Abstract
Bacteria of the SAR11 clade constitute up to one half of all microbial cells in the oxygen-rich surface ocean. SAR11 bacteria are also abundant in oxygen minimum zones (OMZs), where oxygen falls below detection and anaerobic microbes have vital roles in converting bioavailable nitrogen to N2 gas. Anaerobic metabolism has not yet been observed in SAR11, and it remains unknown how these bacteria contribute to OMZ biogeochemical cycling. Here, genomic analysis of single cells from the world’s largest OMZ revealed previously uncharacterized SAR11 lineages with adaptations for life without oxygen, including genes for respiratory nitrate reductases (Nar). SAR11 nar genes were experimentally verified to encode proteins catalysing the nitrite-producing first step of denitrification and constituted ~40% of OMZ nar transcripts, with transcription peaking in the anoxic zone of maximum nitrate reduction activity. These results link SAR11 to pathways of ocean nitrogen loss, redefining the ecological niche of Earth’s most abundant organismal group.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
BioProject
GenBank/EMBL/DDBJ
Data deposits
SAR11 SAG sequences from the ETNP and GoM can be found in the BioProject database under accession numbers PRJNA290513 and PRJNA291283, respectively. The two OMZ metagenomes sequenced have been deposited in the Joint Genome Institute database under accession numbers 1059848 and 1059863. The mutant E. coli genome sequenced has been deposited in the BioProject database under accession number PRJNA322349. Sequences of the clone SAR11 nar operons have been deposited in GenBank under accession numbers KX275213 and KX275214.
References
Brown, M. V., Schwalbach, M. S., Hewson, I. & Fuhrman, J. A. Coupling 16S-ITS rDNA clone libraries and automated ribosomal intergenic spacer analysis to show marine microbial diversity: development and application to a time series. Environ. Microbiol. 7, 1466–1479 (2005)
Carlson, C. A. et al. Seasonal dynamics of SAR11 populations in the euphotic and mesopelagic zones of the northwestern Sargasso Sea. ISME J. 3, 283–295 (2009)
Eiler, A., Hayakawa, D. H., Church, M. J., Karl, D. M. & Rappé, M. S. Dynamics of the SAR11 bacterioplankton lineage in relation to environmental conditions in the oligotrophic North Pacific subtropical gyre. Environ. Microbiol. 11, 2291–2300 (2009)
Morris, R. M. et al. SAR11 clade dominates ocean surface bacterioplankton communities. Nature 420, 806–810 (2002)
Salter, I. et al. Seasonal dynamics of active SAR11 ecotypes in the oligotrophic Northwest Mediterranean Sea. ISME J. 9, 347–360 (2015)
Thrash, J. C. et al. Single-cell enabled comparative genomics of a deep ocean SAR11 bathytype. ISME J. 8, 1440–1451 (2014)
Giovannoni, S. J. et al. Genome streamlining in a cosmopolitan oceanic bacterium. Science 309, 1242–1245 (2005)
Grote, J. et al. Streamlining and core genome conservation among highly divergent members of the SAR11 clade. MBio 3, e00252–12 (2012)
Tripp, H. J. The unique metabolism of SAR11 aquatic bacteria. J. Microbiol. 51, 147–153 (2013)
Konstantinidis, K. T., Braff, J., Karl, D. M. & DeLong, E. F. Comparative metagenomic analysis of a microbial community residing at a depth of 4,000 meters at station ALOHA in the North Pacific subtropical gyre. Appl. Environ. Microbiol. 75, 5345–5355 (2009)
Swan, B. K. et al. Potential for chemolithoautotrophy among ubiquitous bacteria lineages in the dark ocean. Science 333, 1296–1300 (2011)
King, G. M., Smith, C. B., Tolar, B. & Hollibaugh, J. T. Analysis of composition and structure of coastal to mesopelagic bacterioplankton communities in the northern gulf of Mexico. Front. Microbiol. 3, 438 (2013)
