Abstract
Inhibitory neurons regulate the adaptation of neural circuits to sensory experience1, but the molecular mechanisms by which experience controls the connectivity between different types of inhibitory neuron2,3 to regulate cortical plasticity are largely unknown. Here we show that exposure of dark-housed mice to light induces a gene program in cortical vasoactive intestinal peptide (VIP)-expressing neurons that is markedly distinct from that induced in excitatory neurons and other subtypes of inhibitory neuron. We identify Igf1 as one of several activity-regulated genes that are specific to VIP neurons, and demonstrate that IGF1 functions cell-autonomously in VIP neurons to increase inhibitory synaptic input onto these neurons. Our findings further suggest that in cortical VIP neurons, experience-dependent gene transcription regulates visual acuity by activating the expression of IGF1, thus promoting the inhibition of disinhibitory neurons3,4,5 and affecting inhibition onto cortical pyramidal neurons.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hensch, T. K. Critical period plasticity in local cortical circuits. Nature Rev. Neurosci. 6, 877–888 (2005)
Rudy, B., Fishell, G., Lee, S. & Hjerling-Leffler, J. Three groups of interneurons account for nearly 100% of neocortical GABAergic neurons. Dev. Neurobiol. 71, 45–61 (2011)
Pfeffer, C. K., Xue, M., He, M., Huang, Z. J. & Scanziani, M. Inhibition of inhibition in visual cortex: the logic of connections between molecularly distinct interneurons. Nature Neurosci. 16, 1068–1076 (2013)
Lee, S., Kruglikov, I., Huang, Z. J., Fishell, G. & Rudy, B. A disinhibitory circuit mediates motor integration in the somatosensory cortex. Nature Neurosci. 16, 1662–1670 (2013)
Pi, H.-J. et al. Cortical interneurons that specialize in disinhibitory control. Nature 503, 521–524 (2013)
Spiegel, I. et al. Npas4 regulates excitatory-inhibitory balance within neural circuits through cell-type-specific gene programs. Cell 157, 1216–1229 (2014)
Majdan, M. & Shatz, C. J. Effects of visual experience on activity-dependent gene regulation in cortex. Nature Neurosci. 9, 650–659 (2006)
Sanz, E. et al. Cell-type-specific isolation of ribosome-associated mRNA from complex tissues. Proc. Natl Acad. Sci. USA 106, 13939–13944 (2009)
Kubota, Y. et al. Selective coexpression of multiple chemical markers defines discrete populations of neocortical GABAergic neurons. Cereb. Cortex 21, 1803–1817 (2011)
Bondy, C. A. Transient IGF-I gene expression during the maturation of functionally related central projection neurons. J. Neurosci. 11, 3442–3455 (1991)
Liu, J. P., Baker, J., Perkins, A. S., Robertson, E. J. & Efstratiadis, A. Mice carrying null mutations of the genes encoding insulin-like growth factor I (Igf-1) and type 1 IGF receptor (Igf1r). Cell 75, 59–72 (1993)
Cheng, C. M. et al. Insulin-like growth factor 1 is essential for normal dendritic growth. J. Neurosci. Res. 73, 1–9 (2003)
Cao, P., Maximov, A. & Südhof, T. C. Activity-dependent IGF-1 exocytosis is controlled by the Ca2+-sensor synaptotagmin-10. Cell 145, 300–311 (2011)
Nishijima, T. et al. Neuronal activity drives localized blood-brain-barrier transport of serum insulin-like growth factor-I into the CNS. Neuron 67, 834–846 (2010)
Liu, J. L. et al. Insulin-like growth factor-I affects perinatal lethality and postnatal development in a gene dosage-dependent manner: manipulation using the Cre/loxP system in transgenic mice. Mol. Endocrinol. 12, 1452–1462 (1998)
Hede, M. S. et al. E-peptides control bioavailability of IGF-1. PLoS ONE 7, e51152 (2012)
Lin, J. Y., Knutsen, P. M., Muller, A., Kleinfeld, D. & Tsien, R. Y. ReaChR: a red-shifted variant of channelrhodopsin enables deep transcranial optogenetic excitation. Nature Neurosci. 16, 1499–1508 (2013)
Fagiolini, M. & Hensch, T. K. Inhibitory threshold for critical-period activation in primary visual cortex. Nature 404, 183–186 (2000)
