Abstract
During pre-implantation development, the mammalian embryo self-organizes into the blastocyst, which consists of an epithelial layer encapsulating the inner-cell mass (ICM) giving rise to all embryonic tissues1. In mice, oriented cell division, apicobasal polarity and actomyosin contractility are thought to contribute to the formation of the ICM2,3,4,5. However, how these processes work together remains unclear. Here we show that asymmetric segregation of the apical domain generates blastomeres with different contractilities, which triggers their sorting into inner and outer positions. Three-dimensional physical modelling of embryo morphogenesis reveals that cells internalize only when differences in surface contractility exceed a predictable threshold. We validate this prediction using biophysical measurements, and successfully redirect cell sorting within the developing blastocyst using maternal myosin (Myh9)-knockout chimaeric embryos. Finally, we find that loss of contractility causes blastomeres to show ICM-like markers, regardless of their position. In particular, contractility controls Yap subcellular localization6, raising the possibility that mechanosensing occurs during blastocyst lineage specification. We conclude that contractility couples the positioning and fate specification of blastomeres. We propose that this ensures the robust self-organization of blastomeres into the blastocyst, which confers remarkable regulative capacities to mammalian embryos.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wennekamp, S., Mesecke, S., Nédélec, F. & Hiiragi, T. A self-organization framework for symmetry breaking in the mammalian embryo. Nature Rev. Mol. Cell Biol. 14, 452–459 (2013)
Anani, S., Bhat, S., Honma-Yamanaka, N., Krawchuk, D. & Yamanaka, Y. Initiation of Hippo signaling is linked to polarity rather than to cell position in the pre-implantation mouse embryo. Development 141, 2813–2824 (2014)
Samarage, C. R. et al. Cortical tension allocates the first inner cells of the mammalian embryo. Dev. Cell 34, 435–447 (2015)
Hirate, Y. et al. Polarity-dependent distribution of angiomotin localizes Hippo signaling in preimplantation embryos. Curr. Biol. 23, 1181–1194 (2013)
Dard, N., Louvet-Vallée, S. & Maro, B. Orientation of mitotic spindles during the 8- to 16-cell stage transition in mouse embryos. PLoS ONE 4, e8171 (2009)
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–183 (2011)
Johnson, M. H. From mouse egg to mouse embryo: polarities, axes, and tissues. Annu. Rev. Cell Dev. Biol. 25, 483–512 (2009)
Watanabe, T., Biggins, J. S., Tannan, N. B. & Srinivas, S. Limited predictive value of blastomere angle of division in trophectoderm and inner cell mass specification. Development 141, 2279–2288 (2014)
Johnson, M. H. & Ziomek, C. A. The foundation of two distinct cell lineages within the mouse morula. Cell 24, 71–80 (1981)
Matsumoto, M. et al. PKCλ in liver mediates insulin-induced SREBP-1c expression and determines both hepatic lipid content and overall insulin sensitivity. J. Clin. Invest. 112, 935–944 (2003)
Hirate, Y. et al. Par-aPKC-dependent and -independent mechanisms cooperatively control cell polarity, Hippo signaling, and cell positioning in 16-cell stage mouse embryos. Dev. Growth Differ. 57, 544–556 (2015)
Maître, J.-L., Niwayama, R., Turlier, H., Nédélec, F. & Hiiragi, T. Pulsatile cell-autonomous contractility drives compaction in the mouse embryo. Nature Cell Biol. 17, 849–855 (2015)
Lehtonen, E. Changes in cell dimensions and intercellular contacts during cleavage-stage cell cycles in mouse embryonic cells. J. Embryol. Exp. Morphol. 58, 231–249 (1980)
Heisenberg, C.-P. & Bellaïche, Y. Forces in tissue morphogenesis and patterning. Cell 153, 948–962 (2013)
Overholtzer, M. et al. A nonapoptotic cell death process, entosis, that occurs by cell-in-cell invasion. Cell 131, 966–979 (2007)
