Abstract
X-chromosome inactivation (XCI) involves major reorganization of the X chromosome as it becomes silent and heterochromatic. During female mammalian development, XCI is triggered by upregulation of the non-coding Xist RNA from one of the two X chromosomes. Xist coats the chromosome in cis and induces silencing of almost all genes via its A-repeat region1,2, although some genes (constitutive escapees) avoid silencing in most cell types, and others (facultative escapees) escape XCI only in specific contexts3. A role for Xist in organizing the inactive X (Xi) chromosome has been proposed4,5,6. Recent chromosome conformation capture approaches have revealed global loss of local structure on the Xi chromosome and formation of large mega-domains, separated by a region containing the DXZ4 macrosatellite7,8,9,10. However, the molecular architecture of the Xi chromosome, in both the silent and expressed regions, remains unclear. Here we investigate the structure, chromatin accessibility and expression status of the mouse Xi chromosome in highly polymorphic clonal neural progenitors (NPCs) and embryonic stem cells. We demonstrate a crucial role for Xist and the DXZ4-containing boundary in shaping Xi chromosome structure using allele-specific genome-wide chromosome conformation capture (Hi-C) analysis, an assay for transposase-accessible chromatin with high throughput sequencing (ATAC–seq) and RNA sequencing. Deletion of the boundary disrupts mega-domain formation, and induction of Xist RNA initiates formation of the boundary and the loss of DNA accessibility. We also show that in NPCs, the Xi chromosome lacks active/inactive compartments and topologically associating domains (TADs), except around genes that escape XCI. Escapee gene clusters display TAD-like structures and retain DNA accessibility at promoter-proximal and CTCF-binding sites. Furthermore, altered patterns of facultative escape genes in different neural progenitor clones are associated with the presence of different TAD-like structures after XCI. These findings suggest a key role for transcription and CTCF in the formation of TADs in the context of the Xi chromosome in neural progenitors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wutz, A., Rasmussen, T. P. & Jaenisch, R. Chromosomal silencing and localization are mediated by different domains of Xist RNA. Nature Genet. 30, 167–174 (2002)
Chu, C. et al. Systematic discovery of Xist RNA binding proteins. Cell 161, 404–416 (2015)
Peeters, S. B., Cotton, A. M. & Brown, C. J. Variable escape from X-chromosome inactivation: identifying factors that tip the scales towards expression. BioEssays 36, 746–756 (2014)
Chaumeil, J., Le Baccon, P., Wutz, A. & Heard, E. A novel role for Xist RNA in the formation of a repressive nuclear compartment into which genes are recruited when silenced. Genes Dev. 20, 2223–2237 (2006)
Splinter, E. et al. The inactive X chromosome adopts a unique three-dimensional conformation that is dependent on Xist RNA. Genes Dev. 25, 1371–1383 (2011)
Engreitz, J. M. et al. The Xist lncRNA exploits three-dimensional genome architecture to spread across the X chromosome. Science 341, 1237973 (2013)
Rao, S. S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 162, 687–688 (2015)
Deng, X. et al. Bipartite structure of the inactive mouse X chromosome. Genome Biol. 16, 152 (2015)
Minajigi, A. et al. Chromosomes. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation. Science 349, aab2276 (2015)
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012)
Giacalone, J., Friedes, J. & Francke, U. A novel GC-rich human macrosatellite VNTR in Xq24 is differentially methylated on active and inactive X chromosomes. Nature Genet. 1, 137–143 (1992)
