Abstract
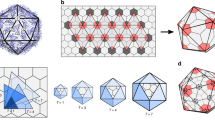
The icosahedron is the largest of the Platonic solids, and icosahedral protein structures are widely used in biological systems for packaging and transport1,2. There has been considerable interest in repurposing such structures3,4,5 for applications ranging from targeted delivery to multivalent immunogen presentation. The ability to design proteins that self-assemble into precisely specified, highly ordered icosahedral structures would open the door to a new generation of protein containers with properties custom-tailored to specific applications. Here we describe the computational design of a 25-nanometre icosahedral nanocage that self-assembles from trimeric protein building blocks. The designed protein was produced in Escherichia coli, and found by electron microscopy to assemble into a homogenous population of icosahedral particles nearly identical to the design model. The particles are stable in 6.7 molar guanidine hydrochloride at up to 80 degrees Celsius, and undergo extremely abrupt, but reversible, disassembly between 2 molar and 2.25 molar guanidinium thiocyanate. The icosahedron is robust to genetic fusions: one or two copies of green fluorescent protein (GFP) can be fused to each of the 60 subunits to create highly fluorescent ‘standard candles’ for use in light microscopy, and a designed protein pentamer can be placed in the centre of each of the 20 pentameric faces to modulate the size of the entrance/exit channels of the cage. Such robust and customizable nanocages should have considerable utility in targeted drug delivery6, vaccine design7 and synthetic biology8.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
06 July 2016
An addition was made to the Acknowledgements section.
19 October 2016
A Correction to this paper has been published: https://doi.org/10.1038/nature20108
References
Zandi, R., Reguera, D., Bruinsma, R. F., Gelbart, W. M. & Rudnick, J. Origin of icosahedral symmetry in viruses. Proc. Natl Acad. Sci. USA 101, 15556–15560 (2004)
Ritsert, K. et al. Studies on the lumazine synthase/riboflavin synthase complex of Bacillus subtilis: crystal structure analysis of reconstituted, icosahedral beta-subunit capsids with bound substrate analogue inhibitor at 2.4 Å resolution. J. Mol. Biol. 253, 151–167 (1995)
Howorka, S. Rationally engineering natural protein assemblies in nanobiotechnology. Curr. Opin. Biotechnol. 22, 485–491 (2011)
Roldão, A., Mellado, M. C. M., Castilho, L. R., Carrondo, M. J. T. & Alves, P. M. Virus-like particles in vaccine development. Expert Rev. Vaccines 9, 1149–1176 (2010)
Effio, C. L. & Hubbuch, J. Next generation vaccines and vectors: designing downstream processes for recombinant protein-based virus-like particles. Biotechnol. J. 10, 715–727 (2015)
Ma, Y., Nolte, R. J. M. & Cornelissen, J. J. L. M. Virus-based nanocarriers for drug delivery. Adv. Drug Deliv. Rev. 64, 811–825 (2012)
Smith, M. L. et al. Modified tobacco mosaic virus particles as scaffolds for display of protein antigens for vaccine applications. Virology 348, 475–488 (2006)
Bauler, P., Huber, G., Leyh, T. & McCammon, J. A. Channeling by proximity: the catalytic advantages of active site colocalization using Brownian dynamics. J. Phys. Chem. Lett. 1, 1332–1335 (2010)
Brodin, J. D. et al. Metal-directed, chemically tunable assembly of one-, two- and three-dimensional crystalline protein arrays. Nat. Chem. 4, 375–382 (2012)
Der, B. S. et al. Metal-mediated affinity and orientation specificity in a computationally designed protein homodimer. J. Am. Chem. Soc. 134, 375–385 (2012)
Fletcher, J. M. et al. Self-assembling cages from coiled-coil peptide modules. Science 340, 595–599 (2013)
Usui, K. et al. Nanoscale elongating control of the self-assembled protein filament with the cysteine-introduced building blocks. Protein Sci. 18, 960–969 (2009)
Raman, S., Machaidze, G., Lustig, A., Aebi, U. & Burkhard, P. Structure-based design of peptides that self-assemble into regular polyhedral nanoparticles. Nanomedicine 2, 95–102 (2006)
Raman, S. et al. Design of peptide nanoparticles using simple protein oligomerization domains. Open Nanomed. J. 2, 15–26 (2009)
Sinclair, J. C., Davies, K. M., Vénien-Bryan, C. & Noble, M. E. M. Generation of protein lattices by fusing proteins with matching rotational symmetry. Nat. Nanotechnol. 6, 558–562 (2011)
