Abstract
Circadian clocks are fundamental to the biology of most eukaryotes, coordinating behaviour and physiology to resonate with the environmental cycle of day and night through complex networks of clock-controlled genes1,2,3. A fundamental knowledge gap exists, however, between circadian gene expression cycles and the biochemical mechanisms that ultimately facilitate circadian regulation of cell biology4,5. Here we report circadian rhythms in the intracellular concentration of magnesium ions, [Mg2+]i, which act as a cell-autonomous timekeeping component to determine key clock properties both in a human cell line and in a unicellular alga that diverged from each other more than 1 billion years ago6. Given the essential role of Mg2+ as a cofactor for ATP, a functional consequence of [Mg2+]i oscillations is dynamic regulation of cellular energy expenditure over the daily cycle. Mechanistically, we find that these rhythms provide bilateral feedback linking rhythmic metabolism to clock-controlled gene expression. The global regulation of nucleotide triphosphate turnover by intracellular Mg2+ availability has potential to impact upon many of the cell’s more than 600 MgATP-dependent enzymes7 and every cellular system where MgNTP hydrolysis becomes rate limiting. Indeed, we find that circadian control of translation by mTOR8 is regulated through [Mg2+]i oscillations. It will now be important to identify which additional biological processes are subject to this form of regulation in tissues of multicellular organisms such as plants and humans, in the context of health and disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Covington, M. F., Maloof, J. N., Straume, M., Kay, S. A. & Harmer, S. L. Global transcriptome analysis reveals circadian regulation of key pathways in plant growth and development. Genome Biol. 9, R130 (2008)
Hughes, M. E. et al. Harmonics of circadian gene transcription in mammals. PLoS Genet. 5, e1000442 (2009)
Endo, M., Shimizu, H., Nohales, M. A., Araki, T. & Kay, S. A. Tissue-specific clocks in Arabidopsis show asymmetric coupling. Nature 515, 419–422 (2014)
Edgar, R. S. et al. Peroxiredoxins are conserved markers of circadian rhythms. Nature 485, 459–464 (2012)
Bass, J. Circadian topology of metabolism. Nature 491, 348–356 (2012)
Hedges, S. B., Dudley, J. & Kumar, S. TimeTree: a public knowledge-base of divergence times among organisms. Bioinformatics 22, 2971–2972 (2006)
de Baaij, J. H., Hoenderop, J. G. & Bindels, R. J. Magnesium in man: implications for health and disease. Physiol. Rev. 95, 1–46 (2015)
Lipton, J. O. et al. The circadian protein BMAL1 regulates translation in response to S6K1-mediated phosphorylation. Cell 161, 1138–1151 (2015)
Dunlap, J. C. Molecular bases for circadian clocks. Cell 96, 271–290 (1999)
Corellou, F. et al. Clocks in the green lineage: comparative functional analysis of the circadian architecture of the picoeukaryote Ostreococcus . Plant Cell 21, 3436–3449 (2009)
Hastings, M. H., Maywood, E. S. & O’Neill, J. S. Cellular circadian pacemaking and the role of cytosolic rhythms. Curr. Biol. 18, R805–R815 (2008)
Olmedo, M. et al. Circadian regulation of olfaction and an evolutionarily conserved, nontranscriptional marker in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 109, 20479–20484 (2012)
O’Neill, J. S. & Reddy, A. B. Circadian clocks in human red blood cells. Nature 469, 498–503 (2011)
O’Neill, J. S. et al. Circadian rhythms persist without transcription in a eukaryote. Nature 469, 554–558 (2011)
van Ooijen, G. & Millar, A. J. Non-transcriptional oscillators in circadian timekeeping. Trends Biochem. Sci. 37, 484–492 (2012)
