Abstract
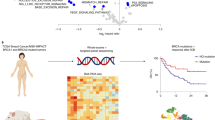
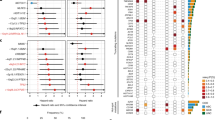
Successful treatment of many patients with advanced cancer using antibodies against programmed cell death 1 (PD-1; also known as PDCD1) and its ligand (PD-L1; also known as CD274) has highlighted the critical importance of PD-1/PD-L1-mediated immune escape in cancer development1,2,3,4,5,6. However, the genetic basis for the immune escape has not been fully elucidated, with the exception of elevated PD-L1 expression by gene amplification and utilization of an ectopic promoter by translocation, as reported in Hodgkin and other B-cell lymphomas, as well as stomach adenocarcinoma6,7,8,9,10. Here we show a unique genetic mechanism of immune escape caused by structural variations (SVs) commonly disrupting the 3′ region of the PD-L1 gene. Widely affecting multiple common human cancer types, including adult T-cell leukaemia/lymphoma (27%), diffuse large B-cell lymphoma (8%), and stomach adenocarcinoma (2%), these SVs invariably lead to a marked elevation of aberrant PD-L1 transcripts that are stabilized by truncation of the 3′-untranslated region (UTR). Disruption of the Pd-l1 3′-UTR in mice enables immune evasion of EG7-OVA tumour cells with elevated Pd-l1 expression in vivo, which is effectively inhibited by Pd-1/Pd-l1 blockade, supporting the role of relevant SVs in clonal selection through immune evasion. Our findings not only unmask a novel regulatory mechanism of PD-L1 expression, but also suggest that PD-L1 3′-UTR disruption could serve as a genetic marker to identify cancers that actively evade anti-tumour immunity through PD-L1 overexpression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Data deposits
Sequencing data have been deposited in the European Genome-phenome Archive (EGA) under accession EGAS00001001296 (https://www.ebi.ac.uk/ega/studies/EGAS00001001296).
References
Iwai, Y. et al. Involvement of PD-L1 on tumor cells in the escape from host immune system and tumor immunotherapy by PD-L1 blockade. Proc. Natl Acad. Sci. USA 99, 12293–12297 (2002)
Topalian, S. L. et al. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N. Engl. J. Med. 366, 2443–2454 (2012)
Brahmer, J. R. et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N. Engl. J. Med. 366, 2455–2465 (2012)
Topalian, S. L., Drake, C. G. & Pardoll, D. M. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer Cell 27, 450–461 (2015)
Page, D. B., Postow, M. A., Callahan, M. K., Allison, J. P. & Wolchok, J. D. Immune modulation in cancer with antibodies. Annu. Rev. Med. 65, 185–202 (2014)
Ansell, S. M. et al. PD-1 blockade with nivolumab in relapsed or refractory Hodgkin’s lymphoma. N. Engl. J. Med. 372, 311–319 (2015)
Green, M. R. et al. Integrative analysis reveals selective 9p24.1 amplification, increased PD-1 ligand expression, and further induction via JAK2 in nodular sclerosing Hodgkin lymphoma and primary mediastinal large B-cell lymphoma. Blood 116, 3268–3277 (2010)
Steidl, C. et al. MHC class II transactivator CIITA is a recurrent gene fusion partner in lymphoid cancers. Nature 471, 377–381 (2011)
Twa, D. D. et al. Genomic rearrangements involving programmed death ligands are recurrent in primary mediastinal large B-cell lymphoma. Blood 123, 2062–2065 (2014)
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513, 202–209 (2014)
Mertens, F., Johansson, B., Fioretos, T. & Mitelman, F. The emerging complexity of gene fusions in cancer. Nat. Rev. Cancer 15, 371–381 (2015)
Garraway, L. A. & Lander, E. S. Lessons from the cancer genome. Cell 153, 17–37 (2013)
Northcott, P. A. et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Nature 511, 428–434 (2014)
Peifer, M. et al. Telomerase activation by genomic rearrangements in high-risk neuroblastoma. Nature 526, 700–704 (2015)
Kataoka, K. et al. Integrated molecular analysis of adult T cell leukemia/lymphoma. Nat. Genet. 47, 1304–1315 (2015)
Beaudoing, E., Freier, S., Wyatt, J. R., Claverie, J. M. & Gautheret, D. Patterns of variant polyadenylation signal usage in human genes. Genome Res. 10, 1001–1010 (2000)
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015)
Hu, Z. et al. Genome-wide profiling of HPV integration in cervical cancer identifies clustered genomic hot spots and a potential microhomology-mediated integration mechanism. Nat. Genet. 47, 158–163 (2015)