Dietrich, J.-E. & Hiiragi, T. Stochastic patterning in the mouse pre-implantation embryo. Development 134, 4219–4231 (2007)
Johnson, M. H. & Ziomek, C. A. Cell interactions influence the fate of mouse blastomeres undergoing the transition from the 16- to the 32-cell stage. Dev. Biol. 95, 211–218 (1983)
Graner, F. & Glazier, J. A. Simulation of biological cell sorting using a two-dimensional extended Potts model. Phys. Rev. Lett. 69, 2013–2016 (1992)
Brodland, G. W. The Differential Interfacial Tension Hypothesis (DITH): a comprehensive theory for the self-rearrangement of embryonic cells and tissues. J. Biomech. Eng. 124, 188–197 (2002)
Guzowski, J., Korczyk, P. M., Jakiela, S. & Garstecki, P. The structure and stability of multiple micro-droplets. Soft Matter 8, 7269–7278 (2012)
Da, F., Batty, C. & Grinspun, E. Multimaterial mesh-based surface tracking. ACM Trans. Graph. 33, 112 (2014)
Krieg, M. et al. Tensile forces govern germ-layer organization in zebrafish. Nature Cell Biol. 10, 429–436 (2008)
Maître, J.-L. et al. Adhesion functions in cell sorting by mechanically coupling the cortices of adhering cells. Science 338, 253–256 (2012)
Jacobelli, J. et al. Confinement-optimized three-dimensional T cell amoeboid motility is modulated via myosin IIA-regulated adhesions. Nature Immunol. 11, 953–961 (2010)
Wang, A. et al. Nonmuscle myosin II isoform and domain specificity during early mouse development. Proc. Natl Acad. Sci. USA 107, 14645–14650 (2010)
Aragona, M. et al. A mechanical checkpoint controls multicellular growth through YAP/TAZ regulation by actin-processing factors. Cell 154, 1047–1059 (2013)
Benham-Pyle, B. W., Pruitt, B. L. & Nelson, W. J. Mechanical strain induces E-cadherin-dependent Yap1 and β-catenin activation to drive cell cycle entry. Science 348, 1024–1027 (2015)
Shin, J.-W. et al. Contractile forces sustain and polarize hematopoiesis from stem and progenitor cells. Cell Stem Cell 14, 81–93 (2014)
Riedl, J. et al. Lifeact mice for studying F-actin dynamics. Nature Methods 7, 168–169 (2010)
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007)
Balbach, S. T. et al. Nuclear reprogramming: kinetics of cell cycle and metabolic progression as determinants of success. PLoS ONE 7, e35322 (2012)
de Vries, W. N. et al. Expression of Cre recombinase in mouse oocytes: a means to study maternal effect genes. Genesis 26, 110–112 (2000)
Ma, X. et al. Conditional ablation of nonmuscle myosin II-B delineates heart defects in adult mice. Circ. Res. 105, 1102–1109 (2009)
Acknowledgements
We are grateful to the Hiiragi laboratory members and the European Molecular Biology Laboratory (EMBL) animal facility for their support. We thank F. Da and C. Batty for discussions on the simulations. We thank Y. Bellaïche for comments on an earlier version of the manuscript. Marie Curie individual fellowships under FP7 and H2020 programs support J.-L.M., H.T. and R.N. under Research Executive Agency grant agreements 329044, 656306 and 326701, respectively. H.T. acknowledges support from the Bettencourt-Schueller and Joachim Herz foundations. The Hiiragi laboratory is supported by EMBL, the European Research Council and VolkswagenStifftung.
Author information
Authors and Affiliations
Contributions
J.-L.M. designed the project and experiments, and wrote the manuscript with input from all authors. J.-L.M. and B.E. performed and analysed the tension and lineage mapping experiments. J.-L.M. and R.I. performed and analysed the remaining experiments. R.N. helped with image analysis of the periodic contractions. H.T. designed the physical model and performed the simulations with help from F.N. T.H. supervised the study and helped design the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks D. Discher, P.-F. Lenne, B. Plusa and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 aPKC antagonizes myosin phosphorylation at the apical domain.