Boyle, A. L. et al. Squaring the circle in peptide assembly: from fibers to discrete nanostructures by de novo design. J. Am. Chem. Soc. 134, 15457–15467 (2012)
Lai, Y.-T. et al. Structure of a designed protein cage that self-assembles into a highly porous cube. Nat. Chem. 6, 1065–1071 (2014)
King, N. P. et al. Computational design of self-assembling protein nanomaterials with atomic level accuracy. Science 336, 1171–1174 (2012)
King, N. P. et al. Accurate design of co-assembling multi-component protein nanomaterials. Nature 510, 103–108 (2014)
Leaver-Fay, A. et al. ROSETTA3: an object-oriented software suite for the simulation and design of macromolecules. Methods Enzymol. 487, 545–574 (2011)
DiMaio, F., Leaver-Fay, A., Bradley, P., Baker, D. & André, I. Modeling symmetric macromolecular structures in Rosetta3. PLoS One 6, e20450 (2011)
Lawrence, M. C. & Colman, P. M. Shape complementarity at protein/protein interfaces. J. Mol. Biol. 234, 946–950 (1993)
Griffiths, J. S. et al. Cloning, isolation and characterization of the Thermotoga maritima KDPG aldolase. Bioorg. Med. Chem. 10, 545–550 (2002)
Fullerton, S. W. B. et al. Mechanism of the class I KDPG aldolase. Bioorg. Med. Chem. 14, 3002–3010 (2006)
Perlmutter, J. D. & Hagan, M. F. Mechanisms of virus assembly. Annu. Rev. Phys. Chem. 66, 217–239 (2015)
Pédelacq, J.-D., Cabantous, S., Tran, T., Terwilliger, T. C. & Waldo, G. S. Engineering and characterization of a superfolder green fluorescent protein. Nat. Biotechnol. 24, 79–88 (2006)
Andrews, B. T., Schoenfish, A. R., Roy, M., Waldo, G. & Jennings, P. A. The rough energy landscape of superfolder GFP is linked to the chromophore. J. Mol. Biol. 373, 476–490 (2007)
Cortese, K., Diaspro, A. & Tacchetti, C. Advanced correlative light/electron microscopy: current methods and new developments using Tokuyasu cryosections. J. Histochem. Cytochem. 57, 1103–1112 (2009)
Huang, P.-S. et al. High thermodynamic stability of parametrically designed helical bundles. Science 346, 481–485 (2014)
Zhou, Z. et al. Genetically encoded short peptide tags for orthogonal protein labeling by Sfp and AcpS phosphopantetheinyl transferases. ACS Chem. Biol. 2, 337–346 (2007)
Baalousha, M. & Lead, J. R. Nanoparticle dispersity in toxicology. Nat. Nanotechnol. 8, 308–309 (2013)
Zulauf, M. & D’Arcy, A. Light scattering of proteins as a criterion for crystallization. J. Cryst. Growth 122, 102–106 (1992)
Nannenga, B. L., Iadanza, M. G., Vollmar, B. S. & Gonen, T. in Current Protocols in Protein Science (eds Coligan, J. E., Dunn, B. M., Speicher, D. W. & Wingfield, P. T. ) Ch. 17.15 (John Wiley & Sons, 2013)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 10, 584–590 (2013)
van Heel, M., Harauz, G., Orlova, E. V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996)
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996)
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012)
Huang, P.-S. et al. RosettaRemodel: a generalized framework for flexible backbone protein design. PLoS One 6, e24109 (2011)
Muller, E. G. D. et al. The organization of the core proteins of the yeast spindle pole body. Mol. Biol. Cell 16, 3341–3352 (2005)
Shimogawa, M. M., Wargacki, M. M., Muller, E. G. & Davis, T. N. Laterally attached kinetochores recruit the checkpoint protein Bub1, but satisfy the spindle checkpoint. Cell Cycle 9, 3619–3628 (2010)
Acknowledgements
This work was supported by the Howard Hughes Medical Institute (D.B. and T.G.), the JRC visitor programme (S.G.), the National Science Foundation CHE-1332907 (D.B.), a UW/Hutch CCSG Pilot Award NCI 5 P30 CA015704-41 (D.B. and N.P.K.), Takeda Pharmaceutical Company (N.P.K.), the Bill and Melinda Gates Foundation OPP1120319 (D.B. and N.P.K.), the National Institutes of Health (NIH) P41 GM103533 (T.N.D.), the Defense Advanced Research Projects Agency (D.B. and N.P.K., grant no. W911NF-14-1-0162) and the Air Force Office of Scientific Research (AFOSR) AFOSR FA950-12-10112 (D.B.). Y.H. was supported in part by a NIH Molecular Biology Training Grant (T32GM008268). U.N. was supported in part by a PHS National Research Service Award (T32GM007270) from NIGMS. J.B.B. was supported in part by an NSF Graduate Research Fellowship (DGE-0718124). We thank the Janelia Research Campus Cryo-EM Facility and J. de la Cruz for their assistance with the Titan Krios.
