Abstract
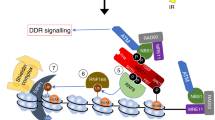
After DNA replication, chromosomal processes including DNA repair and transcription take place in the context of sister chromatids. While cell cycle regulation can guide these processes globally, mechanisms to distinguish pre- and post-replicative states locally remain unknown. Here we reveal that new histones incorporated during DNA replication provide a signature of post-replicative chromatin, read by the human TONSL–MMS22L1,2,3,4 homologous recombination complex. We identify the TONSL ankyrin repeat domain (ARD) as a reader of histone H4 tails unmethylated at K20 (H4K20me0), which are specific to new histones incorporated during DNA replication and mark post-replicative chromatin until the G2/M phase of the cell cycle. Accordingly, TONSL–MMS22L binds new histones H3–H4 both before and after incorporation into nucleosomes, remaining on replicated chromatin until late G2/M. H4K20me0 recognition is required for TONSL–MMS22L binding to chromatin and accumulation at challenged replication forks and DNA lesions. Consequently, TONSL ARD mutants are toxic, compromising genome stability, cell viability and resistance to replication stress. Together, these data reveal a histone-reader-based mechanism for recognizing the post-replicative state, offering a new angle to understand DNA repair with the potential for targeted cancer therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Duro, E. et al. Identification of the MMS22L-TONSL complex that promotes homologous recombination. Mol. Cell 40, 632–644 (2010)
O’Donnell, L. et al. The MMS22L-TONSL complex mediates recovery from replication stress and homologous recombination. Mol. Cell 40, 619–631 (2010)
O’Connell, B. C. et al. A genome-wide camptothecin sensitivity screen identifies a mammalian MMS22L-NFKBIL2 complex required for genomic stability. Mol. Cell 40, 645–657 (2010)
Piwko, W. et al. RNAi-based screening identifies the Mms22L–Nfkbil2 complex as a novel regulator of DNA replication in human cells. EMBO J. 29, 4210–4222 (2010)
Campos, E. I. et al. Analysis of the histone H3.1 interactome: a suitable chaperone for the right event. Mol. Cell 60, 697–709 (2015)
Groth, A. et al. Regulation of replication fork progression through histone supply and demand. Science 318, 1928–1931 (2007)
Huang, H. et al. A unique binding mode enables MCM2 to chaperone histones H3–H4 at replication forks. Nature Struct. Mol. Biol. 22, 618–626 (2015)
Richet, N. et al. Structural insight into how the human helicase subunit MCM2 may act as a histone chaperone together with ASF1 at the replication fork. Nucleic Acids Res. 43, 1905–1917 (2015)
Kalashnikova, A. A., Porter-Goff, M. E., Muthurajan, U. M., Luger, K. & Hansen, J. C. The role of the nucleosome acidic patch in modulating higher order chromatin structure. J. R. Soc. Interface 10, 20121022 (2013)
Collins, R. E. et al. The ankyrin repeats of G9a and GLP histone methyltransferases are mono- and dimethyllysine binding modules. Nature Struct. Mol. Biol. 15, 245–250 (2008)
Jasencakova, Z. et al. Replication stress interferes with histone recycling and predeposition marking of new histones. Mol. Cell 37, 736–743 (2010)
Taipale, M. et al. hMOF histone acetyltransferase is required for histone H4 lysine 16 acetylation in mammalian cells. Mol. Cell. Biol. 25, 6798–6810 (2005)
Rice, J. C. et al. Mitotic-specific methylation of histone H4 Lys 20 follows increased PR-Set7 expression and its localization to mitotic chromosomes. Genes Dev. 16, 2225–2230 (2002)
Pesavento, J. J., Yang, H. & Kelleher, N. L. & Mizzen, C. A. Certain and progressive methylation of histone H4 at lysine 20 during the cell cycle. Mol. Cell. Biol. 28, 468–486 (2008)