Davis, M. F. et al. Inhibitory neuron transplantation into adult visual cortex creates a new critical period that rescues impaired vision. Neuron 86, 1055–1066 (2015)
Fu, Y., Kaneko, M., Tang, Y., Alvarez-Buylla, A. & Stryker, M. P. A cortical disinhibitory circuit for enhancing adult plasticity. eLife 4, e05558 (2015)
Hong, E. J., McCord, A. E. & Greenberg, M. E. A biological function for the neuronal activity-dependent component of Bdnf transcription in the development of cortical inhibition. Neuron 60, 610–624 (2008)
Bloodgood, B. L., Sharma, N., Browne, H. A., Trepman, A. Z. & Greenberg, M. E. The activity-dependent transcription factor NPAS4 regulates domain-specific inhibition. Nature 503, 121–125 (2013)
Turrigiano, G. Too many cooks? Intrinsic and synaptic homeostatic mechanisms in cortical circuit refinement. Annu. Rev. Neurosci. 34, 89–103 (2011)
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001)
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nature Protocols 7, 562–578 (2012)
Rubinson, D. A. et al. A lentivirus-based system to functionally silence genes in primary mammalian cells, stem cells and transgenic mice by RNA interference. Nature Genet. 33, 401–406 (2003)
Schopman, N. C. T., Liu, Y. P., Konstantinova, P., ter Brake, O. & Berkhout, B. Optimization of shRNA inhibitors by variation of the terminal loop sequence. Antiviral Res. 86, 204–211 (2010)
Spiegel, I. et al. A central role for Necl4 (SynCAM4) in Schwann cell-axon interaction and myelination. Nature Neurosci. 10, 861–869 (2007)
Schaeren-Wiemers, N. & Gerfin-Moser, A. A single protocol to detect transcripts of various types and expression levels in neural tissue and cultured cells: in situ hybridization using digoxigenin-labelled cRNA probes. Histochemistry 100, 431–440 (1993)
Tiscornia, G., Singer, O. & Verma, I. M. Production and purification of lentiviral vectors. Nature Protocols 1, 241–245 (2006)
Gordon, J. A. & Stryker, M. P. Experience-dependent plasticity of binocular responses in the primary visual cortex of the mouse. J. Neurosci. 16, 3274–3286 (1996)
Durand, S. et al. NMDA receptor regulation prevents regression of visual cortical function in the absence of Mecp2. Neuron 76, 1078–1090 (2012)
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nature Neurosci. 13, 133–140 (2010)
Gorski, J. A. et al. Cortical excitatory neurons and glia, but not GABAergic neurons, are produced in the Emx1-expressing lineage. J. Neurosci. 22, 6309–6314 (2002)
Hippenmeyer, S. et al. A developmental switch in the response of DRG neurons to ETS transcription factor signaling. PLoS Biol. 3, e159 (2005)
Taniguchi, H. et al. A resource of Cre driver lines for genetic targeting of GABAergic neurons in cerebral cortex. Neuron 71, 995–1013 (2011)
Acknowledgements
We thank C. Chen for help with electrophysiology experiments, E. Griffith and T. Cherry for critical reading of the manuscript, P. Zhang for managing the mouse colony and the HMS Biopolymers Facility Next-Gen Sequencing Core for their expertise in constructing Seq libraries and sequencing of the library samples. The ReachR-tdTomato construct was a gift from J. Lin, and we thank M. Li for production of the ReachR-virus. H.A. is a New York Stem Cell Robertson Investigator. This work was funded by fellowships by the Human Frontiers Science Program and the Swiss National Science Foundation (I.S.) and the National Institute of Health grants R01 NS028829 and P01 NS047572 (M.E.G.).
Author information
Authors and Affiliations
Contributions
Experiments were designed by A.R.M., I.S. and M.E.G. Experiments were conducted and analysed by A.R.M., I.S., E.C., A.P., C.P.T., J.E.B., C.M.B. and D.A.H. Experiments were supervised by H.A., M.F. and M.E.G. The manuscript was prepared by A.R.M., I.S. and M.E.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Validation of the sensory stimulation protocol and the RiboTag-based cell-type-specific purification of mRNA.