Vergin, K. L. et al. High-resolution SAR11 ecotype dynamics at the Bermuda Atlantic Time-series Study site by phylogenetic placement of pyrosequences. ISME J. 7, 1322–1332 (2013)
Paulmier, A. & Ruiz-Pino, D. Oxygen minimum zones (OMZs) in the modern ocean. Prog. Oceanogr. 80, 113–128 (2009)
Tiano, L., Garcia-Robledo, E. & Revsbech, N. P. A new highly sensitive method to assess respiration rates and kinetics of natural planktonic communities by use of the switchable trace oxygen sensor and reduced oxygen concentrations. PLoS One 9, e105399 (2014)
Kalvelage, T. et al. Nitrogen cycling driven by organic matter export in the South Pacific oxygen minimum zone. Nature Geosci. 6, 228–234 (2013)
Stewart, F. J., Ulloa, O. & DeLong, E. F. Microbial metatranscriptomics in a permanent marine oxygen minimum zone. Environ. Microbiol. 14, 23–40 (2012)
Ganesh, S., Parris, D. J., DeLong, E. F. & Stewart, F. J. Metagenomic analysis of size-fractionated picoplankton in a marine oxygen minimum zone. ISME J. 8, 187–211 (2014)
Ganesh, S. et al. Size-fraction partitioning of community gene transcription and nitrogen metabolism in a marine oxygen minimum zone. ISME J. 9, 2682–2696 (2015)
Ulloa, O., Canfield, D. E., DeLong, E. F., Letelier, R. M. & Stewart, F. J. Microbial oceanography of anoxic oxygen minimum zones. Proc. Natl Acad. Sci. USA 109, 15996–16003 (2012)
Codispoti, L. A. et al. The oceanic fixed nitrogen and nitrous oxide budgets: Moving targets as we enter the anthropocene? Sci. Mar. 65, 85–105 (2001)
Gruber, N. in The Ocean Carbon Cycle and Climate (eds Follows, M. & Oguz, T. ) 97–148 (Springer, 2004)
Stewart, F. J., Sharma, A. K., Bryant, J. A., Eppley, J. M. & DeLong, E. F. Community transcriptomics reveals universal patterns of protein sequence conservation in natural microbial communities. Genome Biol. 12, R26 (2011)
Lüke, C., Speth, D. R., Kox, M. A. R., Villanueva, L. & Jetten, M. S. M. Metagenomic analysis of nitrogen and methane cycling in the Arabian Sea oxygen minimum zone. PeerJ 4, e1924 (2016)
Dalsgaard, T. et al. Oxygen at nanomolar levels reversibly suppresses process rates and gene expression in anammox and denitrification in the oxygen minimum zone off northern Chile. MBio 5, e01966–14 (2014)
Kalvelage, T. et al. Oxygen sensitivity of anammox and coupled N-cycle processes in oxygen minimum zones. PLoS ONE 6, e29299 (2011)
Dean, F. B. et al. Comprehensive human genome amplification using multiple displacement amplification. Proc. Natl Acad. Sci. USA 99, 5261–5266 (2002)
Dupont, C. L. et al. Genomic insights to SAR86, an abundant and uncultivated marine bacterial lineage. ISME J. 6, 1186–1199 (2012)
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015)
Thrash, J. C. et al. Phylogenomic evidence for a common ancestor of mitochondria and the SAR11 clade. Sci. Rep. 1, 13 (2011)
Luo, H. Evolutionary origin of a streamlined marine bacterioplankton lineage. ISME J. 9, 1423–1433 (2015)
Rodríguez-Ezpeleta, N. & Embley, T. M. The SAR11 group of alpha-proteobacteria is not related to the origin of mitochondria. PLoS ONE 7, e30520 (2012)
Viklund, J., Martijn, J., Ettema, T. J. G. & Andersson, S. G. E. Comparative and phylogenomic evidence that the alphaproteobacterium HIMB59 is not a member of the oceanic SAR11 clade. PLoS ONE 8, e78858 (2013)
Konstantinidis, K. T. & DeLong, E. F. Genomic patterns of recombination, clonal divergence and environment in marine microbial populations. ISME J. 2, 1052–1065 (2008)
Takami, H. et al. A deeply branching thermophilic bacterium with an ancient acetyl-CoA pathway dominates a subsurface ecosystem. PLoS ONE 7, e30559 (2012)
Kuwahara, H. et al. Reduced genome of the thioautotrophic intracellular symbiont in a deep-sea clam, Calyptogena okutanii. Curr. Biol. 17, 881–886 (2007)
