Abstract
Chemical modifications of RNA have essential roles in a vast range of cellular processes1,2,3. N6-methyladenosine (m6A) is an abundant internal modification in messenger RNA and long non-coding RNA that can be dynamically added and removed by RNA methyltransferases (MTases) and demethylases, respectively2,3,4,5. An MTase complex comprising methyltransferase-like 3 (METTL3) and methyltransferase-like 14 (METTL14) efficiently catalyses methyl group transfer6,7. In contrast to the well-studied DNA MTase8, the exact roles of these two RNA MTases in the complex remain to be elucidated. Here we report the crystal structures of the METTL3–METTL14 heterodimer with MTase domains in the ligand-free, S-adenosyl methionine (AdoMet)-bound and S-adenosyl homocysteine (AdoHcy)-bound states, with resolutions of 1.9, 1.71 and 1.61 Å, respectively. Both METTL3 and METTL14 adopt a class I MTase fold and they interact with each other via an extensive hydrogen bonding network, generating a positively charged groove. Notably, AdoMet was observed in only the METTL3 pocket and not in METTL14. Combined with biochemical analysis, these results suggest that in the m6A MTase complex, METTL3 primarily functions as the catalytic core, while METTL14 serves as an RNA-binding platform, reminiscent of the target recognition domain of DNA N6-adenine MTase9,10. This structural information provides an important framework for the functional investigation of m6A.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Lee, M., Kim, B. & Kim, V. N. Emerging roles of RNA modification: m6A and U-tail. Cell 158, 980–987 (2014)
Meyer, K. D. & Jaffrey, S. R. The dynamic epitranscriptome: N6-methyladenosine and gene expression control. Nature Rev. Mol. Cell Biol. 15, 313–326 (2014)
Fu, Y., Dominissini, D., Rechavi, G. & He, C. Gene expression regulation mediated through reversible m6A RNA methylation. Nature Rev. Genet. 15, 293–306 (2014)
Schwartz, S. Cracking the epitranscriptome. RNA 22, 169–174 (2016)
Liu, N. & Pan, T. N-methyladenosine-encoded epitranscriptomics. Nature Struct. Mol. Biol. 23, 98–102 (2016)
Wang, Y. et al. N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nature Cell Biol. 16, 191–198 (2014)
Liu, J. et al. A METTL3–METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nature Chem. Biol. 10, 93–95 (2014)
Malone, T., Blumenthal, R. M. & Cheng, X. Structure-guided analysis reveals nine sequence motifs conserved among DNA amino-methyltransferases, and suggests a catalytic mechanism for these enzymes. J. Mol. Biol. 253, 618–632 (1995)
Gupta, Y. K., Chan, S. H., Xu, S. Y. & Aggarwal, A. K. Structural basis of asymmetric DNA methylation and ATP-triggered long-range diffusion by EcoP15I. Nat. Commun. 6, 7363 (2015)
Goedecke, K., Pignot, M., Goody, R. S., Scheidig, A. J. & Weinhold, E. Structure of the N6-adenine DNA methyltransferase M.TaqI in complex with DNA and a cofactor analog. Nature Struct. Biol. 8, 121–125 (2001)
Deng, X. et al. Widespread occurrence of N6-methyladenosine in bacterial mRNA. Nucleic Acids Res. 43, 6557–6567 (2015)
Schwartz, S. et al. High-resolution mapping reveals a conserved, widespread, dynamic mRNA methylation program in yeast meiosis. Cell 155, 1409–1421 (2013)
Luo, G. Z. et al. Unique features of the m6A methylome in Arabidopsis thaliana . Nat. Commun. 5, 5630 (2014)
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012)
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012)
Ping, X. L. et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 24, 177–189 (2014)
Fustin, J. M. et al. RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell 155, 793–806 (2013)
Geula, S. et al. Stem cells. m6A mRNA methylation facilitates resolution of naive pluripotency toward differentiation. Science 347, 1002–1006 (2015)