Ko, G. Y., Shi, L. & Ko, M. L. Circadian regulation of ion channels and their functions. J. Neurochem. 110, 1150–1169 (2009)
Njus, D., Sulzman, F. M. & Hastings, J. W. Membrane model for the circadian clock. Nature 248, 116–120 (1974)
Nitabach, M. N., Holmes, T. C. & Blau, J. Membranes, ions, and clocks: testing the Njus-Sulzman-Hastings model of the circadian oscillator. Methods Enzymol. 393, 682–693 (2005)
Danku, J. M., Lahner, B., Yakubova, E. & Salt, D. E. Large-scale plant ionomics. Methods Mol. Biol. 953, 255–276 (2013)
Danku, J. M. C., Gumaelius, L., Baxter, I. & Salt, D. E. A high-throughput method for Saccharomyces cerevisiae (yeast) ionomics. J. Anal. At. Spectrom. 24, 103–107 (2009)
Nishinaga, H. et al. Circadian expression of the Na+/H+ exchanger NHE3 in the mouse renal medulla. Biomed. Res. 30, 87–93 (2009)
Wang, Y. C., Chen, Y. S., Cheng, R. C. & Huang, R. C. Role of Na+/Ca2+ exchanger in Ca2+ homeostasis in rat suprachiasmatic nucleus neurons. J. Neurophysiol. 113, 2114–2126 (2015)
Rubin, H. The logic of the membrane, magnesium, mitosis (MMM) model for the regulation of animal cell proliferation. Arch. Biochem. Biophys. 458, 16–23 (2007)
Pittendrigh, C. S. Circadian rhythms and the circadian organization of living systems. Cold Spring Harb. Symp. Quant. Biol. 25, 159–184 (1960)
Pizarro, A., Hayer, K., Lahens, N. F. & Hogenesch, J. B. CircaDB: a database of mammalian circadian gene expression profiles. Nucleic Acids Res. 41, D1009–D1013 (2013)
Zhang, E. E. et al. A genome-wide RNAi screen for modifiers of the circadian clock in human cells. Cell 139, 199–210 (2009)
Kolisek, M. et al. SLC41A1 is a novel mammalian Mg2+ carrier. J. Biol. Chem. 283, 16235–16247 (2008)
Kucharski, L. M., Lubbe, W. J. & Maguire, M. E. Cation hexaammines are selective and potent inhibitors of the CorA magnesium transport system. J. Biol. Chem. 275, 16767–16773 (2000)
Kolisek, M., Nestler, A., Vormann, J. & Schweigel-Rontgen, M. Human gene SLC41A1 encodes for the Na+/Mg2+ exchanger. Am. J. Physiol. Cell Physiol. 302, C318–C326 (2012)
O’Neill, J. S., Maywood, E. S., Chesham, J. E., Takahashi, J. S. & Hastings, M. H. cAMP-dependent signaling as a core component of the mammalian circadian pacemaker. Science 320, 949–953 (2008)
van Ooijen, G., Dixon, L. E., Troein, C. & Millar, A. J. Proteasome function is required for biological timing throughout the twenty-four hour cycle. Curr. Biol. 21, 869–875 (2011)
van Ooijen, G. et al. Functional analysis of casein kinase 1 in a minimal circadian system. PLoS ONE 8, e70021 (2013)
Le Bihan, T. et al. Label-free quantitative analysis of the casein kinase 2-responsive phosphoproteome of the marine minimal model species Ostreococcus tauri. Proteomics 15, 4135–4144 (2015)
Blanc-Mathieu, R. et al. An improved genome of the model marine alga Ostreococcus tauri unfolds by assessing Illumina de novo assemblies. BMC Genom . 15, 1103 (2014)
Sterck, L., Billiau, K., Abeel, T., Rouze, P. & Van de Peer, Y. ORCAE: online resource for community annotation of eukaryotes. Nature Methods 9, 1041 (2012)
Baker, C. L., Kettenbach, A. N., Loros, J. J., Gerber, S. A. & Dunlap, J. C. Quantitative proteomics reveals a dynamic interactome and phase-specific phosphorylation in the Neurospora circadian clock. Mol. Cell 34, 354–363 (2009)
Valekunja, U. K. et al. Histone methyltransferase MLL3 contributes to genome-scale circadian transcription. Proc. Natl Acad. Sci. USA 110, 1554–1559 (2013)
O’Neill, J. S. & Hastings, M. H. Increased coherence of circadian rhythms in mature fibroblast cultures. J. Biol. Rhythms 23, 483–488 (2008)