Iobbi, C., Santini, C.-L., Bonnefoy, V. & Giordano, G. Biochemical and immunological evidence for a second nitrate reductase in Escherichia coli K12. Eur. J. Biochem. 168, 451–459 (1987)
Iobbi-Nivol, C., Santini, C. L., Blasco, F. & Giordano, G. Purification and further characterization of the second nitrate reductase of Escherichia coli K12. Eur. J. Biochem. 188, 679–687 (1990)
Philippot, L. Denitrifying genes in bacterial and Archaeal genomes. Biochim. Biophys. Acta 1577, 355–376 (2002)
Martinez-Espinosa, R. M. et al. Look on the positive side! The orientation, identification and bioenergetics of ‘Archaeal’ membrane-bound nitrate reductases. FEMS Microbiol. Lett. 276, 129–139 (2007)
Rothery, R. A., Workun, G. J. & Weiner, J. H. The prokaryotic complex iron-sulfur molybdoenzyme family. Biochim. Biophys. Acta 1778, 1897–1929 (2008)
Yoshimatsu, K., Iwasaki, T. & Fujiwara, T. Sequence and electron paramagnetic resonance analyses of nitrate reductase NarGH from a denitrifying halophilic euryarchaeote Haloarcula marismortui. FEBS Lett. 516, 145–150 (2002)
Lücker, S. et al. A Nitrospira metagenome illuminates the physiology and evolution of globally important nitrite-oxidizing bacteria. Proc. Natl Acad. Sci. USA 107, 13479–13484 (2010)
Starkenburg, S. R. et al. Genome sequence of the chemolithoautotrophic nitrite-oxidizing bacterium Nitrobacter winogradskyi Nb-255. Appl. Environ. Microbiol. 72, 2050–2063 (2006)
Sorokin, D. Y. et al. Nitrification expanded: discovery, physiology and genomics of a nitrite-oxidizing bacterium from the phylum Chloroflexi. ISME J. 6, 2245–2256 (2012)
Revsbech, N. P. et al. Determination of ultra-low oxygen concentrations in oxygen minimum zones by the STOX sensor: STOX oxygen sensor. Limnol. Oceanogr. Methods 7, 371–381 (2009)
Glass, J. B. et al. Meta-omic signatures of microbial metal and nitrogen cycling in marine oxygen minimum zones. Front. Microbiol. 6, 998 (2015)
Kopylova, E., Noé, L. & Touzet, H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28, 3211–3217 (2012)
Peng, Y., Leung, H. C. M., Yiu, S. M. & Chin, F. Y. L. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28, 1420–1428 (2012)
Zhu, W., Lomsadze, A. & Borodovsky, M. Ab initio gene identification in metagenomic sequences. Nucleic Acids Res. 38, e132 (2010)
Luo, C., Rodriguez-R, L. M. & Konstantinidis, K. T. MyTaxa: an advanced taxonomic classifier for genomic and metagenomic sequences. Nucleic Acids Res. 42, e73 (2014)
Raghunathan, A. et al. Genomic DNA amplification from a single bacterium. Appl. Environ. Microbiol. 71, 3342–3347 (2005)
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013)
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620 (2014)
Cox, M. P., Peterson, D. A. & Biggs, P. J. SolexaQA: at-a-glance quality assessment of Illumina second-generation sequencing data. BMC Bioinformatics 11, 485 (2010)
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012)
Lagesen, K. et al. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res. 35, 3100–3108 (2007)
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676 (2005)
Finn, R. D. et al. Pfam: the protein families database. Nucleic Acids Res. 42, D222–D230 (2014)
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690 (2006)
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011)
Konstantinidis, K. T. & Tiedje, J. M. Prokaryotic taxonomy and phylogeny in the genomic era: advancements and challenges ahead. Curr. Opin. Microbiol. 10, 504–509 (2007)
Suzek, B. E., Wang, Y., Huang, H., McGarvey, P. B. & Wu, C. H. UniRef clusters: a comprehensive and scalable alternative for improving sequence similarity searches. Bioinformatics 31, 926–932 (2015)
Castelle, C. J. et al. Extraordinary phylogenetic diversity and metabolic versatility in aquifer sediment. Nature Commun. 4, 2120 (2013)