Chadwick, B. P. DXZ4 chromatin adopts an opposing conformation to that of the surrounding chromosome and acquires a novel inactive X-specific role involving CTCF and antisense transcripts. Genome Res. 18, 1259–1269 (2008)
Teller, K. et al. A top-down analysis of Xa- and Xi-territories reveals differences of higher order structure at ≥ 20 Mb genomic length scales. Nucleus 2, 465–477 (2011)
Naughton, C., Sproul, D., Hamilton, C. & Gilbert, N. Analysis of active and inactive X chromosome architecture reveals the independent organization of 30 nm and large-scale chromatin structures. Mol. Cell 40, 397–409 (2010)
Eils, R. et al. Three-dimensional reconstruction of painted human interphase chromosomes: active and inactive X chromosome territories have similar volumes but differ in shape and surface structure. J. Cell Biol. 135, 1427–1440 (1996)
Gendrel, A.-V. et al. Developmental dynamics and disease potential of random monoallelic gene expression. Dev. Cell 28, 366–380 (2014)
Zuin, J. et al. Cohesin and CTCF differentially affect chromatin architecture and gene expression in human cells. Proc. Natl Acad. Sci. USA 111, 996–1001 (2014)
Sofueva, S. et al. Cohesin-mediated interactions organize chromosomal domain architecture. EMBO J. 32, 3119–3129 (2013)
Vietri Rudan, M. et al. Comparative Hi-C reveals that CTCF underlies evolution of chromosomal domain architecture. Cell Reports 10, 1297–1309 (2015)
Schulz, E. G. et al. The two active X chromosomes in female ESCs block exit from the pluripotent state by modulating the ESC signaling network. Cell Stem Cell 14, 203–216 (2014)
Yang, F. et al. The lncRNA Firre anchors the inactive X chromosome to the nucleolus by binding CTCF and maintains H3K27me3 methylation. Genome Biol. 16, 52 (2015)
Wutz, A. & Jaenisch, R. A shift from reversible to irreversible X inactivation is triggered during ES cell differentiation. Mol. Cell 5, 695–705 (2000)
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012)
Corbel, C., Diabangouaya, P., Gendrel, A.-V., Chow, J. C. & Heard, E. Unusual chromatin status and organization of the inactive X chromosome in murine trophoblast giant cells. Development 140, 861–872 (2013)
Patrat, C. et al. Dynamic changes in paternal X-chromosome activity during imprinted X-chromosome inactivation in mice. Proc. Natl Acad. Sci. USA 106, 5198–5203 (2009)
Horakova, A. H. et al. The mouse DXZ4 homolog retains Ctcf binding and proximity to Pls3 despite substantial organizational differences compared to the primate macrosatellite. Genome Biol. 13, R70 (2012)
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009)
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013)
Lajoie, B. R., Dekker, J. & Kaplan, N. The Hitchhiker’s guide to Hi-C analysis: practical guidelines. Methods 72, 65–75 (2015)
Selvaraj, S., R Dixon, J., Bansal, V. & Ren, B. Whole-genome haplotype reconstruction using proximity-ligation and shotgun sequencing. Nat. Biotechnol. 31, 1111–1118 (2013)
Imakaev, M. et al. Iterative correction of Hi-C data reveals hallmarks of chromosome organization. Nat. Methods 9, 999–1003 (2012)
Crane, E. et al. Condensin-driven remodelling of X chromosome topology during dosage compensation. Nature 523, 240–244 (2015)
Chaumeil, J., Augui, S., Chow, J. C. & Heard, E. Combined immunofluorescence, RNA fluorescent in situ hybridization, and DNA fluorescent in situ hybridization to study chromatin changes, transcriptional activity, nuclear organization, and X-chromosome inactivation. The Nucleus 463, 297–308 (2008)
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013)
Eckersley-Maslin, M. A. et al. Random monoallelic gene expression increases upon embryonic stem cell differentiation. Dev. Cell 28, 351–365 (2014)
Berletch, J. B. et al. Escape from X inactivation varies in mouse tissues. PLoS Genet. 11, e1005079 (2015)