Chen, T. et al. m6A RNA methylation is regulated by microRNAs and promotes reprogramming to pluripotency. Cell Stem Cell 16, 289–301 (2015)
Zhou, J. et al. Dynamic m6A mRNA methylation directs translational control of heat shock response. Nature 526, 591–594 (2015)
Liu, N. et al. N6-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature 518, 560–564 (2015)
Wang, X. et al. N6-methyladenosine modulates messenger RNA translation efficiency. Cell 161, 1388–1399 (2015)
Choi, J. et al. N6-methyladenosine in mRNA disrupts tRNA selection and translation-elongation dynamics. Nature Struct. Mol. Biol. 23, 110–115 (2016)
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014)
Alarcón, C. R., Lee, H., Goodarzi, H., Halberg, N. & Tavazoie, S. F. N6-methyladenosine marks primary microRNAs for processing. Nature 519, 482–485 (2015)
Schwartz, S. et al. Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5′ sites. Cell Rep. 8, 284–296 (2014)
Jia, G. et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nature Chem. Biol. 7, 885–887 (2011)
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013)
Xiao, W. et al. Nuclear m6A reader YTHDC1 regulates mRNA splicing. Mol. Cell 61, 507–519 (2016)
Iyer, L. M., Zhang, D. & Aravind, L. Adenine methylation in eukaryotes: Apprehending the complex evolutionary history and functional potential of an epigenetic modification. Bioessays 38, 27–40 (2016)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Collaborative Computational Project The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Schneider, T. R. & Sheldrick, G. M. Substructure solution with SHELXD. Acta Crystallogr. D 58, 1772–1779 (2002)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
DeLano, W. L. The PyMOL molecular graphics system. http://www.pymol.org (2002)
Acknowledgements
We thank B. Sun (SSRF beamline BL17U), R. Zhang (BL19U1), and N. Li (BL19U2) for on-site assistance; S. Fan for data collection support; and research associates at the Center for Protein Research, Huazhong Agricultural University, for technical support. This work was supported by funds from the Ministry of Science and Technology (grants 2015CB910900 and 2013CB910200), Fok Ying-Tong Education Foundation (grant 151021), the Fundamental Research Funds for the Central Universities (Program No. 2014PY026, No. 2015PY219, and No. 2014JQ001), and Huazhong Agricultural University Scientific & Technological Self-innovation Foundation (Program No. 2013RC013).
Author information
Authors and Affiliations
Contributions
X.W., T.Z. and P.Y. designed all experiments. X.W., J.F. and Y.X. performed protein purification and crystallization. Z.Gu. determined all of the structures. X.W., Z.L., Z.Go., Q.W., D.Z., J.H., C.T., T.Z. and P.Y. performed the biochemical assays. All authors analysed the data and contributed to manuscript preparation. X.W., T.Z. and P.Y. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks M. Helm, W. Versées and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Sequence alignment of human METTL3 and METTL14.
Sequence alignment of Homo sapiens METTL3 (UniProt accession Q86U44) and METTL14 (UniProt accession Q9HCE5). The alignment was generated using the MultAlin and ENDscript programs. Secondary structural elements are shown above. Sequence identity is shown in white letters with a red background, and sequence similarity is shown in red letters. The coloured dots highlight functionally important positions. Residues of METTL3 and METTL14 that are involved in protein interactions are indicated by magenta and green dots, respectively. Cyan dots indicate residues that interact with AdoMet that were analysed by mutagenesis in this study. Blue dots represent residues that compose the RNA binding groove. The dots at the top and bottom of the sequences indicate residues from METTL3 and METTL14, respectively. Phosphoserine is highlighted by a red arrow.