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013)
Berger, S. A., Krompass, D. & Stamatakis, A. Performance, accuracy, and Web server for evolutionary placement of short sequence reads under maximum likelihood. Syst. Biol. 60, 291–302 (2011)
Rho, M., Tang, H. & Ye, Y. FragGeneScan: predicting genes in short and error-prone reads. Nucleic Acids Res. 38, e191 (2010)
Reddy, T. B. K. et al. The Genomes OnLine Database (GOLD) v.5: a metadata management system based on a four level (meta)genome project classification. Nucleic Acids Res. 43, D1099–D1106 (2015)
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990)
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nature Methods 12, 59–60 (2015)
Potter, L. C., Millington, P., Griffiths, L., Thomas, G. H. & Cole, J. A. Competition between Escherichia coli strains expressing either a periplasmic or a membrane-bound nitrate reductase: does Nap confer a selective advantage during nitrate-limited growth? Biochem. J. 344, 77–84 (1999)
Khlebnikov, A. & Keasling, J. D. Effect of lacY expression on homogeneity of induction from the Ptac and Ptrc promoters by natural and synthetic inducers. Biotechnol. Prog. 18, 672–674 (2002)
Alberge, F. et al. Dynamic subcellular localization of a respiratory complex controls bacterial respiratio n. eLife 4, e05357 (2015)
Hajaya, M. G. & Pavlostathis, S. G. Fate and effect of benzalkonium chlorides in a continuous-flow biological nitrogen removal system treating poultry processing wastewater. Bioresour. Technol. 118, 73–81 (2012)
Bender, K. S. et al. Identification, characterization, and classification of genes encoding perchlorate reductase. J. Bacteriol. 187, 5090–5096 (2005)
Jormakka, M., Richardson, D., Byrne, B. & Iwata, S. Architecture of NarGH reveals a structural classification of Mo-bisMGD enzymes. Structure 12, 95–104 (2004)
Acknowledgements
This work was supported by the National Science Foundation (1151698 to F.J.S. and 1416673 to K.T.K.), the NASA Exobiology Program (NNX14AJ87G to J.B.G. and F.J.S.), the Sloan Foundation (RC944 to F.J.S.), and a Community Science Program grant from the Department of Energy (DOE; to F.J.S. and K.T.K.). The work conducted by the US DOE Joint Genome Institute, a DOE Office of Science User Facility, is supported under contract no. DE-AC02-05CH11231. L.A.B., M.L. and B.T. were supported by a European Research Council Advanced Grant (OXYGEN, 267233) and by the Danish National Research Foundation (DNRF53). D.T. acknowledges the support of Onassis Foundation Fellowship. We are grateful for the generosity of J. Cole, A. Magalon, C. Sohaskey and F. Sargent for providing E. coli mutant strains, S. Pavlostathis for the ion chromatography methods and J. Spain for his suggestions on the heterologous expression experiment.
Author information
Authors and Affiliations
Contributions
D.T. conducted bioinformatics analyses. J.W., P.R. and N.S. conducted next-generation sequencing. C.C.P. and B.K.S. conducted qPCR analyses. L.M.R.-R developed additional bioinformatic methods for SAG contamination evaluation. R.R.M. and T.W. conducted cell sorting and SAG generation. L.A.B. and B.T. conducted process rate measurements. M.L. conducted STOX oxygen measurements. D.T., S.D., S.N., J.B.G. and A.S.B. conducted the heterologous expression experiments. F.J.S. and K.T.K. designed the study. F.J.S. and D.T. analysed the data and wrote the paper. All authors discussed the results and helped edit the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information
Nature thanks R. Kiene, D. Kirchman, R. Lasken and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Evaluation of contamination based on MyTaxa taxonomic affiliations.