Beck, D. B., Oda, H., Shen, S. S. & Reinberg, D. PR-Set7 and H4K20me1: at the crossroads of genome integrity, cell cycle, chromosome condensation, and transcription. Genes Dev. 26, 325–337 (2012)
Jørgensen, S., Schotta, G. & Sørensen, C. S. Histone H4 lysine 20 methylation: key player in epigenetic regulation of genomic integrity. Nucleic Acids Res. 41, 2797–2806 (2013)
Loyola, A., Bonaldi, T., Roche, D., Imhof, A. & Almouzni, G. PTMs on H3 variants before chromatin assembly potentiate their final epigenetic state. Mol. Cell 24, 309–316 (2006)
Alabert, C. et al. Two distinct modes for propagation of histone PTMs across the cell cycle. Genes Dev. 29, 585–590 (2015)
Alabert, C. et al. Nascent chromatin capture proteomics determines chromatin dynamics during DNA replication and identifies unknown fork components. Nature Cell Biol. 16, 281–293 (2014)
Prasanth, S. G., Méndez, J., Prasanth, K. V. & Stillman, B. Dynamics of pre-replication complex proteins during the cell division cycle. Phil. Trans. R. Soc. B 359, 7–16 (2004)
Takahashi, T. S., Wigley, D. B. & Walter, J. C. Pumps, paradoxes and ploughshares: mechanism of the MCM2–7 DNA helicase. Trends Biochem. Sci. 30, 437–444 (2005)
Seeber, A., Hauer, M. & Gasser, S. M. Nucleosome remodelers in double-strand break repair. Curr. Opin. Genet. Dev. 23, 174–184 (2013)
Burgess, R. J. & Zhang, Z. Histone chaperones in nucleosome assembly and human disease. Nature Struct. Mol. Biol. 20, 14–22 (2013)
Botuyan, M. V. et al. Structural basis for the methylation state-specific recognition of histone H4–K20 by 53BP1 and Crb2 in DNA repair. Cell 127, 1361–1373 (2006)
Panier, S. & Boulton, S. J. Double-strand break repair: 53BP1 comes into focus. Nature Rev. Mol. Cell Biol. 15, 7–18 (2014)
Fox, D., III et al. Crystal structure of the BARD1 ankyrin repeat domain and its functional consequences. J. Biol. Chem. 283, 21179–21186 (2008)
Laufer, M. et al. Structural requirements for the BARD1 tumor suppressor in chromosomal stability and homology-directed DNA repair. J. Biol. Chem. 282, 34325–34333 (2007)
Ishitobi, M. et al. Mutational analysis of BARD1 in familial breast cancer patients in Japan. Cancer Lett. 200, 1–7 (2003)
Iacovoni, J. S. et al. High-resolution profiling of γH2AX around DNA double strand breaks in the mammalian genome. EMBO J. 29, 1446–1457 (2010)
Cejka, P. & Kowalczykowski, S. C. The full-length Saccharomyces cerevisiae Sgs1 protein is a vigorous DNA helicase that preferentially unwinds holliday junctions. J. Biol. Chem. 285, 8290–8301 (2010)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Dyer, P. N. et al. Reconstitution of nucleosome core particles from recombinant histones and DNA. Methods Enzymol. 375, 23–44 (2004)
Bartke, T. et al. Nucleosome-interacting proteins regulated by DNA and histone methylation. Cell 143, 470–484 (2010)
Bodor, D. L., Valente, L. P., Mata, J. F., Black, B. E. & Jansen, L. E. Assembly in G1 phase and long-term stability are unique intrinsic features of CENP-A nucleosomes. Mol. Biol. Cell 24, 923–932 (2013)
Jørgensen, S. et al. The histone methyltransferase SET8 is required for S-phase progression. J. Cell Biol. 179, 1337–1345 (2007)
Mosbech, A., Lukas, C., Bekker-Jensen, S. & Mailand, N. The deubiquitylating enzyme USP44 counteracts the DNA double-strand break response mediated by the RNF8 and RNF168 ubiquitin ligases. J. Biol. Chem. 288, 16579–16587 (2013)
Nakamura, K. et al. Regulation of homologous recombination by RNF20-dependent H2B ubiquitination. Mol. Cell 41, 515–528 (2011)
Jakobsen, J. S. et al. Temporal mapping of CEBPA and CEBPB binding during liver regeneration reveals dynamic occupancy and specific regulatory codes for homeostatic and cell cycle gene batteries. Genome Res. 23, 592–603 (2013)
Aymard, F. et al. Transcriptionally active chromatin recruits homologous recombination at DNA double-strand breaks. Nature Struct. Mol. Biol. 21, 366–374 (2014)