van der Horst, G. T. et al. Mammalian Cry1 and Cry2 are essential for maintenance of circadian rhythms. Nature 398, 627–630 (1999)
Xu, J. in Current Protocols in Molecular Biology (eds Ausubel, F. M. et al.) Ch. 28, Unit 28.1 (Wiley, 2005)
David, A. et al. Nuclear translation visualized by ribosome-bound nascent chain puromycylation. J. Cell Biol. 197, 45–57 (2012)
Fisher, N. I. Statistical Analysis of Circular Data, xviii + 1–277 (Cambridge Univ. Press, 1993)
Jammalamadaka, S. R. & SenGupta, A. Topics in Circular Statistics 1–322 (World Scientific, 2001)
Hirota, T. et al. A chemical biology approach reveals period shortening of the mammalian circadian clock by specific inhibition of GSK-3β. Proc. Natl Acad. Sci. USA 105, 20746–20751 (2008)
Monnier, A. et al. Orchestrated transcription of biological processes in the marine picoeukaryote Ostreococcus exposed to light/dark cycles. BMC Genom . 11, 192 (2010)
Wu, C. et al. BioGPS: an extensible and customizable portal for querying and organizing gene annotation resources. Genome Biol. 10, R130 (2009)
Boratyn, G. M. et al. Domain enhanced lookup time accelerated BLAST. Biol. Direct 7, 12 (2012)
Acknowledgements
G.v.O. is supported by a Royal Society University Research Fellowship (UF110173) and research grants (RS120372 and RS140275). J.S.O. is supported by the Medical Research Council (MC_UP_1201/4) and the Wellcome Trust (093734/Z/10/Z). M.P. is funded by KWF BUIT 2014-6637. L.F.L. and C.O.Y. are supported by Millennium Nucleus for Fungal Integrative and Synthetic Biology (NC120043), and Fondo Nacional de Desarrollo Científico y Tecnológico (FONDECYT 1131030). S. K. Hodge is acknowledged for managing imaging facilities for algal experiments. At the MRC LMB, we are grateful to the Biomedical Services Group for animal care; M. Hastings and J. Chesham for supplying mouse tissue; P. Margiotta for assistance with figures. We also thank D. E. Salt, M. Knight, E. Grünewald, P. Crosby, L. Hewitt and B. Cross for criticism. The anti-puromycin ascites was a gift from M. Hegde.
Author information
Authors and Affiliations
Contributions
G.v.O. and J.S.O. conceived the approach and designed the study. L.F.L. and C.O.Y. generated the Neurospora result. J.D. and L.E. performed ICP analyses. G.v.O. and L.L.H. performed Ostreococcus experiments. Human U2OS cell experiments were performed by K.A.F. M.P. performed mouse fibroblast experiments. N.P.H. provided analytical and intellectual contributions. G.v.O. and J.S.O. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
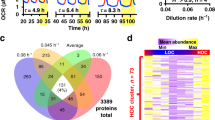
Extended Data Figure 1 Additional ICP-MS data and controls (Ostreococcus).
a, b, ICP-MS analyses of cell lysates from 12 h:12 h light/dark cycles (a) or on the second day of constant light (b). P values report significance by one-way ANOVA (mean ± s.e.m., n = 3). c, ICP-MS analyses on cell lysates compared with media control (no cells) and membrane fractions (lysed cells) (mean ± s.d. plotted, n = 2 biological replicates), indicating that magnesium signal in a and Fig. 1 comes predominantly from the intracellular space. Groups are significantly different by one-way ANOVA (P < 0.0001); Tukey’s multiple comparison P values are indicated. d, Fluctuations in measured concentrations are not related to fluctuations in cell size over time. No significance of time as source of variation in cell size was observed by fluorescence-activated cell sorting analyses (mean ± s.d. plotted, one-way ANOVA P value is indicated, n = 5 biological replicates).