a, Representative MyTaxa plots to test for contamination based on taxonomic affiliations of predicted genes. The MyTaxa algorithm51 predicts the taxonomic affiliation on the basis of a weighted classification scheme that takes into account the phylogenetic signal of each protein family. Each gene is assigned to the deepest taxonomic resolution (out of phylum, genus and species) for which a high-confidence value can be obtained (score 0.5). Each MyTaxa scan represents taxonomic distributions of all the predicted genes for one genome, given in windows of 10 genes, and sorted based on their position in the concatenated assembly of the genome (when a partial genome is used). a, b, White space in the histograms represents genes that could not be assigned to a given taxon due to (1) lack of BLASTP hits against the reference database (a collection of closed and draft genomes) or (2) lack of high confidence scores. Notice that for the representative OMZ SAG E5, more than 80% of the genes can be classified as Candidatus Pelagibacter (SAR11), with an additional 10% assigned to Proteobacteria. Note there are no genome representatives for this taxon (that is, SAR11 subclade IIa.A) in the database upon which MyTaxa is based. Similar results are obtained for the bathytype SAR11 SAG6, as this genome also lacks representatives. The closed genome from a coastal isolate HTCC1002 is shown for comparison to demonstrate a typical pattern for cases when close relatives of the query genome are available in the reference database, as is the case for this isolate. b, Taxonomic classifications of genes from the 19 SAGs analysed here. Each distribution was obtained from the MyTaxa scans performed for each SAG. The percentage of the total genes that could be taxonomically classified with MyTaxa was on average ~60%, and varied depending on the completeness of the genome (that is, partial genes are less likely to be assigned taxonomy with high confidence). These values are also reported in Supplementary Table 1. Of the genes that could be classified, the majority (>90%) were classified to SAR11 taxa.
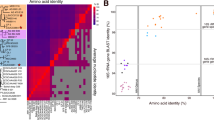
Extended Data Figure 2 Microdiversity within the SAR11 populations.
a, Recruitment plot of metagenomic reads from the ETNP OMZ 300 m sample, against scaffolds from SAG E4. Notice that the recruited reads vary in identities from 100% down to 85%, indicating the presence of closely affiliated clades, as well as extensive microdiversity within the same clade (that is, reads sharing >95% identity). b, Phylogenetic reconstruction of reference RpoB protein sequences from SAR11 genomes, and placement of identified RpoB metagenomic sequences (denoted with the cross symbols). The alignment length was 1,406 columns with 5.9% gaps or undetermined sites. The presence of multiple divergent rpoB reads within the same subclade (predominantly for subclades IIa.A and Ic) suggests high abundance but also extensive microdiversity within those populations (rather than clonal populations).
Extended Data Figure 3 nar genes encoded by SAR11 populations of OMZs.
a, nar operon and adjacent genes identified in SAR11 SAGs from the ETNP OMZ, and in assemblies from the 85 m and 300 m ETNP OMZ metagenomes. narG sequences with at least 97% amino acid similarity are represented with the same colour. b, c, Representative maximum likelihood phylogeny to show sequence variation among full-length or near full-length narG (b) and narH (c) amino acid sequences identified in the SAGs. A subset of cytoplasm-oriented Nar and Nxr enzymes from publicly available genomes is also included. A comprehensive phylogeny showing the placement of SAR11 nar sequences relative to enzymes (n = 392) of the DMSO family is in Fig. 2a. Coloured pies represent the placement of shorter narG/narH gene fragments identified in the SAGs. Bootstrap values over 50 are shown. Outgroups (arrows) are E. coli dmsA (b) and dmsB (c). Note that the Gamma-type nar-containing contig recovered in E4 (Fig. 2a) contains narHJI, but not narG; E4 Gamma-type is therefore not represented in Fig. 3b. All genes co-localized in the nar-containing contigs are listed in Supplementary Table 5. The p-numbers are gene identifiers given by the gene prediction software, consistent with those in Supplementary Table 5.