Acknowledgements
We thank the beam staff at the synchrotrons at the Argonne National Laboratory (NE-CAT) for technical assistance. We thank J. Rouse, D. Durocher, G. Legube and C. Storgaard Sørensen for reagents, G. Montoya for assistance with circular dichroism, C. B. Strømme, A. Strandsby, K. Nakamura, S.-b. Lee and M. Hödl for help with experiments, and Y. Antoku for assistance with microscopy. We thank J. Lukas for comments on the manuscript and Z. Jasencakova for illustrations. G.S. was supported by European Commission Marie Curie ITN FP7 ‘aDDRess’. D.J.P. was supported in part by grants from the Leukemia and Lymphoma Society and the STARR foundation. A.G. is an EMBO Young Investigator and her research is supported by the European Research Council (ERC StG, no. 281765), the Danish National Research Foundation to the Center for Epigenetics (DNRF82), the Danish Cancer Society, the Danish Medical Research Council, the Novo Nordisk Foundation and the Lundbeck Foundation. A.I. is supported by the European Commission FP7 Network of Excellence EpiGeneSys (project 257082), the DFG Excellence Clusters CIPSM and SyNergy, as well as the DFG Collaborative Research Center 1064 (projects A3 and Z3). T.B. is supported by the Medical Research Council and the European Research Council (ERC StG, no. 309952).
Author information
Authors and Affiliations
Contributions
G.S. and A.G. conceived and led the functional studies. H.H conceived and led the generation of cassette to crystallize the complex, and H.H. solved the structure and performed the ITC under the supervision of D.J.P. C.M.H. performed peptide pull-downs with ARD and recombinant nucleosomes. C.A. performed SET8 experiments and NCC. S.B.-J. and N.M. analysed recruitment to laser-induced DNA damage. N.R.-G. prepared histones for mass spectrometry and performed ChIP analysis, I.F. analysed histone modifications by mass spectrometry under the supervision of A.I. B.M.F and T.B. prepared modified recombinant nucleosomes. L.M. and P.C. prepared recombinant TONSL–MMS22L. G.S., H.H., D.J.P. and A.G. wrote the manuscript and all authors commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
G.S., H.H., C.M.H., D.J.P. and A.G. are inventors on a filed patent application covering the discoveries presented in this manuscript.
Additional information
Reviewer Information Nature thanks T. Kutateladze and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 TONSL binding to histones in vivo and in vitro.
a, Histones bridge the interaction between TONSL–MMS22L and MCM2 in cell extracts as shown by co-immunoprecipitation of Flag–HA–MCM2 wild type or histone-binding mutant (Y81A, Y90A)7. U-2-OS cell inducible for Flag–HA–MCM2 wild type or Y81A, Y90A7 were induced for 24 h before immunoprecipitation with Flag antibodies (one representative experiment out of two is shown). b, Immunoprecipitation of GFP–TONSL from solubilized chromatin of HeLa cells transiently transfected with GFP–TONSL plasmid, showing that TONSL associates with nucleosomal histones H3 and H2B (one representative experiment out of two is shown). c, Domain structure of TONSL1,2,3,4. LRR, leucine-rich repeats; TPR, tetratricopeptide repeats; UBL, ubiquitin-like domain. d, Pull-down of GST–ARD with recombinant histones H3–H4 tetramers. e, f, Pull-down of in vitro-translated full-length TONSL with recombinant histones H3–H4 tetramers (e) or H2A–H2B dimers (f) coupled to NHS-activated sepharose beads (one representative experiment out of three (e) and two (f) is shown). ASF1a wild type and histone-binding mutant (V94R) were included as controls. g, TONSL ARD consists of four ankyrin repeats and uses its elongated concave surface to target the H4 tail spanning residues 12 to 23.