Martynoga, B. et al. Epigenomic enhancer annotation reveals a key role for NFIX in neural stem cell quiescence. Genes Dev. 27, 1769–1786 (2013)
McHugh, C. A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015)
Acknowledgements
We thank members of the Heard, Dekker, and Chang laboratories for their help and critical insights; PICT-IBiSA@BDD (UMR3215/U934) Imaging facility of the Institut Curie. L.G. would like to thank L. Mirny for discussing gyration tensor analysis. Supported by grants from the National Institutes of Health (P50-HG007735) and Scleroderma Research Foundation (to H.Y.C.), from the National Human Genome Research Institute (R01 HG003143) and the National Institutes of Health Common Fund, National Institute of Diabetes and Digestive and Kidney Diseases (U54 DK107980) to J.D., the Human Frontier Science Program to N.K., an EMBO Fellowship to L.G., an ERC Advanced Investigator award (ERC-2014-AdG no. 671027), EU FP7 grants SYBOSS (EU 7th Framework G.A. no. 242129) and MODHEP (EU 7th Framework G.A. no. 259743), La Ligue, Fondation de France, Labex DEEP (ANR-11-LBX-0044) part of the IDEX Idex PSL (ANR-10-IDEX-0001-02 PSL) and ABS4NGS (ANR-11-BINF-0001) to E.H. J.D. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
E.H. and J.D. conceived the original strategy. For Hi-C, M.A. and L.G. prepared the ES cell and NPC samples, Y.Z. performed the Hi-C experiments; for ATAC–seq, H.Y.C., E.H. and J.D. designed the experiments; A.C.C. and J.X. prepared the samples and performed the experiments; for RNA-seq, M.A. and L.G. prepared the samples and performed the experiments and C.J.C. analysed the data. Integrated analysis of Hi-C sequencing, RNA-seq and ATAC–seq data was performed by B.R.L., J.X., L.G. and A.C.C., with assistance from C.J.C. and N.K. and input from J.D.; L.G. and E.H. designed the FISH experiments, L.G. and M.A. performed FISH experiment and L.G. analysed data. M.A., L.G. and E.H. designed the NPC and DXZ4 mutant strategy, M.A. and L.G. performed the experiments and analysed them. L.G., B.R.L., A.C.C., E.H. and J.D. wrote the manuscript with input from H.Y.C.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Description of allele-specific Hi-C and ATAC–seq.
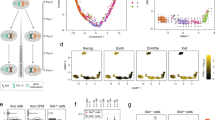
a, Schematic of hybrid mouse strains used for all experiments. b, Top, scheme outlining differentiation of ES cells to NPCs and picking of clones. Bottom, scheme outlining CRISPR deletion of the mega-domain boundary in ES cells, differentiation to NPCs and the picking of clones. c, Schematic of Hi-C library generation. d, Schematic of the Hi-C alignment strategy. Paired-end reads are aligned to a ‘diploid’ genome consisting of 22 chromosomes from Cast, and 22 chromosomes from 129 (1–19 X, Y, M). The interaction row shows all possible paired-end read combinations between the 129, Cast and ambiguous (AMB) genomes. e, Schematic showing the re-assignment of a subset of ‘cis’ interactions. Paired-end reads in which one side uniquely aligned to an allele and the other side aligned equally to both alleles (AMB), were re-classified as allelic reads, only if both reads aligned to the same chromosome (cis). f, Cartoon explaining the re-assignment of 129:amb or cast:amb cis interactions. g, Scheme for ATAC–seq library preparation. Cells are lysed followed by incubation with adaptor-loaded hyperactive Tn5 transposase. The transposase integrates into accessible DNA, and these fragments are then directly amplified and sequenced. h, Scheme for allele-specific ATAC–seq data analysis. i, SNP-mapping simulation. For each SNP location on the X chromosome, all overlapping 50 bp reads were extracted (50 total) for each of the 129 and Cast alleles. All reads were then processed through the Hi-C and ATAC–seq mapping pipelines described in the Methods to measure assignment accuracy. Results are shown in the table.