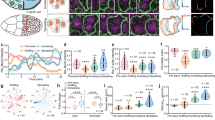
a–c, Immunostaining of 16- (a) and 8-cell-stage wild-type (b) and aPKC-knockout (c) embryos showing aPKC (red), phalloidin (blue) and bi-phosphorylated myosin regulatory light chain (ppMRLC; green). Enlarged images of ppMRLC are shown on the far right. d–f, Cortical intensity profiles under the dotted lines in the far right panels of a–c. Apical domains are highlighted in orange and non-apical regions in blue. g, h, Box plot of unpolarized/polarized blastomere intensity ratio at the 16-cell stage (43 neighbouring blastomeres from 35 embryos from 3 experiments) and non-apical/apical intensity ratio at the 8-cell stage for wild-type (WT) and aPKC-knockout (KO) embryos (68 and 58 blastomeres from 22 and 12 embryos from 2 and 3 experiments, respectively). ppMRLC is in green, aPKC in red and phalloidin in blue. Student’s t-test P values between wild type and aPKC knockout. At the 16-cell stage (a, d, g), blastomeres showing accumulations of ppMRLC and phalloidin at their cell-medium interfaces have less aPKC. At the 8-cell stage, the apical domain does not occupy the entirety of the cell-medium interface (b, e). The aPKC-rich apical domain shows less myosin than the aPKC-poor region of the cortex (b, e, h). In aPKC-knockout embryos (c, f, h), no aPKC-rich, nor ppMRLC-poor regions can be observed at the cell-medium interface of blastomeres. i, Immunostaining of 8-cell-stage blastomeres showing aPKC (red), phalloidin (blue), ppMRLC (green) and merged staining. The apical domain is highlighted in orange, the non-apical cortex in blue. Scale bar, 10 μm. j, Intensity profile along the cell perimeter showing aPKC (red), phalloidin (blue) and ppMRLC (green). The apical intensity is highlighted in orange, the non-apical cortex in blue. k, Box plot of apical and non-apical intensity ratio of cortical aPKC (red), phalloidin (blue) and ppMRLC (green) for 27 blastomeres from 3 experiments. Blastomeres isolated at the 8-cell stage show an aPKC-rich region (i–k), which has less cortical ppMRLC and phalloidin than the aPKC-poor region of the cell-medium interface. The ppMRLC- and actin-rich regions are distinct from the basolateral domain of cells (their cell–cell contact), which have less ppMRLC and actin (a–c, l)12. This ppMRLC cortical region is therefore labelled ‘non-apical’. l, Immunostaining of doublets of 16-cell-stage blastomeres showing aPKC (red), phalloidin (blue), ppMRLC (green) and merged staining. The polarized blastomere is highlighted in orange, the unpolarized one in blue. Scale bar, 10 μm. m, Intensity profile along the doublet perimeter showing aPKC (red), phalloidin (blue) and ppMRLC (green). The polarized blastomere is highlighted in orange, the unpolarized one in blue. Blastomeres isolated at the 8-cell stage divide to give rise to doublets of 16-cell-stage blastomeres2,16. The polarized sister cell shows high aPKC and low ppMRCL/phalloidin at their cell-medium interfaces when compared to the non-polarized sister cell (l, m). n, Cortical intensity ratio of ppMRLC (green) and phalloidin (blue) between the inner and outer cells as a function of the inner contact angles θ1 (Pearson R = 0.464 and 0.614, n = 67 doublets from 2 experiments, P < 0.001). During the 16-cell stage, polarized blastomeres can envelop their unpolarized sister blastomeres (Supplementary Video 5)2,16. As envelopment occurs, the internal contact angles change (Extended Data Fig. 2). As the internal contact angles change, the asymmetry in cortical ppMRLC and phalloidin between sister blastomeres changes. After another division, a cyst consisting of four blastomeres forms (Supplementary Video 5). This structure is equivalent to the blastocyst in terms of gene expression16 (Fig. 4). o, p, Immunostaining of 16- (o) and 8-cell-stage (p) embryos showing aPKC (red), phalloidin (blue) and myosin heavy chain phosphorylated on S1943 (pMyh9; green). Enlarged images of ppMRLC are shown on the far right. q, r, Cortical intensity profiles under the dotted lines on the far right of o, p. Apical domains are highlighted in orange and non-apical regions in blue. s, t, Box plot of unpolarized/polarized blastomere intensity ratio at the 16-cell stage (24 neighbouring blastomeres from 16 embryos from 3 experiments) and non-apical/apical intensity ratio at the 8-cell stage (34 blastomeres from 10 embryos from 3 experiments). pMyh9 in green, aPKC in red and phalloidin in blue.