Author information
Authors and Affiliations
Contributions
J.B.B., N.P.K., and W.S. developed the computational design methodology. Y.H. and J.B.B. performed the design of the icosahedra. Y.H. performed all other unlisted experiments. S.G. and D.S. performed the cryo-EM experiments. K.K.F. performed the fluorescence microscopy experiments. U.N. performed the negative-stain electron microscopy experiments. C.X. provided the pentamer sequence for I3-01(HB). P.-S.H. created the computational methodology to model fusions to I3-01. R.R. produced I3-01(HB) proteins. S.Y. produced T33-21 sfGFP fusion proteins.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 I3-01 tolerance to temperature.
DLS measurements as I3-01 is subjected to heating to 90 °C (solid line), then cooling to 25 °C (dotted line) in TBS (a), 6.7 M GuHCl (b) and 2 M GITC (c). Under all three conditions, any indications of aggregation or increase in size due to temperature appear to be completely reversible.
Extended Data Figure 2 Reproducibility of I3-01 transition in 2 M to 2.25 M GITC.
Four examples each of independent measurements at 2 M (blue) and 2.25 M (red) GITC using DLS show the reproducibility of the cage disassociation. Histograms are plotted offset by 1% intensity from each other for clarity.
Extended Data Figure 3 SEC of T33-21 and I3-01 fused with sfGFP.
Size exclusion chromatography traces for T33-21 (12mer in red and 24mer in blue) and I3-01 (60mer in green and 120mer in purple) sfGFP fusions, display increased particle sizes with increasing copies of GFP, but retain monodispersed populations. The N-terminal fusion of sfGFP (dashed line) is expected to extend mostly outward from the icosahedron, thus greatly increasing the hydrodynamic radius while the C-terminal fusion is predicted to occupy the internal void space. A230, ultraviolet absorbance at 230 nm; mAU, milli-absorbance units.
Extended Data Figure 4 Tolerance of I3-01–sfGFP fusions to GuHCl.
N-terminal (red) and C-terminal (blue) sfGFP fusions were equilibrated to 0–6.4 M GuHCl. Ultraviolet absorbance at 490 nm (A490) monitors the unfolding of sfGFP (top, solid line and crosses). DLS experiments (top, dotted line and dots) reveal as sfGFP unfolds, the hydrodynamic radius increases slightly, and then stabilizes. The bottom panels show that in 1 M GuHCl (solid line) and in 6 M GuHCl (dotted line), the icosahedral assemblies remain relatively monodisperse.
Extended Data Figure 5 I3-01 C-terminal fusions with other fluorescent proteins.
Fluorescent proteins mTurquoise2 (in blue) or sYFP2 (in green) were fused to the C terminus of I3-01. The field of view using widefield fluorescence microscopy shows distinct signals of each type when the two types are mixed together.
Extended Data Figure 6 I3-01 retains native enzyme activity.
Coupled KDPG aldolase assay showing native-like enzymatic activity in I3-01. The K129A knockout shows no enzyme activity, similar to buffer alone. UV339, absorbance at 339 nm; error bars are standard deviation.
Supplementary information
Supplementary Information
This zipped file contains the protein sequences, the design structure PDB file and an example script. (ZIP 3559 kb)
Rights and permissions
About this article
Cite this article
Hsia, Y., Bale, J., Gonen, S. et al. Design of a hyperstable 60-subunit protein icosahedron. Nature 535, 136–139 (2016). https://doi.org/10.1038/nature18010
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature18010
This article is cited by
-
Mosaic quadrivalent influenza vaccine single nanoparticle characterization
Scientific Reports (2024)
-
Higher-order protein assembly controls kinetochore formation
Nature Cell Biology (2024)
-
Asymmetric oligomerization state and sequence patterning can tune multiphase condensate miscibility
Nature Chemistry (2024)
-
DNA-origami-directed virus capsid polymorphism
Nature Nanotechnology (2023)
-
A supramolecular system mimicking the infection process of an enveloped virus through membrane fusion
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.