Parfenov, M. et al. Characterization of HPV and host genome interactions in primary head and neck cancers. Proc. Natl Acad. Sci. USA 111, 15544–15549 (2014)
Maddalo, D. et al. In vivo engineering of oncogenic chromosomal rearrangements with the CRISPR/Cas9 system. Nature 516, 423–427 (2014)
Matsumoto, M. et al. Defined TLR3-specific adjuvant that induces NK and CTL activation without significant cytokine production in vivo. Nat. Commun. 6, 6280 (2015)
Mayr, C. & Bartel, D. P. Widespread shortening of 3′UTRs by alternative cleavage and polyadenylation activates oncogenes in cancer cells. Cell 138, 673–684 (2009)
Schoenberg, D. R. & Maquat, L. E. Regulation of cytoplasmic mRNA decay. Nat. Rev. Genet. 13, 246–259 (2012)
Gong, A. Y. et al. MicroRNA-513 regulates B7-H1 translation and is involved in IFN-γ-induced B7-H1 expression in cholangiocytes. J. Immunol. 182, 1325–1333 (2009)
Wang, W. et al. A miR-570 binding site polymorphism in the B7-H1 gene is associated with the risk of gastric adenocarcinoma. Hum. Genet. 132, 641–648 (2013)
Wang, X. et al. Tumor suppressor miR-34a targets PD-L1 and functions as a potential immunotherapeutic target in acute myeloid leukemia. Cell. Signal. 27, 443–452 (2015)
Chen, L. et al. Metastasis is regulated via microRNA-200/ZEB1 axis control of tumour cell PD-L1 expression and intratumoral immunosuppression. Nat. Commun. 5, 5241 (2014)
Cortez, M. A. et al. PDL1 Regulation by p53 via miR-34. J. Natl. Cancer Inst. 108, djv303 (2015)
Tsukasaki, K. et al. Definition, prognostic factors, treatment, and response criteria of adult T-cell leukemia-lymphoma: a proposal from an international consensus meeting. J. Clin. Oncol. 27, 453–459 (2009)
Swerdllow, S. et al. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues 4th edn, Vol. 2 (IARC Press, 2008)
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013)
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Brister, J. R., Ako-Adjei, D., Bao, Y. & Blinkova, O. NCBI viral genomes resource. Nucleic Acids Res. 43, D571–D577 (2015)
Kent, W. J. BLAT–the BLAST-like alignment tool. Genome Res. 12, 656–664, 2002)
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-seq. Nat. Methods 5, 621–628 (2008)
Nannya, Y. et al. A robust algorithm for copy number detection using high-density oligonucleotide single nucleotide polymorphism genotyping arrays. Cancer Res. 65, 6071–6079 (2005)
Yamamoto, G. et al. Highly sensitive method for genomewide detection of allelic composition in nonpaired, primary tumor specimens by use of affymetrix single-nucleotide-polymorphism genotyping microarrays. Am. J. Hum. Genet. 81, 114–126 (2007)
Van Loo, P. et al. Allele-specific copy number analysis of tumors. Proc. Natl Acad. Sci. USA 107, 16910–16915 (2010)
Horn, P. A., Topp, M. S., Morris, J. C., Riddell, S. R. & Kiem, H. P. Highly efficient gene transfer into baboon marrow repopulating cells using GALV-pseudotype oncoretroviral vectors produced by human packaging cells. Blood 100, 3960–3967 (2002)
Grillo, G. et al. UTRdb and UTRsite (RELEASE 2010): a collection of sequences and regulatory motifs of the untranslated regions of eukaryotic mRNAs. Nucleic Acids Res. 38, D75–D80 (2010)
Acknowledgements
This work was supported by Grant-in-Aid from the Japan Agency for Medical Research and Development (Practical Research for Innovative Cancer Control (15Ack0106014h0002) and Medical Research and Development Programs Focused on Technology Transfer (15im0210102h0001)), Grant-in-Aid for Scientific Research (KAKENHI 22134006, 15H05909, 25250020), and National Cancer Center Research and Development Funds (26-A-6). We thank M. Sago, M. Nakamura and S. Baba for technical assistance, and R. Velaga for English editing. The supercomputing resources were provided by the Human Genome Center, the Institute of Medical Science, the University of Tokyo. This research also used computational resources of the K computer provided by the RIKEN Advanced Institute for Computational Science through the HPCI System Research project (hp140230, hp160219, and hp150232). The results shown here are partly based on data generated by the TCGA Research Network (http://cancergenome.nih.gov/).