Extended Data Figure 2 Cortical asymmetries intensify during the 16-cell stage.
a, Time-lapse of mTmG (magenta) and LifeAct–GFP (green) expressing doublets of 16-cell-stage blastomeres. Scale bar, 10 μm. b, c, External (θc), and internal (θ1 and θ2) contact angles (b) and cortical LifeAct–GFP intensities of unpolarized I1 and polarized I2 blastomeres and intensity ratio I1/I2 (c) over time for the doublet shown in a. Blastomeres isolated at the 8-cell stage can divide asymmetrically to give rise to a polarized blastomere that will envelop its unpolarized sister blastomere (a). The external contact angle θc shows a rapid re-compaction of the cell doublet after division (b). The internal contact angles θ1 and θ2 indicate the progression of the envelopment process (b). As this happens, the cortical intensity of LifeAct–GFP of the internalizing blastomere I1 increases while the one of the enveloping blastomere I2 remains comparably more stable (c). This increases the cortical asymmetry I1/I2 (c). d, Initial cortical asymmetry over internalization time of doublets of 16-cell-stage blastomeres (Pearson R = 0.064, n = 16 doublets from 4 experiments, P > 0.1). The initial cortical asymmetry, calculated within 30 min after division, is 1.0 ± 0.1 (mean ± s.d., n = 17 doublets from 4 experiments) and does not control the time it takes for envelopment to occur (d). e, Intensity ratio as a function of the contact angle θ1 of doublets throughout the 16-cell-stage blastomeres (Pearson R = 0.573, 186 measurements on 17 asymmetric doublets (purple), P < 0.001 and Pearson R = 0.266, 69 measurements on 3 symmetric (green) doublets, P < 0.1, from 4 experiments). As the internal contact angles change, the asymmetry in cortical LifeAct–GFP between sister blastomeres with distinct polarity changes. f, Cortical intensity ratio increase rate as a function of the contact angle θ1 increase rate (Pearson R = 0.824, n = 17 asymmetric (purple), P < 0.001, and Pearson R = 0.393, n = 3 symmetric (green) doublets, P > 0.1, from 4 experiments). g, Cortical intensity increase rate as a function of the contact angle increase rate for the polarized (orange, Pearson R = −0.026, n = 17 asymmetric doublets, P > 0.1) and unpolarized blastomere (blue, Pearson R = 0.658, n = 17 asymmetric doublets, P < 0.01) of a doublet resulting from asymmetric division or of two polarized cells resulting from a symmetric division (green, Pearson R = −0.011, n = 3 symmetric doublets, P > 0.1), from 4 experiments. The rates are correlated, which suggests that the dynamics of internalization and the dynamics of building up of cortical asymmetries are linked.
Extended Data Figure 3 Cell size has no influence on internalization.
Phase diagram describing the mechanical equilibrium of a cell within a doublet or embryo as function of the cell size asymmetry parameter β and the tension asymmetry parameter δ, for a fixed compaction parameter α = 0.25. The colour code measures the degree of internalization, defined as the proportion of internalized volume Vin/V1, which equals 1 for the internalized cell. The dotted line indicates the threshold value δc at which internalization occurs. An example of internalization with β = 0.5 is indicated in black (from A to E). Changing the volume asymmetry does not change the internalization threshold. Internalization of a doublet with β = 0.5 obtained with the analytical model for the same values of δ as indicated in the diagram from A to E.
Extended Data Figure 4 Contractility is required for internalization.
Brightfield images of tension measurement on wild-type (WT; top) and mMyh9 (bottom) 8-cell-stage embryos. Scale bar, 10 μm. Mean ± s.d. of 25 blastomeres from 4 wild-type embryos and 26 blastomeres from 7 mMyh9 embryos from 2 experiments, Student’s t-test P < 10−9.
Extended Data Figure 5 Control of pYap and Cdx2 localization by contractility in a dose-dependent manner.
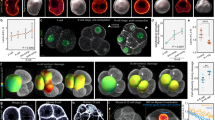
a–g, Immunostaining of wild-type embryos treated for 3 h with Bb(+) at 5 (a), 12.5 (c) or 25 (e) μM or with Bb(−) at 5 (b), 12.5 (d) or 25 (f) μM or of mMyh9 embryos (g) showing pYap (green), Cdx2 (red) and phalloidin (blue). h–u, Nucleus-to-cytoplasm intensity ratio of pYap (h–n) or Cdx2 (o–u) as a function of the distance from the surface for wild-type embryo treated with Bb(+) (outer cells in orange and inner cells in blue) at 5 (h, o, corresponding embryo shown in a), 12.5 (j, q, corresponding embryo shown in c) or 25 (l, s, corresponding embryo shown in e) μM or with Bb(−) (outer cells in red and inner cells in pink) at 5 (i, p, corresponding embryo shown in b), 12.5 (k, r, corresponding embryo shown in d) or 25 (m, t, corresponding embryo shown in f) μM and for mMyh9 embryos (n, u, corresponding embryo shown in g). v, w, Mean ± s.e.m. Pearson correlation values between the nucleus to cytoplasm intensity ratio of pYap (v) or Cdx2 (w) as a function of the distance from the surface from individual embryos. Two-hundred and seven blastomeres from 20 embryos for Bb(+) 5 μM, 252 blastomeres from 29 embryos for Bb(+) 12.5 μM, 179 blastomeres from 18 embryos for Bb(−) 5 μM and 267 blastomeres from 28 embryos for Bb(−) 12.5 μM from 3 experiments each. Two-hundred and eighty-one cells from 28 embryos from 5 experiments for pYap and 136 cells from 13 embryos from 4 experiments for Cdx2 for 25 μM Bb(+), 241 cells from 32 embryos from 5 experiments for pYap and 192 cells from 22 embryos from 4 experiments for Cdx2 for 25 μM Bb(−), and 349 cells from 32 embryos from 6 experiments for pYap and 217 cells from 21 embryos from 3 experiments for Cdx2 for mMyh9. Student’s t-test P values; NS, not significant.