Extended Data Figure 2 Additional ICP-MS data (U2OS cells) and controls.
a, ICP-MS analyses of U2OS cell extracts for stable isotopes of several biologically relevant ions (mean ± s.e.m., grey/black, n ≥ 4 biological replicates), with insets showing standards that indicate linearity over the observed concentration ranges (mean ± percentage coefficient of variation). We compared how well a straight-line + damped sine wave model (adapted from ref. 44) fit to each time series compared with a straight-line only (null hypothesis, no rhythm). The null hypothesis was preferred in each case except for Mg and K (analysed by 24Mg and 39K), where the sinusoidal fit with a circadian period was preferred (blue line, R2 and fit period ± s.e.m. are reported). b, Bmal1:luc bioluminescence data showing no effect of serum concentration on circadian period in U2OS cells in the presence of B-27 supplement, mean ± s.e.m. (n = 3 biological replicates). c, Quantification of period and amplitude for data shown in b (one-way ANOVA for period, P = 0.79; one-way ANOVA for amplitude, P = 0.01). d, Cellular impedance measurements indicate that U2OS cells do not proliferate upon reaching stationary phase under our assay conditions, reported doubling times (Td) were calculated from data collected between the dotted lines.
Extended Data Figure 3 Circadian rhythms of [Mg2+]i in N. crassa and mouse fibroblasts.
a, Circadian regulation of [Mg2+]i detected by ICP-MS in the fungus N. crassa under constant darkness (mean ± s.e.m., n = 3 biological replicates). b, Representative (out of three) FRQ immunoblot sampled in parallel. c, Quantification of FRQ abundance (mean ± s.e.m., n = 3 biological replicates). d, Circadian regulation of [Mg2+]i measured by luciferase-based assay is dependent upon CRYPTOCHROME in immortalized adult mouse fibroblasts under constant conditions (mean ± s.e.m., n = 3 biological replicates).
Extended Data Figure 4 Rhythms of [Mg2+]i entrain to relevant external cues and are temperature-compensated.
a, Inversion of 12 h:12 h light/dark entrainment cycles is sufficient to entrain the phase of [Mg2+]i in Ostreococcus cells, measured by luciferase assay under constant light (mean ± s.e.m., n = 3 biological replicates). b, From the start of the experiment (S), 3 days of 12 h:12 h temperature cycles between 32 and 37 °C, followed by a change to air medium (M), are sufficient to entrain the phase of [Mg2+]i in U2OS cells measured by ICP-MS over two circadian cycles under constant conditions (mean ± s.e.m., n = 3 biological replicates). c, Ostreococcus bioluminescence recordings (CCA1-LUC) at the indicated temperatures (n = 8 replicate wells). Vertical dotted lines indicate sampling window for Mg2+ assays reported in d: assays performed during the second cycle under constant conditions show that circadian [Mg2+]i rhythms are temperature-compensated (n = 4 biological replicates). Each data set was fitted with a Lorentzian curve to estimate peak [Mg2+]i. e, No significant difference in temperature compensation (Q10) between CCA1-LUC rhythms and the timing of the second Mg2+i peak; unpaired t-test P value is reported.
Extended Data Figure 5 Human magnesium transporters and conservation in O. tauri.
a, Ubiquitously expressed human proteins with a clearly defined Mg2+ transport activity7 are listed. Note that many additional putative Mg2+-transporters are annotated, with several of these also being circadian-regulated in multiple mouse tissues. b, Expression profiles of Ostreococcus homologues of mammalian Mg2+ channels and transporters listed in a, mined from publically available microarray data45. *From BioGPS26,46. †From CircaDB25 with JTK cycle P < 0.05. ‡From the Orcae service34,35. §Percentage sequence identity/similarity with human protein sequence (E value). DELTA-BLAST47 performed using default settings. ||From micro-array data45 shown in b.