a, Quantitative real-time PCR (qPCR) for known experience-regulated genes on RNA isolated from the visual cortex of mice that were dark-housed for varying durations (overnight, 3 days, 7 days or 14 days) and then either euthanized in the dark or exposed to light for 1, 3 or 7.5 h, and then euthanized. Data are normalized to the maximal value in each data set and represent the mean and standard error of four biological replicates. b, qPCR for known experience-regulated genes on RNA isolated from the visual cortex of mice that were dark-housed for 14 days and then either exposed to light for 1, 3 or 7.5 h (dark + light, black) or kept in the dark during these hours (dark + dark, red). All mice of a given time point were dissected in very close temporal proximity. Data are normalized to the maximal value in each data set and represent the mean and standard error of four biological replicates. c, qPCR for known cell-type-specific marker genes on RNA isolated from RiboTag mice expressing Cre in distinct neuronal subtypes. Data are normalized to the maximal value in each data set and represent the mean and standard error of three biological replicates. d, qPCR for known early-induced transcription factors on RNA isolated from RiboTag mice expressing Cre in distinct neuronal subtypes. Data are normalized to the maximal value in each data set and represent the mean and standard error of three biological replicates.
Extended Data Figure 2 Validation of the RiboTag-seq approach.
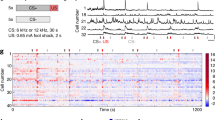
a, Matrix of Spearman correlation coefficients between biological replicates across all samples (scale of correlation coefficients extends from 0.7 to 1, see colour bar) (S, standard housing; 0, dark-housed only; 1/3/7.5, 1/3/7.5 h of light exposure after dark housing, respectively). b, Line plots of RNA-seq data showing the expression values (normalized reads across all exons of a gene) for cell-type-specific marker genes and ubiquitously expressed house-keeping genes in different Cre lines (Emx1-cre, red; Gad2-cre, blue; Pv-cre, brown; Sst-cre, green; Vip-cre, purple) across all time points of the experiment. Data represent the mean and standard error of three biological replicates.
Extended Data Figure 3 Characterization of the experience-induced gene programs in subtypes of cortical neurons.
a, Average matrix of Spearman correlation coefficients computed from the expression values of 1000 random sets of 602 genes (including experience-regulated genes, with replacement). b, Matrix of the Spearman correlation coefficients computed from the expression levels of control transcripts that match the expression distribution of experience-regulated genes (n = 602). c, Cladogram resulting from hierarchical clustering of all samples (except samples from Gad2-cre). Cladograms were computed using the mean expression values (that is, normalized reads across all exons of a gene) for all expressed transcripts (n = 14,280). d, Cladogram resulting from hierarchical clustering of the mean expression values of a set of control transcripts that match the expression distribution of experience-regulated genes (n = 602). e, Pie charts showing the subdivision of experience-regulated genes on the basis of kinetics in each set of Cre lines (red, rapidly induced; grey, induced with delayed kinetics; orange, induced only after two weeks of dark housing; green, rapidly suppressed; magenta, suppressed with delayed kinetics; blue, suppressed only after two weeks of dark housing). f, Left, matrix of Spearman correlation coefficients between Cre lines computed using the mean expression values (normalized reads across all exons of a gene) of early-induced genes one hour after light exposure. Right, matrix of Spearman correlation coefficients between Cre lines computed using the mean expression values of late-induced genes 7.5 h after light exposure. For each matrix, the correlations upon permuting the expression values are also shown (colour scale at right, scale begins at zero.)
Extended Data Figure 4 Characterization of cell-type-specific and experience-induced genes in subtypes of cortical neurons.
a, Scatter plots showing the log10 expression values for each expressed gene in a given Cre line (x axis) plotted against the maximum log10 expression values for that gene found in all other Cre lines (y axis). Black line denotes unity, and the red line is the 5.5-fold enrichment threshold set to include Vglut1 as a cell-type-specific gene in Emx1-cre. Data represent the mean values of three biological replicates. b, Scatter plots of all expressed genes, for each Cre line plotting the mean log2 fold enrichment in that Cre line (x axis) against the mean log2 of the absolute value of the maximum fold change observed in that Cre line. Data represent the mean values of three biological replicates. Genes that pass both enrichment and induction thresholds in 3 of 3 biological replicates are shown in red. c, Bar graph showing the maximum expression value (in normalized reads) for VIP-neuron-specific experience-regulated genes.
Extended Data Figure 5 Conditional knockout of Igf1 in VIP neurons does not affect cortical morphology or gross morphology of VIP neurons.
a, Example image of cortices from Igf1 wild-type (WT) (Vip-cre/+, LSL-tdTomato/+, Igf1WT/WT) or conditional-knockout (cKO) (Vip-cre/+, LSL-tdTomato/+, Igf1fl/fl) mice. VIP neurons are labelled in white, with DAPI shown in blue (cortical layers are indicated on the left, scale bar, 200 μm). b–d, Bar graphs showing the area of each cortical layer (b), number of VIP neurons per image per layer (c), or soma size of VIP neurons (d) in Igf1 wild-type (black) or conditional-knockout (red) mice. Data represent the mean and standard error of three biological replicates.