Extended Data Figure 2 Structure of the Xa chromosome in ES cells, compartment analysis and characterization of interactions between genes that escape XCI.
a, Allele-specific Hi-C contact maps for X chromosome in ES cells at 500-kb resolution (top), and for a ~40-Mb region centred around the DXZ4-containing locus at 40-kb resolution (bottom). The insulation score is plotted at the bottom of each 40-kb heat map as in Fig. 1a. Purple shaded areas indicate the IQR of insulation scores along the chromosome. b, Zoomed in view of three regions showing Hi-C interactions, RNA-seq and ATAC–seq signal on the Cast and 129 Xa chromosomes in ES cells. c, Compartment profiles of X chromosome in ES cells and NPCs. The first eigenvector (PC1) of each allele-specific Hi-C contact map, obtained with principal component analysis, is shown, together with the difference in chromosome-wide insulation score between the 129 and Cast allele. A/B-compartments are evident in ES cells and NPCs along both Xa (red and blue signal), whereas the first eigenvector corresponds to the two mega-domains for the Xi chromosome in NPCs. In ES cells, both Xa chromosomes display comparable insulation profiles (difference is close to 0 along the chromosome), whereas in NPCs large differences are observed (difference in insulation fluctuates along the chromosome). Grey areas indicate regions with low SNP density that were excluded from analysis. d, Same plots as in Fig. 2e for the same set of genes that escape XCI in NPCs, but using Hi-C data obtained in ES cells. e, As in d, where the interactions between loci that escape XCI on the wild-type Xi chromosome were divided in three different groups, according to whether pairs of loci belong to the same or different mega-domains. Interactions across the mega-domain boundary only occur on the wild-type Xi chromosome and are lost on the ΔFT Xi chromosome concomitant with loss of transcription of constitutive escapees.
Extended Data Figure 3 Comparison of allele-specific Hi-C maps of chromosomes X and 13 in ES cells, NPCs and boundary-mutant ΔFT NPCs.
a, Hi-C data, insulation scores, and the difference in insulation scores (129 Cast) are shown for ES cells, NPCs and mutant ΔFT NPCs for both alleles (Cast and 129) for the X chromosome. Large dips in the insulation vector are found at TAD boundaries. Peaks in the insulation vector are found towards the centre of each TAD. The insulation difference plot highlights areas of differential TAD structure between the alleles (many differences along the X chromosome in NPCs as compared to the allelic differences along autosomes, see b). b, Same as a for chromosome 13. The insulation difference plot highlights areas of differential TAD structure between the alleles (rare).
Extended Data Figure 4 Quantitative analysis of 3D DNA FISH on the Xa and Xi chromosomes.
a, Top left, scheme of the procedure used to quantify the Pearson correlation. A background is generated for each xy plane in a three-dimensional z-stack by morphological opening the image with a circle of 5 pixels in radius, and subtracted from it. Pearson correlation between red and green pixel intensities is measured inside a fixed-size region of 40 × 40 × 20 pixels (5.16 × 5.16 × 4 μm) centred on each FISH signal. To demonstrate that background subtraction does not affect the measured correlations, we show here a line-scan of 10 μm across a typical DNA FISH signal (top right). The shape of the signals along the line scan, as well as their relative intensities, is not affected by background subtraction (bottom). Pre- and post- refer to line scans taken on images before and after background subtraction, respectively. b, In more than 80% of nuclei in NPCs, Pearson correlations are higher on the Xi than on the Xa chromosome. Shown is NPC clone C2 (the same where Hi-C was performed). c, Same quantification as in Fig. 1c (and b) for an independent NPC clone (E1) where the active X is on the 129 allele and the inactive X chromosome on the Cast, and in astrocytes derived from NPC clone C2. n denotes the number of cells analysed in DNA FISH. Centre lines represent medians; crosses denote experimental points that were considered as outliers (larger than (q3 + 1.5 × (q3 − q1) or smaller than q1 − w(q3 − q1), where q1 and q3 are the twenty-fifth and seventy-fifth percentiles of the data distribution, respectively). d, Scheme of the gyration tensor based analysis of FISH volumes (see Methods). e, Left, gyration radii of DNA FISH signals from probes a, b and c. Probe b was used in combination with both probes a and c separately in two independent experiments. Statistical significance was assessed by Wilcoxon’s rank sum test (*P < 0.05, **P < 1 × 10−5). The mean gyration radii for Xa and Xi chromosome signals are indicated by dotted lines as a guide for the eye. On the Xi chromosome, the mean gyration radius is approximately 6% smaller than on the Xa chromosome corresponding to approximately 20% smaller volume. Right, representative images of probe a, showing smaller size and increased roundness of the Xi chromosome signals. f, Left, scheme of the thresholding-based method for volume quantification. Thirty increasing threshold levels were imposed, starting from the residual grayscale background level surrounding the signal, up to the minimum between the red and green channel grayscale maxima. For each of these thresholds we determined the number of voxels in each channel, where the greyscale intensity was higher than the threshold. Centre, the fraction of cells where the Xa chromosome signal is larger than the Xi chromosome is between 60% and 80% in the entire threshold range. Right, in a wide range of thresholds, the volume of Xa chromosome signals is approximately 25% bigger than Xi chromosome signals. Results are shown here for probes a and b; the same holds for probe c (not shown). n denotes the number of cells analysed in DNA FISH. Centre lines: medians. Boxes: middle 50% of data points. Two biological replicates were analysed for each experiment.