Extended Data Figure 6 Chronic CPA treatment dose-dependently lengthens period.
a–f, Traces (a, b) of the CCA1-LUC (Ostreococcus) or Per2:luc (U2OS cells) reporters, showing the effect of inhibition of magnesium transport by Co(NH3)5Cl2+ (CPA) upon period dose–response (c, d) and upon [Mg2+]i (e, f). All plots show mean ± s.e.m., with biological replicate numbers (n) indicated. P values report significance by one-way ANOVA and post-test for linear trend.
Extended Data Figure 7 Period lengthening by CHA and SLC41 knockdown is dependent upon extracellular magnesium.
a, Extracellular magnesium-depletion and CHA act synergistically to lengthen circadian period in Ostreococcus cells (mean ± s.e.m., n = 4 biological replicates). b, Quantification of period lengthening by CHA at different concentrations of extracellular magnesium (mean ± s.e.m., n = 4 biological replicates). P value for two-way ANOVA (interaction effect) is reported. c, Extracellular magnesium-depletion and CHA act synergistically to lengthen circadian period in human U2OS cells (mean ± s.e.m., n = 6 biological replicates). d, Quantification of period lengthening by CHA in Mg2+-depleted versus normal media (mean ± s.e.m., n = 4 biological replicates). P values for two-way ANOVA (interaction effect) and Fisher’s exact test are reported. e, Period lengthening due to knockdown of plasma membrane Mg2+/Na+ antiporter SLC41A1 is attenuated by depletion of extracellular magnesium (mean ± s.e.m., n = 8 biological replicates). f, Quantification of period lengthening due to knockdown of SLC41A1 in normal versus Mg2+-depleted media (mean ± s.e.m., n = 8 biological replicates); two-way ANOVA interaction effect, P < 0.0001. P values for Sidak’s multiple comparisons test are also reported. g, Quantification of SLC41A1 knockdown efficacy. Unpaired t-test P values are reported, a representative immunoblot (of three) is shown (mean ± s.e.m., n = 3 biological replicates).
Extended Data Figure 8 Bioluminescence data of wedge experiment.
Peak expression phase of the clock protein CCA1 was analysed upon re-introduction of magnesium to cultures in low extracellular magnesium, to test whether the phase of cellular rhythms is dictated by the prior phase of entrainment or by this enforced transition from low to high [Mg2+]i. a, Bioluminescence traces showing that circadian rhythms in Ostreococcus are reversibly attenuated by depletion of extracellular Mg2+, and restored by Mg2+ wash-in. b, e, Bioluminescence traces from cells in low extracellular magnesium (b; 5 μM, e; 20 μM) with rhythms rescued by release into media containing normal physiological concentrations of magnesium at the indicated times (vertical dotted lines) in constant light (LL), compared with their respective controls where no magnesium was added in (blue traces). Data from seven or eight replicate wells are shown in each panel. c, f, Summary graphs where results from b, e are plotted in circadian wedge graphs: peak phases of CCA1-LUC rhythms in untreated control cells (grey dots) are compared with peak phase of rhythms reinstated by introduction of physiological magnesium following depletion to 5 μM (c, red dots) or 20 μM (f, orange dots), revealing that the phase of resulting rhythms is dictated solely by the phase of magnesium reintroduction (blue line). d, g, Radial plots of phase shift (mean ± s.d., circumferential axis) depicted in c and f, versus phase before addition of Mg2+ to normal levels (old phase, radial axis and colour). The expected phase responses for type 0 resetting (black dotted line) and no resetting (red dotted line) are indicated. The goodness of fit (R2) and y intercept (Y0) to the type 0 model are shown. Dose-dependent effects of intracellular magnesium on a critical clock parameter are confirmed by the observation that resetting was less strong when magnesium was reintroduced to cells adapted to intermediate levels of extracellular magnesium (e–g) compared with lowest extracellular magnesium (b–d).
