loading control and reflects the decrease in cell viability after addition of OA (data not shown). g, Western blot against His6-tagged ubiquitin of in vitro securin ubiquitination assays performed with either APC/C or APC/COA in the presence or absence of Cdc20. h, The input sample for the ubiquitination assays performed in this study is shown. i, Western blot against securin of in vitro securin ubiquitination assays performed with APC/COA and Cdc20 with or without either MCC or miniMCC. j, SDS–PAGE of APC/CMCC reconstituted with MBP-TEV-Cdc20APC/C and untagged Cdc20MCC. MBP-TEV-Cdc20APC/C TEV cleavage products are indicated. k, SDS–PAGE gels of reconstituted APC/CΔApc15-MCC complex. l, Western blot against His6-tagged ubiquitin of in vitro securin ubiquitination assays performed with APC/CΔApc15 and Cdc20 with or without MCC. m, Western blot against His6-tagged ubiquitin of in vitro Cdc20 ubiquitination assays performed with APC/CMCC and increasing concentrations of UbcH10. Experiments in g, l and m were replicated three times, and those in i were replicated four times. See Supplementary Fig. 1 for gel source data.
Extended Data Figure 2 Stability of APC/CMCC complex, negative-stain EM reconstructions of APC/CMCC wild-type and mutant complexes and cryo-EM analysis.
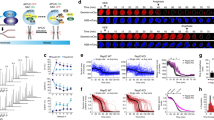
a, Top, chromatogram showing the elution profile of the APC/CMCC complex run on a Superose 6 column. Apo APC/COA and thyroglobulin standard molecular mass marker (669 kDa) are indicated. Bottom, SDS–PAGE of the eluted fractions. APC/CMCC elutes earlier than APC/COA. b, Negative-stain EM reconstructions performed for this study and EMD-1591 (ref. 31) are shown. APC/C (grey) and MCC–Cdc20APC/C module (red) are highlighted. The APC/CMCC and APC/CApc2ΔWHB-MCC reconstructions are also shown in the same orientation as in Fig. 4 to facilitate comparisons. c, A typical cryo-EM micrograph of APC/CMCC-closed representative of 20,234 micrographs. d, Gallery of two-dimensional class averages of APC/CMCC-closed showing different views representative of 50 two-dimensional classes. e, Density quality for secondary structures. The APC/CMCC-closed map was filtered to 4.0 Å.
Extended Data Figure 3 Resolution and other cryo-EM features of APC/CMCC complexes.
a–d, Fourier shell correlation (FSC) curves (a), and local resolution maps calculated with RESMAP63 (b–d) are shown for all the cryo-EM reconstructions determined in this study. b, Ribbon representations of structures shown. c, Overall views of local resolution maps. d, Close up of platform region (Apc4, Apc5 and Apc15). All the maps shown in c and d are filtered to 8.5 Å. Local resolution colour scheme is indicated in the bar at the bottom of d. e, The APC/CUbcH10-MCC reconstruction filtered at 12 Å and shown at different threshold levels. The lowest threshold is the same as in Fig. 5b.
Extended Data Figure 4 Three-dimensional classification of APC/CMCC.
a, b, 3D class averages obtained by classification with local searches (see Methods) are shown in a. Classes 1–5 (56%) show density for the MCC–Cdc20APC/C module with classes 1–3 and 4–5 having a closed-like and an open-like conformation, respectively (framed in red, see Methods). Particles from classes 1–3 and 4–5 were separately refined and re-classified using a mask (yellow, see Methods) shown in b to isolate the best quality particles for APC/CMCC-closed (mask 1: left) and APC/CMCC-open (mask 2: right). Percentages for each of the classes relative to the total number of selected particles are indicated. The percentages relative to the total number of APC/CMCC particles are shown in parentheses.
Extended Data Figure 5 Features of the APC/CMCC structure.
a–c, Cryo-EM density and fitted coordinates for the Cdc20APC/C IR tail (Cdc20APC/C_IR; a), the Cdc20APC/C NTD (Cdc20APC/C_NTD); b) and C-Mad2 (c) are shown. Colours for each subunit are as in Fig. 1. d, Overall superposition of the APC/CCdc20-Hsl1 structure (red) with the APC/CMCC-closed structure (green). The Cdc20WD40 change of position is illustrated, and the blades forming the D-box (yellow) binding-pocket are highlighted. e, Superposition of Apc4, Apc5 and Apc15 between APC/CCdh1-Hsl1-UbcH10-Ub structure (grey) and APC/CMCC (subunit colours as in Fig. 1) shows the marked conformational change of Apc4HBD, Apc5NTD and Apc15NTH induced by Cdc20MCC binding to its indicated binding site on Apc4HBD. f, Close up view of the Cdc20MCC CRY box recognition site of Cdc20APC/C. The CRY box also contacts BubR1 in proximity to D1. Colours for each subunit are as in Fig. 1. g, Superposition of the Apc2WHB domains from APC/CMCC and APC/CCdh1-Hsl1-UbcH10-Ub structures and the corresponding interacting regions of BubR1TPR and UbcH10 are shown. Bottom left, the residues mutated in BubR1Wm that contact Apc2WHB and used in the ubiquitination assay shown in Fig. 5d are indicated (red). Bottom right, residues of UbcH10 (red) that contact the corresponding site on Apc2WHB ablate APC/C UbcH10-dependent ubiquitination activity44.