Extended Data Figure 5 Integrative analysis of allele-specific Hi-C, RNA-seq and ATAC–seq.
a, X-chromosome-wide ATAC–seq and RNA-seq in ES cells and NPCs. ATAC shows signal for ambiguous, 129- and Cast-specific reads in ES cells and NPCs. RNA-seq shows total signal as well as expressed gene calls. ATAC–seq shows global loss of chromatin accessibility and expression on the Xi chromosome, except at specific locations that mostly overlap with escape genes. Dotted line denotes mega-domain boundary. Position of constitutive escapees was adapted from ref. 3. b, Escape genes on the Xi chromosome (as determined by RNA-seq) fall within regions with high ATAC–seq signal (Kolmogorov–Smirnov test P < 2.2 × 10−16). c, Pie charts showing the distribution of peaks that escape XCI versus the peaks that are unique to the Xa chromosome. Peaks are classified into those that are promoter-proximal (within 5 kb of TSS) and distal (>5 kb from TSS). Annotations are based on binding sites identified by ChIP–seq37,38. d, Scatter plot showing the relationship between TAD signal (insulation score) and number of expressed genes of the wild-type NPC 129 (Xi) Hi-C data set. The y axis shows the mean insulation score in a 480-kb window with a 40-kb step size. The x axis shows the number of expressed genes in a 480-kb window with a 40-kb step size. The Pearson R value (0.395) is shown above and is calculated on only the middle 90% (red points) of the residuals. Outliers are shown in black (bottom 5% and top 5% of residuals)
Extended Data Figure 6 Analysis of wild-type and boundary-mutant ΔFT NPCs by FISH and RT–PCR followed by pyrosequencing.
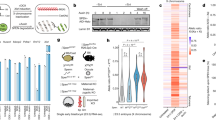
a, Scheme of the strategy used to delete the mega-domain boundary region in ES cells and to derive ΔFT NPCs. b, RNA FISH against constitutive and facultative escapees confirms RNA-seq and ATAC–seq results in the ΔFT NPC clone D9B2. Top, the positions of BAC probes (RP23-328M22 and RP23-436K) are shown relative to the escape genes that they span. Coloured gene names correspond to transcripts that were detected with specific fosmid probes. Bottom, sample RNA FISH images showing that expression of facultative (Mecp2 and BAC probes) but not constitutive (Jarid1c) escapees is lost on the ΔFT Xi chromosome. c, Quantification of the RNA FISH experiment in b. d, RT–PCR followed by pyrosequencing of Xist, two facultative escape genes (Huwe1 and Mecp2) and the constitutive escapee Jarid1c indicate the percentage of expression from the 129 allele (red) or the Cast allele (blue) in 56 wild-type and 17 ΔFT NPC clones. Absence of Mecp2 and Huwe1 escape is observed in a moderately higher proportion of ΔFT than wild-type clones. Absence of Jarid1c escape is never observed.