Author information
Authors and Affiliations
Contributions
K.Kataoka, Y.S., H.T., K.C., S.I., and S.Miyano performed sequencing data analyses. H.S., T.Y., Y.Totoki, H.N., N.H. and T.Shibata assisted sequencing data analyses. K.Kataoka, Y.N., Y.W., N.K., K.Y., M.S. and K.Kashiwase performed sequencing experiments. K.Kataoka, S.N., T.M., K.M., N.M., H.K., and Y.A. performed functional assays. S.Mizuno. and S.T. designed sgRNAs. Y.Takeda, M.M., and T.Seya performed in vivo experiments. S.S. and K.T. performed IHC assay. A.K., H.I., Y.I., W.M., K.Shide, Y.K., T.H., T.K., K.I., A.T.-K., Y.M., and K.Shimoda collected specimens. K.Kataoka, Y.S., and S.O. generated figures and tables and wrote the manuscript. S.O. led the entire project. All authors participated in discussions and interpretation of the data and results.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer information Nature thanks J. Cools, M. Meyerson and A. Ribas for their contribution to the peer review of this work
Extended data figures and tables
Extended Data Figure 1 PD-L1 SVs affecting 3′-UTR in ATL.
a, Diagram showing numbers of ATL samples investigated by WGS/RNA-seq. b, Genome-wide distribution of SV (without deletion) breakpoints in 49 ATL samples, showing a prominent peak at the PD-L1 locus. c, Summary of different types of SVs commonly affecting 3′ region of the PD-L1 gene are shown by indicated colours. d, 3′-truncated PD-L1 mRNA transcripts observed in ATL cases. RNA-seq data are visualized by IGV for ATL samples with or without PD-L1 SVs. Aberrant PD-L1 transcripts are shown in red.
Extended Data Figure 2 Fusion genes involving PD-L1 and non-coding sequences identified in ATL.
a, ATL020 fused sequence: the sequences derived from PD-L1 exons 1–6 and INSL6 intron 1 are marked in yellow and green, respectively. The non-template sequence is marked in blue, and stop codon (UAG), poly-A signal (AAUAAA) and poly-A in red. The putative translated region is underlined. b, Genomic structure of the rearranged PD-L1 locus and transcription in two representative cases (ATL012 and ATL079) with 3′-UTR-truncated PD-L1 transcripts, in which PD-L1 ORF is terminated before exon 6 or 7, and merged into an intergenic sequence. Breakpoints (blue dotted lines) are shown with accompanying copy number alterations. c, Structure and breakpoint sequence of PD-L1 fusion transcripts (top) with Sanger sequencing chromatogram (bottom). d, Length of abnormal PD-L1 transcripts identified in ATL samples with PD-L1 SVs, compared with wild-type PD-L1. e, Genomic and transcript sequences from the PD-L1 locus containing 327 bp inversion within the last exon identified in case ATL017. Aberrant PD-L1 transcripts have a putative poly-A signal sequence in the inverted region, followed by poly-A tract.