Extended Data Figure 2 Models and sequence alignment of TONSL ARD.
a, Pull-down assay of recombinant ARD with GST–H3 tail (amino acids 1–59) and GST–H4 tail (amino acids 1–31). b, Modelling of TONSL ARD on the co-chaperone structure of MCM2 HBD and ASF1 in complex with an H3–H4 dimer. When comparing the structure of the TONSL-ARD–MCM2-HBD–H3–H4 tetramer complex with our previous structure of the MCM2-HBD–H3–H4-dimer–ASF1 complex7 (Protein Data Bank accession 5BNX), the common parts of both structures superimposed well with a small root mean squared deviation (r.m.s.d.) of 0.44 Å. A model of the quinary complex composed of one molecule of each protein, TONSL ARD, MCM2 HBD, ASF1, H3 and H4, was made after superposition. This model shows that TONSL ARD, MCM2 HBD and ASF1 could simultaneously bind an H3–H4 dimer without steric clash. c, Model of TONSL ARD on the structure of the nucleosome. The model was generated by a direct superposition of the H3–H4 tetramer in the structure of the TONSL ARD–MCM2 HBD–H3–H4 tetramer complex onto the H3–H4 tetramer in the nucleosome structure (Protein Data Bank accession 3AV2). There was no adjustment in the conformation of the model and no steric clash in the model. The MCM2 HBD molecules were omitted from the model for clarity. d, Alignment of TONSL ARD (512–692) sequences from Homo sapiens, Mus musculus, Xenopus laevis and Danio rerio. The secondary structures of human TONSL ARD are showed on top of the sequence alignment. Asterisks indicate the highly conserved residues that constitute the H4 tail-binding surface of TONSL ARD and the three strictly conserved acidic residues forming hydrogen bonds with the key residue H4 Lys20 are highlighted with red asterisks.
Extended Data Figure 3 Interaction details of TONSL ARD and GST pull-downs.
a, b, Molecular details of the interactions of TONSL ARD with H4 tail region residues 12–15 (a) and residues 21–23 (b). The Lys12-Gly13-Gly14-Ala15 segment of H4 is positioned within a narrow surface channel of the TONSL ARD scaffold. The intermolecular contacts spanning the Lys12-Gly13-Gly14-Ala15 segment of H4 include hydrophobic interactions between residues Gly13, Gly14 and Ala15 of H4 and residues Asn507, Cys508, Trp641, Tyr645 and Leu649 of ARD, as well as hydrogen bonds between the main-chain O of H4 Gly14 and Nε1 of ARD Trp641, and between the main-chain N of H4 Ala15 and Oη of ARD Tyr645 (a; Fig. 1c). The main-chain O of H4 Lys16 hydrogen bonds with the Nδ2 of ARD Asn571, while the side chain of H4 Lys16 forms contacts with ARD Asn607 and electrostatic interactions with the side chain of ARD Glu597 (Fig. 1c). The side chain of H4 Arg17 stacks over the side chains of ARD Tyr572 and Cys608, while its Nη1 atom forms two hydrogen bonds with main-chain O and Oδ1 of ARD Asn571 (Fig. 1c, e). The side chain of H4 H18 penetrates into a pocket lined by four strictly conserved residues (Trp563, Glu568, Asn571 and Asp604) and is positioned over His567 of ARD (Fig. 1c, f). The side chain of H4 His18 is stacked between Trp563 and Asn571 and forms hydrogen bonds to Glu568 and Asp604 of ARD (Fig. 1f). The main-chain O of H4 Arg19 forms a hydrogen bond with Nε1 of Trp563 and its side chain forms contacts with Cys561 and Gly595 of ARD (Fig. 1c). Interactions with the key residue H4 Lys20 are described in the text (Fig. 1g). The intermolecular contacts spanning the Val21-Leu22-Arg23 segment of H4 include contacts between side chains of H4 Val21 with Tyr560 and Cys561 of ARD (b), while H4 Leu22 interacts with Asp527 and Met528 of ARD. The main-chain N of H4 Arg23 forms a hydrogen bond with the main-chain O of Asp527 of ARD, while the side chain packs against the side chain of Tyr560 of ARD (b). c, Pull-down of recombinant histones H3–H4 with GST–TONSL ARD wild type or indicated mutants. d, Pull-down of pre-purified MCM2 HBD–H3–H4 tetramer complex with GST–TONSL ARD wild type or indicated mutants. e, Circular dichroism analysis of TONSL ARD wild type and the indicated ARD mutants.