Extended Data Figure 7 Allele-specific Hi-C analysis of wild-type and boundary-mutant ΔFT NPCs.
a, Hi-C data, insulation scores, and the difference in insulation scores are shown to compare the wild-type Xi chromosome (NPC 129) and the ΔFT Xi chromosome (NPC 129). Top, the Cast allele (Xa chromosome) for both samples. Bottom, the 129 allele (Xi chromosome) for both samples. Large dips in the insulation vector are indicative of TAD boundaries. Peaks in the insulation vector are found towards the centre of each TAD. The insulation difference plot highlights areas of differential TAD structure between the wild-type and ΔFT NPCs. b, Cumulative plots of TAD strength at the wild-type expressed versus the wild-type silenced genes on Cast and 129 chromosomes, for ES cells, wild-type and ΔFT NPCs. Escapee genes on the Xi chromosome (NPC 129, ΔFT NPC 129) show higher insulation scores as compared to silenced genes.
Extended Data Figure 8 3D DNA FISH and integrative ATAC–seq/RNA-seq analysis in ΔFT NPCs.
a, Top left, scheme of the DNA FISH probe sets (a–b: inside the same mega-domain, b–c: across the boundary). Bottom, loci detected by probe set b–c are more interacting in the ΔFT than in the wild-type Xi chromosome both in Hi-C (left) and in 3D-DNA FISH (right), showing loss of mega-domain boundary. *P < 2 × 10−4, **P < 1 × 10−5 (Wilcoxon’s rank sum test corrected with Bonferroni for multiple hypothesis testing). Right, sample RNA/DNA FISH images showing that signals from probe set b–c are more overlapping on the ΔFT Xi than on the wild-type Xi chromosome. Two biological replicates were analysed. b, Chromosome-wide ATAC–seq and RNA-seq signal generated with ambiguous (mm9), 129- and Cast-specific reads in wild-type NPCs and ΔFT NPCs, showing global loss of chromatin accessibility on the ΔFT Xi chromosome except at the X-inactivation centre (Xic) region and constitutive escape genes. Dotted line indicates the position of the mega-domain boundary. c, Zoomed-in view of a region on the ΔFT Xi chromosome encompassing the Xic showing Hi-C interactions, RNA-seq and ATAC–seq signal. ATAC–seq from wild-type NPCs is included for reference (previously shown in Fig. 1).
Extended Data Figure 9 3D DNA FISH and integrative Hi-C/ATAC-seq analysis of inducible Xist ES cell lines.
a, Left, schematic representation of TXY and TXY:ΔA male cell lines, carrying a tetracycline-inducible wild-type and A-repeat mutant Xist, respectively, at the endogenous Xist locus. Centre right, RNA FISH against G6pdx and a group of genes recognized by the RP23-436K BAC (see Extended Data Fig. 6b) showing that expression of X-chromosome-linked genes is lost upon induction of wild-type but not A-repeat mutant Xist in male ES cells. TXY and TXY:ΔA were treated with doxycycline for 48 h. b, Representative images from RNA/DNA FISH experiments in TXY and TXY:ΔA male ES cell lines, showing increased overlap of probes a–b on the Xist-coated X chromosome after 48 h of wild-type, but not ΔA Xist induction. c, Left, schematic representation of TX1072 female ES cells in which Xist expression can be induced via a tetracycline-responsive promoter at one of the two endogenous Xist loci. Center right, RNA/DNA FISH in TX1072 cells treated for 3 days with doxycycline. Probes a–b overlap more on the Xist-coated than on the wild-type X chromosome, whereas signals from b–c show lower overlap and partitioning of the Xist-coated chromosome into two separate domains. *P < 1 × 10−7 (Wilcoxon’s rank sum test corrected with Bonferroni for multiple hypothesis testing). n denotes the number of cells analysed in DNA FISH. Centre lines: medians. Boxes: middle 50% of data points. Two biological replicates were analysed. d, Correlation between changes in Hi-C interactions and ATAC–seq signal following 48-h wild-type Xist induction in male ES cells (TXY cell line). log2(+dox/−dox) was calculated for bins of 40 kb (left), 100 kb (middle), and 500 kb (right). e, Model of mega-domain boundary-mediated control of chromosome folding and facultative escape. Xist coating causes gene silencing and initiates chromosome-wide conformational changes in a manner dependent on its A-repeat region, which result in formation of mega-domains, overall compaction of chromosome folding, and loss of TADs. During differentiation, transient interactions with the mega-domain boundary may occur and may contribute to facultative escape and re-establishment of TADs at facultative escape loci, although the presence of residual TADs at escapee loci on the Xi chromosome is not necessarily due to the mega-domain boundary.