Extended Data Figure 6 Contractility controls Yap subcellular localization.
a–c, Immunostaining of wild-type embryos treated with 25 μM Bb(+) (a; an inactive enantiomere of the inhibitor) or Bb(−) (b; the selective inhibitor of myosin II ATPase activity) for 3 h or mMyh9 embryos (c) showing Yap (green), pYap (red) and phalloidin (blue). d–f, Nucleus-to-cytoplasm intensity ratio of pYap (left) and Yap (right) as a function of the distance from the surface for wild-type embryo treated with 25 μM Bb(+) (d; outer cells in orange and inner cells in blue, corresponding embryo shown in a) or Bb(−) (e; outer cells in magenta and inner cells in red, corresponding embryo shown in b) or mMyh9 embryo (f; outer cells in dark green and inner cells in light green, corresponding embryo shown in c). g, Mean ± s.e.m. Pearson correlation values between the nucleus to cytoplasm intensity ratio of Yap as a function of the distance from the surface from individual embryos. Two-hundred and fifty-two cells from 29 embryos for Bb(+), 201 cells from 26 embryos for Bb(−) and 132 cells from 12 embryos for mMyh9 from 3 experiments each. Student’s t-test P value is shown; NS, not significant.
Extended Data Figure 7 Quantitative comparison between analytical and numerical results.
a, Comparison of the surface areas of the cell medium (blue) and cell–cell interfaces (green) between the simulations (crosses) and the analytical model (lines) for different values of the compaction parameter α between 0 and 1. A schematic diagram of a cell doublet defining the cell medium and cell–cell surface tensions γ1, γ2 and γc and areas A1, A2 and Ac are shown as an inset. b, Configurations of doublets as predicted by the analytical model and simulations for the discrete values of α corresponding to the plot in a. c, Comparison of the surface areas of the cell-medium interfaces of cell 1 (blue), 2 (orange) and of the cell–cell interface (green) between the simulations (crosses) and the analytical model (lines) for different values of the tension asymmetry parameter δ between 1 and 1.6 fixed compaction parameter α = 0.25. d, Configurations of doublets as predicted by the analytical model and simulations for the discrete values of δ corresponding to the plot in c.