Extended Data Figure 3 Elevated PD-L1 mRNA expression in ATL according to PD-L1 SV state.
Diagonal plots between PD-L1 exon 3 expression (RPKM) and its relative value to that of 3′-UTR (exon 3 to 3′-UTR ratio) for 43 ATL cases. SV (+) cases are indicated by corresponding colours to each SV type.
Extended Data Figure 4 PD-L1 fusion proteins identified in ATL.
a, Amino acid sequence alignment of wild-type and truncated PD-L1 proteins. Transmembrane domain is shaded in blue. Conserved regions are shown in red. b, Western blot analysis with antibodies against the N-terminal and C-terminal domains of PD-L1 in PC-9 cells transduced with indicated PD-L1 constructs. c, d, Flow cytometry plots for PD-L1 surface expression (c) and PD-1 Ig binding (d) in PC-9 cells transduced with indicated PD-L1 constructs. e, Western blot of PD-L1 SV(+) cases harbouring intact or truncated ORFs, compared with SV(−) cases using antibodies specifically detecting N-terminal and C-terminal domains of PD-L1. b–e, Representative of three independent experiments.
Extended Data Figure 5 Flow chart for detecting abnormal PD-L1 transcripts in the TCGA cohort.
In total, 10,210 tumour samples in 33 tumour types, for which RNA-seq data were available in TCGA, were assessed by exon-level expression quantification and fusion detection by STAR algorithm. Viral integration within or near the PD-L1 gene was also searched in this cohort. After manual review by IGV, a total of 32 cases with aberrant PD-L1 transcription were identified.
Extended Data Figure 6 3′-truncated PD-L1 mRNA transcripts in the TCGA cohort.
RNA-seq data are visualized by IGV for TCGA samples with abnormal PD-L1 transcription. Aberrant PD-L1 transcripts were shown in red.
Extended Data Figure 7 Aberrant transcription affecting PD-L1 3′-UTR and associated genomic alterations identified in multiple cancers.
a, Genomic structure of the rearranged PD-L1 locus and transcription in FA-A4XK-01 (DLBC), BP-4983-01 (KIRC), L5-L4OE-01 (ESCA), and F5-6814-01 (READ), showing loss of PD-L1 3′-UTR transcription and fusion transcripts between PD-L1 and intronic or intergenic segments. Breakpoints (blue dotted lines) are shown with accompanying copy number alterations. Del, deletion. b, PD-L1 DNA copy number versus JAK2 mRNA expression across 48 DLBC (left), 415 STAD (middle), and 43 ATL (right) samples. SV(+) samples (red) and those with 9p24.1 copy number gains involving both JAK2 and PD-L1 genes (orange) are indicated. P values for the effects of PD-L1 SVs and copy number on JAK2 expression (GLM) are shown. c, Genomic structure of the rearranged PD-L1 locus and transcription in two cases with viral integrations around the PD-L1 gene; a STAD case (FP-7998-01) with an EBV integration (top) and an HNSC case (CV-5443-01), showing HPV16 integration, which was described previously19, and premature termination of PD-L1 transcripts within intron 4 (bottom).
Extended Data Figure 8 Induction of Pd-l1 3′-UTR deletions and inversions in mouse cell lines using the CRISPR-Cas9 system.
a, Positions of targeting sgRNAs used for CRISPR-Cas9-mediated disruption of Pd-l1 3′-UTR are indicated by arrows. b, Pd-l1 surface expression in EG7-OVA cells transfected with Cas9 and no, single, or pairwise sgRNAs. Representative of three independent experiments. c, d, PCR detection of the Pd-l1 3′-UTR deletion (c) or inversion (d) breakpoint junction from EG7-OVA, P815, and B16-F10 cells in which Cas9 was expressed without (parental) or with no sgRNA (mock), or a pair of Pd-l1 sgRNAs. e, Sequence chromatogram of the detected Pd-l1 3′-UTR deletions from sgPd-l1-transfected EG7-OVA, P815, and B16-F10 cells. f, g, Pd-l1 exon 4 mRNA expression (RPKM) was calculated from the RNA-seq data for EG7-OVA, P815, and B16-F10 cells in which Cas9 was expressed without (parental) or with no sgRNA (mock), or a pair of Pd-l1 sgRNAs (f). RNA-seq reads within the Pd-l1 gene were visualized by IGV (g).