Supplementary information
Supplementary Table
This table contains all relevant RNA-Seq data for all locations (genes) along the X chromosome. The table columns are as follows: 1, xloc; 2, chr; 3, start; 4, end; 5, gene; 6, B129T3__129S1__category; 7, B129T3__129S1__pval; 8, B129T3__129S1__reads; 9, B129T3__129S1__rpkm; 10, B129T3__129S1__status; 11, B129T3__CAST__category; 12, B129T3__CAST__pval; 13, B129T3__CAST__reads; 14, B129T3__CAST__rpkm; 15, B129T3__CAST__status; 16, GEI.72b__129S1__category; 17, GEI.72b__129S1__pval; 18, GEI.72b__129S1__reads; 19, GEI.72b__129S1__rpkm; 20, GEI.72b__129S1__status; 21, GEI.72b__CAST__category; 22, GEI.72b__CAST__pval; 23, GEI.72b__CAST__reads; 24, GEI.72b__CAST__rpkm; 25, GEI.72b__CAST__status; 26, GUR.2d__129S1__category; 27, GUR.2d__129S1__pval; 28, GUR.2d__129S1__reads; 29, GUR.2d__129S1__rpkm; 30, GUR.2d__129S1__status; 31, GUR.2d__CAST__category; 32, GUR.2d__CAST__pval; 33, GUR.2d__CAST__reads; 34, GUR.2d__CAST__rpkm; 35, GUR.2d__CAST__status. xloc is a numerical ID for each gene location. chr is the chromosome. start is the start position of the gene. end is the end position of the gene. (for positions, start > end, not re-oriented by strand) gene is the gene name. The remaining columns are broken down into groups of 5 per sample, per allele. NNNN is the sample name. ESC = GUR.2d; WT NPC = GEI.72b; ΔFT NPC = B129T3 (D9B2). XXXX is the allele, 129S1 for the 129 allele, CAST for the Cast allele. The five columns are: NNNN__XXXX_category, category assignment of expression (bi,mono,biased,na, see ref 20) NNNN__XXXX__pval, p-value of the allelic assignment. NNNN__XXXX__reads, number of allelic reads. NNNN__XXXX__rpkm, RPKM value for the allelic gene. NNNN__XXXX__status, expression status of the gene, expressed or silenced. We defined expressed as ≥ 3 RPKM. (XLSX 5591 kb)
Rights and permissions
About this article
Cite this article
Giorgetti, L., Lajoie, B., Carter, A. et al. Structural organization of the inactive X chromosome in the mouse. Nature 535, 575–579 (2016). https://doi.org/10.1038/nature18589
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature18589
This article is cited by
-
Computational methods for analysing multiscale 3D genome organization
Nature Reviews Genetics (2024)
-
TLR8 escapes X chromosome inactivation in human monocytes and CD4+ T cells
Biology of Sex Differences (2023)
-
Replication dynamics identifies the folding principles of the inactive X chromosome
Nature Structural & Molecular Biology (2023)
-
Determining chromatin architecture with Micro Capture-C
Nature Protocols (2023)
-
LncRNA XIST regulates breast cancer stem cells by activating proinflammatory IL-6/STAT3 signaling
Oncogene (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.