Extended Data Figure 2 The MTase domains of METTL3 and METTL14 adopt the class I MTase fold.
a, Diagram of the METTL3–METTL14 secondary structure profiles. METTL3 (magenta) and METTL14 (green) are boxed with a light teal background and a wheat background, respectively. The MTase domain contains an eight-stranded β-sheet (triangles) flanked by four α-helices and three 310-helices (circles). Structural elements are numbered by their linear order in the sequence. The loops in the front are indicated by black lines, and loops in the back are indicated by black dashed lines. b, Structural comparison of METTL3 and METTL14. Two perpendicular views of superimposed METTL3 and METTL14 coloured magenta and green, respectively. The NHM and CTM of METTL14 are coloured cyan and yellow, respectively. The main differences between the MTase domains of METTL3 and METTL14 are the two gate loops (orange) and the interface loop (blue).
Extended Data Figure 3 Extensive hydrogen network between METTL3 and METTL14.
a, The main interface of the METTL3–METTL14 heterodimer comprises interface 1 (boxed with orange and green rectangles) and interface 2 (boxed with a cyan rectangle), which generate an extensive water-mediated hydrogen network. METTL3 and METTL14 are coloured wheat and silver, respectively. The interface loop of METTL3 (blue) primarily contributes to the heterodimer interaction. b, Details of interfaces 1 and 2. Water is shown as a red ball. Hydrogen bonds are represented by red dashed lines. Residues from METTL3 (magenta) and METTL14 (green) that are involved in interactions are shown as sticks.
Extended Data Figure 4 One AdoMet was located at the AdoMet binding site of METTL3.
a, Lattice packing of AdoMet-bound complex. One AdoMet (green sphere) was coordinated by METTL3 (purple) but not METTL14 (green). The arrow shows the putative AdoMet-binding pocket. b, Stereo views of electron density map of AdoMet binding site of METTL3. 2Fo − Fc electron density (grey) of AdoMet binding site in METTL3, contoured at 1.0σ. AdoMet is show as green balls-and-sticks and surrounding residues in magenta with the DPPW motif (orange). c, Representative 2Fo − Fc electron density (grey) of AdoMet binding site in METTL14, contoured at 1.0σ. The electron density of METTL14 (grey) is clearly visible and the EPPL motif is coloured orange. No additional apparent electron density was observed in the putative AdoMet binding site of METTL14.
Extended Data Figure 5 Mutagenesis analysis of the METTL3–METTL14–AdoMet interaction.
a, Characterization of METTL3–METTL14 mutations affecting MTase activity. The indicated point mutations were introduced into METTL3. Each METTL3 mutant was co-expressed and purified with wild-type METTL14 as a binary complex and used for the MTase and ITC assays. Methylation yields were calculated based on the c.p.m. of the extracted tritium-labelled RNA probe. The c.p.m. of the extracted RNA was measured in a scintillation counter. The data are shown as mean ± s.e.m. from experiments that were independently repeated at least three times. All alanine substitutions resulted in remarkable decreases in activity. b, c, Measurement of the binding affinity between AdoMet and the METTL3–METTL14 complex (wild-type and D377A for METTL3 and D395A for METTL3) using ITC. Individual peaks from titrations were integrated and presented in a Wiseman plot. The first dot was removed from the analysis. The dissociation constant (Kd) and the binding stoichiometry (N) of the wild type were approximately 1.5 μM and 1.15, respectively. The mutants exhibited undetectable AdoMet binding activity.
Extended Data Figure 6 Biochemical analysis of the role of the potential RNA binding groove.
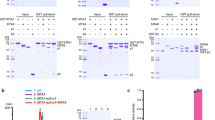
a, RNA binding activity of the METTL3–METTL14 complex revealed by EMSA. The final concentrations of proteins in each set of five lanes (1–5, 6–10, 11–15, 16–20 and 21–25) were 0, 0.19, 0.56, 1.67 and 5 μM, respectively. ‘Well’ indicates the top of native gel. The RNA-bound complex is highlighted by a black asterisk. The wild-type complex binds to the substrate RNA probe weakly (the dissociation constant is approximately 10 μM). All of the mutants showed moderately reduced RNA binding activity. These results suggested that the positively charged groove is involved in RNA interactions. For uncropped gels, see Supplementary Fig. 1. b, Measurement of the binding affinity between AdoMet and the METTL3–METTL14 complex mutants using ITC. These mutations in METTL3 or METTL14 had little effect on AdoMet binding activity.