Extended Data Figure 9 Induction of PD-L1 3′-UTR deletions and inversions in human cell lines using the CRISPR-Cas9 system.
a, PCR detection of the PD-L1 3′-UTR deletion breakpoint junction from T2 cells in which Cas9 was expressed without (parental) or with no sgRNA (mock), or a pair of PD-L1 sgRNAs. b, Sequence chromatogram of the detected PD-L1 3′-UTR deletions from sgPD-L1-transfected HEK293T and T2 cells. c, PCR detection of the PD-L1 3′-UTR inversion breakpoint junction from HEK293T, T2, and PC-9 cells in which Cas9 was expressed without (parental) or with no sgRNA (mock), or a pair of PD-L1 sgRNAs. d, Sequence chromatogram of the detected PD-L1 3′-UTR inversions from sgPD-L1-transfected HEK293T, T2, and PC-9 cells. e, Visualization of RNA-seq reads within the PD-L1 gene for T2 and PC-9 cells in which Cas9 was expressed without (parental) or with no sgRNA (mock), or a pair of PD-L1 sgRNAs. f, Flow cytometric analysis of PD-L1 surface expression in parental or sgPD-L1-transfected PC-9 cells stimulated with IFN-γ (100 or 300 U ml−1) for 48 h. Representative of three independent experiments.
Extended Data Figure 10 Tumour-intrinsic Pd-l1 activation by 3′-UTR loss suppresses CD8+ cytotoxic T lymphocyte recruitment within the tumour microenvironment.
a, Strategy for evaluating the effect of Pd-l1 3′-UTR disruption on anti-tumour immunity. b, Representative immunofluorescence images (from experiments in Fig. 4c) of CD8 (green) and DAPI (purple) staining in mock- and sgPd-l1-transfected EG7-OVA tumours treated with PBS or poly(I:C). c, Flow cytometric analysis showing frequency of CD8+ T cells infiltrating into mock- and sgPd-l1-transfected EG7-OVA tumours treated with PBS or poly(I:C) (n = 6 per group; Welch’s t-test). Data represent mean ± s.e.m. d, Flow cytometric analysis showing frequency of CD8+ T cells infiltrating into sgPd-l1-transfected, poly(I:C)-treated EG7-OVA tumours treated with isotype control or anti-Pd-l1 antibody (n = 7 per group; Welch’s t-test). Data represent mean ± s.e.m.
Supplementary information
Supplementary Figure
This file contains a Supplementary Figure showing uncropped images with size marker indications. (PDF 719 kb)
Supplementary Tables
This file contains Supplementary Tables 1-8. (XLSX 394 kb)
Source data
Rights and permissions
About this article
Cite this article
Kataoka, K., Shiraishi, Y., Takeda, Y. et al. Aberrant PD-L1 expression through 3′-UTR disruption in multiple cancers. Nature 534, 402–406 (2016). https://doi.org/10.1038/nature18294
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature18294
This article is cited by
-
DNA mismatch repair system regulates the expression of PD-L1 through DNMTs in cervical cancer
Cancer Cell International (2024)
-
Cancer immune evasion through KRAS and PD-L1 and potential therapeutic interventions
Cell Communication and Signaling (2023)
-
Dynamics in the expression of programmed death ligand 1 and cluster of differentiation 163 in the tumor microenvironment of uterine cervical cancer: a single-center retrospective study
Radiation Oncology (2023)
-
Inhibition of PD-L1 and tumor growth in triple-negative breast cancer using a magnetic nanovector with microRNA34a
Cancer Nanotechnology (2023)
-
Programmed death ligand 1 expression in diffuse large B cell lymphoma: correlation with clinicopathological prognostic factors
Journal of the Egyptian National Cancer Institute (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.