Extended Data Figure 4 Identified NarG in SAR11 SAGs are members of the DMSO superfamily of oxidoreductases.
a, Phylogenetic reconstruction of NarG and DMSO enzymes. The tree shown in Fig. 2 is presented here but has been expanded to include diverse DMSO oxidoreductases for direct comparison with the NarG/NxrA enzymes. Notice that both OP1 (green, blue, grey) and Gamma-type (red, orange) variants cluster within the cytoplasmically oriented Nar and Nxr enzymes. Six-hundred and ninety-seven NarG/NxrA proteins were identified from UniRef63, and from those, 321 full-length sequences were selected to represent all the diverse clades. An additional 71 non-NarG/NxrA proteins, representative of the diverse enzymes of the DMSO superfamily were also included in the collection. The full-length amino acid sequences were aligned with Clustal Omega61 and the phylogenetic tree was constructed by maximum likelihood and 1,000 bootstraps using RAxML60. The alignment length was 1,803 columns, out of which 31.2% were gaps or undetermined. Partial NarG sequences identified in the SAGs were placed on the tree using the epa algorithm from RAxML66. The same collection of proteins was used to train the Rocker models and quantify the narG metagenomic fragments, and can be found in the enve-omics website (http://enve-omics.ce.gatech.edu/rocker/models). b, Alignment of NarG sequences from OMZ SAR11 with representative sequences from the DMSO superfamily of oxidoreductases. The protein motifs in the second and third panels are present in all functional Nar enzymes (NarG) and Nxr enzymes (NxrA) but not in closely related enzymes of the DMSO superfamily. The first panel shows the presence/absence of the TAT signal peptide (SRRSFLK), whose presence typically denotes a protein excreted to the outer membrane40,41. SAR11 NarG is instead oriented towards the cytoplasm (lack of TAT). The second panel shows the cysteine-rich motif typically found in the N terminus of the type-II DMSO superfamily oxidoreductases75 and believed to enable the formation of a [4Fe–4S] cluster in these proteins76. The Asn in position 158 of the alignment is typically found in catalytic subunits of nitrite reductases and DMSO oxidoreductases (DmsA) but not in other DMSO family enzymes. The third panel shows the Gln(Q) and Thr(T) in positions 398 and 399 within the putative substrate entry channel of the protein, which differentiate the Nar proteins from all other oxidoreductases of the DMSO family40.
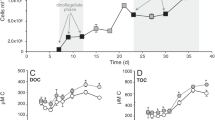
Extended Data Figure 5 Functional characterization of the SAR11 nar operons in the E. coli heterologous expression system.
a, Genotype of the E. coli triple mutant confirmed by whole-genome sequencing. The triple mutant lacks complete functional operons of all three NO3− reductase enzymes, and thus is incapable of NO3− reduction. b, Anaerobic growth of triple-mutant clones, complemented with the SAR11 nar operons. For each strain three independent clones were monitored, and data from the replicate growth curves were fitted into a logistic model. Shaded areas represent the 95% confidence intervals of optical density readings (OD600 nm) in the fitted logistic growth models. NO3− and NO2− were measured in parallel with ion chromatography. Note that the Gamma-type SAR11 operon complements the triple-mutant phenotype, growing anaerobically by reducing NO3− to NO2−. E. coli encodes functional nitrite reductases, thus the accumulated NO2− can be further reduced to ammonia, accounting for the non-stoichiometric NO2− production. c, Whole-cell NO2− production assays under aerobic conditions. Eight independent clones (columns A–H) of each type (C1–C5) were inoculated in Luria–Bertani (LB) broth supplemented with 30 mM NO3− and different isopropyl-β-d-thiogalactoside (IPTG) concentrations, and the well plate was incubated for 2 days at room temperature. Griess reagent was added, and development of pink colour indicated NO2− production.
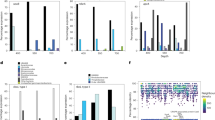
Extended Data Figure 6 Relative abundance of narG variants in ETNP OMZ metagenomes and metatranscriptomes and various other ocean metagenomes.