Extended Data Figure 4 Structural comparison of the ARDs of TONSL and GLP.
a, b, Representative view of the TONSL ARD with histone H4 tail (a; this work), and crystal structure of the GLP ARD in complex with histone H3 tail dimethylated at Lys9 (b; ref. 10). Both TONSL ARD and GLP ARD use the concave surface to bind their cognate target H4 tail and H3 tail, respectively. TONSL ARD recognizes H4K20me0 mainly through three strong hydrogen bonds with acidic residues Glu530, Asp559 and Glu568, while GLP ARD recognizes H3K9me2 mainly through an aromatic cage forming by residues Trp839, Trp844, Glu847 and Trp877. c, ITC analysis of TONSL ARD binding to H3K9me1 peptide. d, ITC analysis of TONSL acidic stretch and ARD (amino acids 450–692) with H3K9me1 (amino acids 1–21) and H4 (amino acids 9–25) peptides.
Extended Data Figure 5 Effect of SET8 and MOF depletion on TONSL chromatin binding.
a, Immunoprecipitation of GFP–TONSL from solubilized chromatin of GFP–TONSL U-2-OS cells (one representative experiment out of two is shown). Same exposures are shown for input and immunoprecipitation western blots of H3 and H4K16ac. b, TONSL ARD preference for H4K16ac could be mediated by I599 through hydrophobic association with the K16 acetyl group as I599E ARD mutation preferentially reduces binding to H4K16ac peptides as compared to the unmodified H4 tail. Left, pull-down of GFP–TONSL from cell extracts with biotinylated H4 tail peptides. Right, quantification of the western blot, GFP–TONSL binding to the H4K16ac peptide is shown relative to the unmodified peptide. Means with individual data points are shown (n = 2). c, High-content quantitative imaging of TONSL in pre-extracted U-2-OS cells. Plots show total chromatin-bound TONSL and DAPI intensities in cells treated with control or TONSL siRNA, confirming the specificity of TONSL antibody staining. Each dot represents one nucleus. d–f, Analysis of TONSL chromatin-binding in MOF-depleted (d), SET8-depleted (e) and ionizing radiation (IR)-treated cells (f). Chromatin-bound TONSL was quantified by high content imaging of pre-extracted U-2-OS cells stained for endogenous TONSL. Mean TONSL intensity is shown. AU, arbitrary units. d, e, Knockdown efficiency and expected effect on histone modification were confirmed by western blotting (representative of two experiments). e, f, G1 cells were defined by gating on DAPI and EdU intensity. f, TONSL is not recruited to DNA damage in G1 cells, supporting that TONSL accumulation in SET8-depleted cells is due to lack of H4K20me1 and not DNA damage. Cells were irradiated (1.5 Gy) and analysed 1.5 h later (representative of two experiments). d, f, Error bars indicate s.d.; d, from left, n = 4,920, 2,341, 3,608, 2,917; f, n = 382 (−IR), 523(+IR).