Extended Data Figure 6 Conservation analysis on BubR1A1-K2 and BubR1TPR regions.
a, Similarities in modes of binding of BubR1 to two Cdc20 subunits of APC/CMCC (left) and Acm1 to two Cdh1 subunits in the Acm1–Cdh1 heterotrimer38 (right). D-box, KEN box, NEN box and ABBA motif are labelled as D, K, NEN and A, respectively. BubR1 (colour-ramped from blue to red indicating N to C terminus) mediates a Cdc20 dimer interface, whereas Acm1 mediates a Cdh1 dimer interface. b, Local sequence alignment performed with BubR1A1-K2 region sequences from several species (described on the left as: sequence identifier_protein name_species/residue number) and the Saccharomyces cerevisiae Acm1A-KEN region. A D-box-like feature (corresponding to Emi1D-box 7–10 positions)32 precedes the first ABBA motif (A1). A 21–33-residue long linker connects the A1 to the second KEN-box (K2). Conserved positions are highlighted in orange. c, ConSurf analysis of the BubR1TPR region highlighting conserved residues on the Cdc20APC/C binding pocket (left) and on the Apc2WHB domain pocket (right). The Cdc20APC/C binding pocket is required for a functional SAC67. This site interacts with residues of BubR1 immediately N-terminal to KEN-2, thereby reinforcing their contacts with Cdc20APC/C. Residue conservation is indicated in a gradient from cyan to purple. BubR1, Cdc20APC/C and Apc2WHB are coloured as in Fig. 1.
Extended Data Figure 7 Three-dimensional classification of APC/CΔApc15-MCC and APC/CUbcH10-MCC.
a, 3D class averages obtained by classification with local searches (see Methods) are shown for APC/CΔApc15-MCC. Particles from classes 1–3 were refined together for obtaining the final APC/CΔApc15-MCC reconstruction shown in b. The APC/CΔApc15-MCC map was filtered to 4.8 Å. c, 3D class averages obtained by classification using a mask (yellow, see Methods) are shown for APC/CUbcH10-MCC. Class 1 was used for the final reconstruction. Percentages relative to the total amount of particles are indicated for each of the classes.
Supplementary information
Supplementary Figure
This file contains original source images for all data obtained by electrophoretic separation: Coomassie stained SDS PAGE and western blots. (PDF 4011 kb)
Structure of APC/CMCC-Closed and MCC-mediated inhibition of degron recognition
The video shows the overall structure of APC/CMCC-Closed, architecture of the MCC-Cdc20APC/C module, the interactions of the BubR1 D box, KEN box and ABBA motif pseudo-degrons with Cdc20APC/C and Cdc20MCC, and how and Cdc20APC/C and Cdc20MCC interact with Apc3 and Apc8. (MOV 28539 kb)
MCC-mediated inhibition of the APC/C catalytic module and conformational changes between 'closed', 'open' and 'UbcH10-bound' APC/CMCC structures.
The video shows the conformational change of the MCC-Cdc20APC/C module that is dependent on Apc15. Apc15 stabilizes APC/CMCC-Open over the APC/CMCC-Closed state that is observed as the major state in wild type APC/CMCC. APC/CMCC-Open resembles the conformation of APC/CMCC with UbcH10. (MOV 7132 kb)
Rights and permissions
About this article
Cite this article
Alfieri, C., Chang, L., Zhang, Z. et al. Molecular basis of APC/C regulation by the spindle assembly checkpoint. Nature 536, 431–436 (2016). https://doi.org/10.1038/nature19083
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19083
This article is cited by
-
PROTAC’ing oncoproteins: targeted protein degradation for cancer therapy
Molecular Cancer (2023)
-
An unexpected timer for cell division
Nature (2023)
-
Principles and dynamics of spindle assembly checkpoint signalling
Nature Reviews Molecular Cell Biology (2023)
-
Lactate regulates cell cycle by remodelling the anaphase promoting complex
Nature (2023)
-
Cell cycle control in cancer
Nature Reviews Molecular Cell Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.