a, Relative abundance and diversity of NarG/NxrA enzymes as revealed by phylogenetic placement of identified narG metagenomic reads (coloured pies). All identified short metagenomic narG reads from various oceanic metagenomes were placed within a reconstructed reference NarG tree to estimate the abundance of the different narG variants. The results of the placement are presented in five separate trees, based on the origin of the analysed metagenomic reads (ETSP metagenomes, ETNP metagenomes and metatranscriptomes, oxic bathypelagic and oxic surface metagenomes) for clarity. In each of the five trees, the coloured pies represent the abundance (normalized for data set size) of the short metagenomic reads clustering in the respective node. Specifically, the pie radius reflects read abundance as a percentage of the total narG genome equivalents identified (that is, number of narG reads compared to number of rpoB reads, normalized for gene length and total number of reads in each metagenome), with the size of grey pies representing the highest and lowest relative abundance, respectively. The reference tree is the same as in Fig. 3a. Scale bars represent substitutions per amino acid. Notice that the two narG variants affiliated with the SAR11 SAGs (highlighted in orange for the OP1 type and blue for the Gamma type) are only abundant in the metagenomes and metatranscriptomes from the OMZ, where they comprise more than 70% of the total narG read pool, as can also be observed in Fig. 3b and c. The number of narG reads of the OP1 or Gamma type are also given in Supplementary Table 1. b, qPCR-based abundance of SAR11-affiliated narG genes in the ETNP OMZ relative to NO2−, NO3− and O2 concentrations and qPCR-based counts of 16S rRNA. Counts of total bacterial 16S rRNA, OP1-type narG, and Gamma-type narG genes at three stations (map on legend) west of Manzanillo, Mexico in May 2014. Map was created with Ocean Data View (http://odv.awi.de). All assays were performed in triplicates, and the bars represent s.e.m. Note that counts of OP1- and Gamma-type narG variants are probably underestimates given the observed microdiversity in the community (Extended Data Figs 2 and 7), and therefore there is a possibility that our primers did not match all OP1- and Gamma-type variants.
Extended Data Figure 7 Diversity of OP1 and Gamma-type narG amino acid sequences in the ETNP OMZ metagenome.
a, Phylogenies showing all full-length narG sequences recovered in the ETNP OMZ metagenomes (85, 100, 125, 300 m), as well as those from the SAR11 SAGs and corresponding narG reference sequences, with the left tree showing OP1-type variants and the right tree showing Gamma-type variants. NarG sequences are colour-coded based on the taxonomic classification of adjacent genes in the same metagenomic scaffolds, as show in Supplementary Table 6. b, Recruitment of metagenomic reads (predicted open reading frames) from the OMZ 300 m sample, against OP1- (left) or Gamma- (right) type narG sequences from the SAR11 SAGs. The metagenomic reads used for recruitment were identified as ‘narG’ using the ROCker pipeline, and their identity was further confirmed by phylogenetic placement within the narG clade on a reference DMSO superfamily protein tree, to minimize non-specific recruitments in conserved protein regions. Note that based on this analysis, the OP1-type narG variants are highly diverse in the OMZ metagenome.
Extended Data Figure 8 Transcriptional profile of predicted genes from the SAR11 OMZ SAG-D9.
Transcriptomic reads with >99% identity matches were counted for each gene, and the counts were normalized for the data set size. Note that the nar operon genes are among the most actively transcribed in the ETNP 300 m OMZ sample.
Supplementary information
Supplementary Information
This file contains Supplementary Results and Discussion, additional references and full legends for Supplementary Tables 1-6. (PDF 368 kb)
Supplementary Data
This file contains Supplementary Tables 1-6 – see the Supplementary Information document for full table legends. (XLSX 125 kb)
Rights and permissions
About this article
Cite this article
Tsementzi, D., Wu, J., Deutsch, S. et al. SAR11 bacteria linked to ocean anoxia and nitrogen loss. Nature 536, 179–183 (2016). https://doi.org/10.1038/nature19068
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19068
This article is cited by
-
Stratified microbial communities in Australia’s only anchialine cave are taxonomically novel and drive chemotrophic energy production via coupled nitrogen-sulphur cycling
Microbiome (2023)
-
Ecophysiology and genomics of the brackish water adapted SAR11 subclade IIIa
The ISME Journal (2023)
-
Partitioning of the denitrification pathway and other nitrite metabolisms within global oxygen deficient zones
ISME Communications (2023)
-
A compendium of bacterial and archaeal single-cell amplified genomes from oxygen deficient marine waters
Scientific Data (2023)
-
Function and distribution of nitrogen-cycling microbial communities in the Napahai plateau wetland
Archives of Microbiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.