Extended Data Figure 7 There is little conformational change in overall structure between the AdoMe-bound and AdoHcy-bound states.
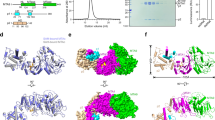
a, Electron density maps of AdoHcy showing 2Fo − Fc electron density (red) of AdoHcy adjacent to the DPPW motif (orange) contoured at 1.0σ. The DPPW motif is shown as sticks. AdoHcy is shown as cyan sticks. b, Structural comparison of AdoHcy (cyan) and AdoMet (green); the electron densities are shown as red and blue meshes, respectively. AdoHcy and AdoMet exhibited nearly identical configurations except for ribose. c, SAXS measurements reveal little structural difference among the ligand-free, AdoMe-bound and AdoHcy-bound states. Superposition of the SAXS curves of ligand-free protein complex (black), and in the presence of AdoMet (red) or AdoHcy (blue).
Extended Data Figure 8 Potential role of METTL14.
a, Structural comparison with the DNA-free (PDB: 2ADM) and DNA-bound (PDB: 1G38) states of M.TaqI. M.TaqI contains the target recognition domain (TRD, green), DNA (orange) and MTase domain (slate). The TRD functions as a scaffold for substrate DNA recognition, and the MTase domain functions as an enzyme. Adenine (magenta) is flipped out and points to the ligand-binding pocket. Black arrows highlight the loop conformational changes, which are similar to those of gate loops 1 and 2 in the METTL3–METTL14 complex. b, Ribbon representation of the DNA-bound state of EcoP15I (PDB: 4ZCF). The TRD (green) of ModA recognizes DNA, while the MTase (slate) of ModB methylates the target adenine. c, The putative AdoMet-binding site of METTL14 (green) is highlighted by a red dashed ellipse. AdoMet coordinated by METTL3 (magenta) is shown as a space-filling representation. The surface electrostatic potential around the putative AdoMet-binding site of METTL14 revealed a negative charge (black dashed ellipse) and suggests a dispensable role for this region in RNA binding. d, Most of the putative AdoMet-binding site residues were conserved between METTL3 (cyan) and METTL14 (yellow). e, Each complex containing alanine substitution mutants of residues in METTL14 (D173 and E192) that correspond to critical residues in METTL3 (D377 and D395) displayed similar methylation activity to the wild type. The average (± s.e.m.) c.p.m. was determined from three independent experiments. f, The complex mutants exhibited similar AdoMet-binding activities to the wild-type complex.
Extended Data Figure 9 Substrate sequence preference of the METTL3–METTL14 complex.
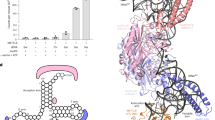
The 20-nucleotide RNA substrate contains four repeats of the consensus sequence GGACU. Each site was substituted by the other three kinds of nucleotide. The average (± s.e.m.) c.p.m. was determined from three independent experiments.
Supplementary information
Supplementary Figure
This file contains Supplementary Figure 1, the uncropped gel images for Extended Data Figure 6a. (PDF 1200 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Feng, J., Xue, Y. et al. Structural basis of N6-adenosine methylation by the METTL3–METTL14 complex. Nature 534, 575–578 (2016). https://doi.org/10.1038/nature18298
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature18298
This article is cited by
-
Intrafamily heterooligomerization as an emerging mechanism of methyltransferase regulation
Epigenetics & Chromatin (2024)
-
The m6A methyltransferase METTL3 drives thyroid cancer progression and lymph node metastasis by targeting LINC00894
Cancer Cell International (2024)
-
RNA m6A methylation and regulatory proteins in pulmonary arterial hypertension
Hypertension Research (2024)
-
Epitranscriptomic modifications in mesenchymal stem cell differentiation: advances, mechanistic insights, and beyond
Cell Death & Differentiation (2024)
-
Advances in brain epitranscriptomics research and translational opportunities
Molecular Psychiatry (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.