Extended Data Figure 9 The effects of magnesium depletion and role of mTOR.
a, Extracellular lactate was measured in U2OS cells after 24 h in Mg2+-depleted compared with normal media. b, c, Combined action of extracellular magnesium depletion and mTOR inhibition using torin1 (b, n = 3 biological replicates) or rapamycin (c, n = 6 biological replicates) to lengthen circadian period in U2OS cells is less than additive (mean ± s.e.m.). d, e, Quantification of period lengthening due to torin1 (d, n = 3) and rapamycin (e, n = 6) in Mg2+-depleted compared with normal media (mean ± s.e.m.). Note the apparent ‘ceiling effect’ at high concentrations of both drugs, such that Mg2+ depletion elicits no additional lengthening of cellular circadian period. Two-way ANOVA interaction effect: P < 0.0001 for both drugs versus Mg2+. Selected P values for Sidak’s multiple comparisons test are also reported (NS, P > 0.33).
Extended Data Figure 10 Factors potentially contributing to maintenance of membrane electroneutrality in light of [Mg2+]i oscillations.
a, Model indicating potential ion fluxes that might explain how clock-regulated [Mg2+]i oscillations impact on global cellular metabolism while membrane electroneutrality is maintained, during the day versus the night. The observed phase dependency of acute CHA was different between Ostreococcus and U2OS cells (Fig. 4a–c), and is consistent with the very different environmental niches inhabited by a marine alga compared with a peripheral human tissue. In Ostreococcus, CHA maintained [Mg2+]i at daytime levels when added before the normal trough, resulting in increased night-time translation and a concomitant reduction in relative ATP levels. This result suggests that Ostreococcus pumps magnesium out of the cell during the dark period, against a large electrochemical potential gradient (magnesium is the second most abundant cation in seawater, at 50 mM in this study) to globally down-tune ATP turnover. In U2OS cells, CHA treatment significantly reduced [Mg2+]i accumulation and translation rates as well as significantly increasing ATP levels when added before the [Mg2+]i peak. Human cells inhabit an environment where nutrient availability is homeostatically regulated (0.8 mM magnesium in cell culture medium). As such, circadian regulation of increased magnesium transport into the cell during the feeding, active phase of day serves to facilitate higher metabolic rate constants. Note that light has no direct effect on the clock in human peripheral cells, instead being mediated by systemic cues. b, c, [Mg2+]i oscillations persist in transcriptionally inactive Ostreococcus cells kept in constant darkness, as analysed both by ICP-MS (b) and by luciferase assay (c), indicating that circadian regulation of ion transport can occur post-translationally in addition to its transcriptional regulation (mean ± s.e.m., n = 3 for ICP-MS data and n = 4 for luciferase assays (biological replicates)).
Rights and permissions
About this article
Cite this article
Feeney, K., Hansen, L., Putker, M. et al. Daily magnesium fluxes regulate cellular timekeeping and energy balance. Nature 532, 375–379 (2016). https://doi.org/10.1038/nature17407
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17407
This article is cited by
-
Nanotandem-rocket releases messenger to disrupt metabolic communication for antitumor immunotherapy
Nano Research (2023)
-
Magnesium Supplementation Stimulates Autophagy to Reduce Lipid Accumulation in Hepatocytes via the AMPK/mTOR Pathway
Biological Trace Element Research (2023)
-
The circadian clock ticks in plant stress responses
Stress Biology (2022)
-
Comment to “Recommendation on an updated standardization of serum magnesium reference ranges”
European Journal of Nutrition (2022)
-
The association between low-carbohydrate diet score and sleep duration among Iranian adults
Sleep and Biological Rhythms (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.