Extended Data Figure 6 TONSL binding to chromatin during the cell cycle.
a, b, H4K20 methylation levels on new and old histones analysed by NCC-pulse-SILAC (data are extracted from ref. 18). Cells grown in light SILAC medium were released into S phase in heavy medium and pulsed with b-dUTP. Chromatin was fixed, sonicated and b-dUTP-labelled fragments isolated on streptavidine beads by NCC. Histones were isolated and analysed by mass spectrometry for modifications on new (heavy) and old (light) histones. For clarity a 24 h (G1/S) chase time point is included. Error bars indicate s.d.; n = 9 (S), 3 (S/G2, M), 5 (G1), 3 (G1/S). Data for M (old histones) is shown as the mean of n = 2, as light peptides were not detected in one of the three biological replicates. c, H4K20 methylation levels measured by mass spectrometry in synchronized TIG3 fibroblasts. d, Plot of mean EdU and total DAPI intensities from TIG3 fibroblasts as in Fig. 3b, with the intensity of chromatin-bound TONSL shown in the third dimension as a colour gradient. AU, arbitrary units. Each dot represents one nucleus. Note that a population of G2 cells (EdU negative) retain TONSL on chromatin. e, High-content quantitative imaging of pre-extracted U-2-OS cells stained for EdU and TONSL analysed as in Fig. 3b. f, Analysis of TONSL chromatin binding by cellular fractionation. U-2-OS cells released from a nocodazole block were followed by fluorescence-activated cell sorting (FACS) analysis of DNA content and analysed by western blotting of soluble (CSK-Triton extracted) and chromatin (pellet) fractions (representative of two experiments).
Extended Data Figure 7 Analysis of GFP–TONSL localization.
a, Colocalization analysis of chromatin-bound GFP–TONSL with MCM2 analysed by deconvolution microscopy and measurement of Pearson coefficient in single cells. Error bars indicate s.d., n = 13 from two independent experiments. Representative image, Fig. 3c. b, c, Representative images for the analysis shown in Fig. 3d. Cells were either pulsed with EdU (40 μM) for 15 min (b) or synchronized in G1/S and released into S phase in the continuous presence of EdU (5 μM) (c). Images are representative of b: n = 9 (very early), 16 (early/mid), 10 (mid/late); c: 9 (very early), 27 (early/mid), 36 (mid/late). Scale bar, 5 μm. b, EdU and MCM2 staining was used to determine the cell cycle state in asynchronous (asyn) cell populations. c, Progression through S phase was followed by FACS analysis of DNA content. d, Chromatin-binding of GFP–TONSL analysed by cellular fractionation in inducible U-2-OS cells as quantified in Fig. 3e. C, chromatin; S, soluble. e, f, Chromatin-binding analysis as in Fig. 3f. U-2-OS cells conditional for GFP–TONSL ARD wild type (WT) and mutant were directly fixed or pre-extracted to remove soluble proteins. Data are representative of three (e) and two (f) experiments, fields of cells in e are representative of (from left) n = 16, 18, 17 and 17 images. Scale bar, 20 μm. g, Asynchronous U-2-OS cells conditional for GFP–TONSL were pulsed with 40 μM EdU for 15 min and soluble proteins were extracted. Representative images of EdU-positive cells are shown (n = 30 for wild type and N571A), for the specific patterns of TONSL wild type see Fig. 3d. Scale bar, 5 μm.
Extended Data Figure 8 TONSL–MMS22L recruitment to damaged DNA.