Supplementary information
Supplementary Information
This file contains Supplementary Text and Data, Supplementary Figures 1-2 and additional references. (PDF 1577 kb)
Periodic contractions in a polarized 8-cell stage blastomere
Montage of mTmG (left) and local curvature measurement (right) of a polarized 8-cell stage blastomere showing periodic contractions. Time-lapse imaging consists of an image taken every 5 s and is displayed at 10 fps. Scale bar 10 μm. (AVI 1290 kb)
Periodic contractions of in a 16-cell stage doublet
Montage of mTmG (left) and local curvature measurement (right) of a doublet of a polarized and unpolarized 16-cell stage blastomeres showing periodic contractions. Time-lapse imaging consists of an image taken every 5 s and is displayed at 5 fps. Scale bar 10 μm. (AVI 1355 kb)
Numerical simulation of the compaction and internalization of a doublet
Numerical simulation of a cell doublet undergoing compaction (compaction parameter α decreasing from 0.9 to 0.25) followed by internalization (tension asymmetry parameter δ increasing from 1.0 to 1.6 with α kept at 0.25). (MP4 59 kb)
Numerical simulation of the compaction of a 16-cell embryo followed by the internalization of one cell
Numerical simulation of a 16-cell embryo undergoing compaction (compaction parameter α decreasing from 0.9 to 0.25) followed by the internalization of one cell (tension asymmetry parameter δ increasing from 1.0 to 1.6 with α kept at 0.25). (MP4 220 kb)
Internalization of an asymmetric doublet
Montage of mTmG (left), LifeAct-GFP (middle), merged image (right) of a polarized 8-cell stage blastomere undergoing asymmetric division to form an doublet of a polarized and unpolarized blastomere. The top panels show a single confocal slice whereas maximum projections of all slices are shown on the bottom. The polarized blastomere envelops the unpolarized one before the next division, which gives rise to a mini-blastocyst equivalent to a 32-cell stage embryo composed of four blasotmeres. Time-lapse imaging consists of an image taken every 30 min and is displayed at 5 fps. Scale bar 10 μm. (AVI 11749 kb)
Surface tension and lineage mapping
Fluorescent imaging of a H2B-GFP (magenta) and mTmG (green) expressing embryo from the 8- to 32-cell stage. Bright-field imaging shows the micropipette used for surface tension measurement. Time-lapse imaging consists of an image taken every 30 min and is displayed at 5 fps except when surface tension measurements and cell positions in the blastocyst are shown. Scale bar 10 μm. (AVI 21087 kb)
Pre-implantation development of mzMyh10 embryos
Bright-field imaging of maternal zygotic Myh10 knockout (mzMyh10) embryos developing from the 4- to 32-cell stage. Time-lapse imaging consists of an image taken every 30 min and is displayed at 10 fps. Scale bar 10 μm. (AVI 11164 kb)
Development of a mMy9h embryo
Fluorescent (green) and bright-field (grey) imaging of a mMyh9 mTmG embryo developing from the 4- to 32-cell stage. Time-lapse imaging consists of an image taken every 30 min and is displayed at 10 fps. Scale bar 10 μm. (AVI 5054 kb)
Internalization of a WT blastomere within a mMyh9 host
Fluorescent imaging of a WT mG (magenta) blastomere internalizing a mMyh9 mTmG (green) host. Time-lapse imaging consists of an image taken every 30 min and is displayed at 5 fps. Scale bar 10 μm. (AVI 3900 kb)
Numerical simulation of the internalization of a cell of higher contractility grafted onto a 16-cell compacted embryo
Numerical simulation the internalization of a cell of higher contractility (fixed tension asymmetry parameter δ = 1.6 and compaction parameter decreasing from α = 0.8 to 0.25, both relative to the embryo) grafted onto a homogeneous and compacted 16-cell stage embryo (fixed internal compaction parameter α = 0.25). (MP4 107 kb)
Spreading of a mMyh9 blastomere onto a WT host
Fluorescent imaging of a mMyh9 mTmG (green) blastomere grafted onto a WT mG (magenta) host. Time-lapse imaging consists of an image taken every 30 min and is displayed at 5 fps. Scale bar 10 μm. (AVI 2562 kb)
Numerical simulation of the spreading of a cell of lower contractility grafted onto a compacted 16-cell stage embryo
Numerical simulation of the spreading of a cell of lower contractility (fixed tension asymmetry parameter δ = 0.5 and compaction parameter decreasing from α = 0.8 to 0.25, both relative to the embryo) grafted onto a homogeneous and compacted 16-cell stage embryo (fixed internal compaction parameter α = 0.25). (MP4 107 kb)
Internalization of a WT blastomere within a WT host after cell division
Fluorescent imaging of a WT mTmG (magenta) blastomere internalizing a WT mG (cyan) host. Time-lapse imaging consists of an image taken every 30 min and is displayed at 5 fps. Scale bar 10 μm. (AVI 2151 kb)
Rights and permissions
About this article
Cite this article
Maître, JL., Turlier, H., Illukkumbura, R. et al. Asymmetric division of contractile domains couples cell positioning and fate specification. Nature 536, 344–348 (2016). https://doi.org/10.1038/nature18958
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature18958
This article is cited by
-
SimuCell3D: three-dimensional simulation of tissue mechanics with cell polarization
Nature Computational Science (2024)
-
Mechanics of human embryo compaction
Nature (2024)
-
Embryo mechanics cartography: inference of 3D force atlases from fluorescence microscopy
Nature Methods (2023)
-
The nuclear lamina couples mechanical forces to cell fate in the preimplantation embryo via actin organization
Nature Communications (2023)
-
A monoastral mitotic spindle determines lineage fate and position in the mouse embryo
Nature Cell Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.