a, Left, ChIP-qPCR analysis of GFP–TONSL recruitment to site-specific DSBs induced by AsiSI, as shown in Fig. 4d but with additional controls. Note that the colours have been changed for clarity. Mean of technical duplicates is shown. Right, dot plot illustrating the relative enrichment of GFP–TONSL wild type (WT) and N571A obtained in four independent ChIPs performed on two biologically independent chromatin preparations. Each experiment was normalized to GFP–TONSL wild-type enrichment at DSB-I_80bp. Mean is shown with two-sided Mann–Whitney test; ***P < 0.001; not significant, P > 0.05; n = 24. Two-sided Mann–Whitney analysis of individual experiments gave similar results. b, U-2-OS cells conditional for GFP–TONSL were laser microirradiated. 53BP1 and cyclin B staining was used as markers of DNA damage and cells in S/G2 phase, respectively. Representative of three experiments as quantified Fig. 4e. Filled arrowheads indicate GFP–TONSL recruitment; open arrowheads indicate no recruitment. Scale bars, 10 μm c, U-2-OS cells transiently transfected with GFP–TONSL wild type or the indicated mutants were laser microirradiated and processed for γH2A.X immunofluorescence. Representative cells are shown (n = 200 cells per condition from two independent experiments). d, U-2-OS cells conditional for GFP–TONSL were laser microirradiated. γH2A.X and RPA staining was used as markers of DNA damage and cells undergoing resection in S/G2 phase, respectively. The percentage of GFP–TONSL cells with recruitment to RPA-positive (+) and RPA-negative (−) laser tracks is indicated. Data are representative of two independent experiments, a total of 118 cells were counted. e, Top, U-2-OS cells conditional for GFP–TONSL wild type and N571A were laser microirradiated. γH2A.X and EdU staining was used as markers of DNA damage and S phase cells, respectively. Bottom, quantification of GFP–TONSL cells with recruitment to laser tracks. Mean with individual data points are shown (n = 2, a total of 138 (wild type) and 174 (N571A) cells were counted). f, H4K20 methylation levels measured by mass spectrometry in synchronized TIG3 cells as in Extended Data Fig. 6c. Cell were released into S phase for 3 h and treated with hydroxyurea (HU; 3 mM) or CPT (1 μM) for 3 h or left untreated (6 h). Mean with individual data points are shown (n = 2). g, Colony formation in cells treated with control or TONSL siRNA and induced to express GFP–TONSL. As shown in Fig. 4f, but including additional mutants. Two cell concentrations in technical triplicate from two (E568A, D559A) or four (wild type, N571A) biological replicates are shown. h, Representation of the complementation analysis from Fig. 4f in a single panel including both CPT-treated and untreated cells. This illustrates that the toxicity of the TONSL ARD mutant is comparable to CPT treatment of cells expressing wild-type TONSL. i, Analysis of GFP–TONSL and MMS22L by cellular fractionation in cells inducible for GFP–TONSL ARD wild type and mutant. Representative experiment of the quantification shown in Fig. 4i.
Extended Data Figure 9 Similarity of the ARDs in TONSL and BARD1, and protein inputs.
a, Superposition of the structures of TONSL ARD and BARD1 ARD (Protein Data Bank accession 3C5R)26. The main residues involved in TONSL ARD interactions with the H4 tail are compared to the corresponding residues of BARD1 ARD. The two ARDs show highly similar topology and conservation of the histone-binding surface. b, Input material of the experiment in Fig. 1h. c, Input material of the experiment in Fig. 1j. d, Spot assay with biotinylated H4 tail (amino acids 14–33) peptides confirming equal input into pull-down reactions. e, Input material of the experiment in Fig. 4b. f, Input material of the NCC experiment in Fig. 4c. Note that because ARD mutation disrupts chromatin binding in the presence and absence of CPT (Figs 3e, f and 4a), GFP–TONSL N517 levels are low in the input chromatin. The NCC experiment in Fig. 4c supports our microscopy-based data (Fig. 4a) and further shows that there is no local accumulation of the GFP–TONSL ARD mutant at damaged forks that could have been missed in our microscopy-based quantification of total TONSL on chromatin.
Supplementary information
Supplementary Figure
This file contains Supplementary Figure 1, the uncropped scans with size marker indications. (PDF 1581 kb)
Rights and permissions
About this article
Cite this article
Saredi, G., Huang, H., Hammond, C. et al. H4K20me0 marks post-replicative chromatin and recruits the TONSL–MMS22L DNA repair complex. Nature 534, 714–718 (2016). https://doi.org/10.1038/nature18312
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature18312
This article is cited by
-
H3.1K27me1 loss confers Arabidopsis resistance to Geminivirus by sequestering DNA repair proteins onto host genome
Nature Communications (2023)
-
Roles for the methyltransferase SETD8 in DNA damage repair
Clinical Epigenetics (2022)
-
Mechanisms of BRCA1–BARD1 nucleosome recognition and ubiquitylation
Nature (2021)
-
RPA shields inherited DNA lesions for post-mitotic DNA synthesis
Nature Communications (2021)
-
Genome-wide and sister chromatid-resolved profiling of protein occupancy in replicated chromatin with ChOR-seq and SCAR-seq
Nature Protocols (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.