Extended Data Figure 6 Igf1 knockdown in VIP neurons affects inhibitory inputs onto VIP neurons but not onto neighbouring neurons.
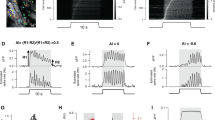
a, AAV shRNA constructs. shRNA cassettes against Igf1 or a control gene (Luc) were cloned downstream of the U6 promoter into an AAV vector that drives Cre-dependent expression of eGFP. b, qPCR validation of the efficacy of Igf1 shRNA constructs. Cultured cortical neurons were infected with lentiviral constructs either expressing no shRNA (vector only), a control shRNA (against Luc) or shRNAs against Igf1. Four days post-infection the cultures were quieted overnight with TTX and AP-5 and then harvested either before or after being depolarized with 55 mM KCl for 6 h; RNA was then isolated and qPCR was performed. Data are normalized to the maximal value in each replicate and represent the mean and standard error of three biological replicates. c, Bar graph showing normalized soma size of P21 visual cortex VIP neurons infected with control shRNA or shRNA targeting Igf1 (shRNA control, n = 103; shRNA Igf1, n = 174; P = 0.41, Mann–Whitney U-test). d, Bar graphs showing the levels of IGF1 in the serum of P20 mice that were injected intracortically with AAVs driving the expression of control shRNA (black) or Igf1 shRNA. Data represent the mean and s.e.m. of the serum IGF1 levels of four mice per group. e–l, Bar graphs and cumulative distribution plots showing mIPSC amplitudes and frequency/inter-event interval upon early widespread knockdown of Igf1 in VIP (e, f), SST (g, h), PV (i, j) and excitatory (k, l) neurons after injection of AAVs into P3 cortices of the respective Cre mice. VIP neurons (identified as eGFP-positive cells in Vip-cre mice): control and Igf1 shRNA, amplitude P = 0.96, frequency P = 0.04. SST neurons (identified as eGFP-positive cells in Sst-cre mice): control and Igf1 shRNA, amplitude P = 0.89, frequency P = 0.55. PV neurons (identified as eGFP-positive cells in Pv-cre mice): control and Igf1 shRNA, amplitude P = 0.084, frequency P = 0.93. Pyramidal neurons (identified by morphology): control and Igf1 shRNA, amplitude P = 0.84, frequency P = 0.15). All P values are derived from Mann–Whitney U-tests; numbers inside bars indicate the number of cells recorded.
Extended Data Figure 7 Effects of IGF1 overexpression in excitatory and SST-positive neurons.
a, RiboTag-seq identifies Igf1.4 as the major Igf1 isoform expressed in VIP neurons. Representative tracks of histograms of the RNA-seq reads in each Cre line across the Igf1 genomic locus. Data are from the 7.5 h light exposure RiboTag-seq data sets. b, AAV constructs for the Cre-dependent expression of HA-/Myc-tagged IGF1 (Igf1.4) and eGFP (that is, IGF1-OE, top) or of eGFP alone (that is, control, bottom). F2A, Furin cleavage site followed by the 2A peptide; black and white triangles represent a Cre-dependent Flex-switch. c, Western blot analysis of IGF1-receptor activation status in lysates of serum starved HEK293T cells that were stimulated with conditioned media (CM) containing epitope-tagged isoforms of IGF1. CM was produced by transfecting HEK293T cells with the respective construct and collecting the culture media. IGF1-receptor is detected with antibodies against either activated IGF1 receptor (anti-IGF1R pTyr1136/1138) or total IGF1 receptor (anti-IGF1R total). Molecular weight markers are on the right and the arrow indicates the band of the IGF1 receptor. d, e, Bar graphs and cumulative distribution plots showing mIPSC frequency/inter-event interval (d) and amplitude (e) of mIPSCs recorded from eGFP-positive neurons in P20 Sst-cre mice that were intracortically injected with AAVs driving the expression of control (C) or IGF1-OE (OE) constructs (amplitude, P = 0.16; frequency, P = 0.01; Mann–Whitney U-test; numbers inside bars indicate the number of cells recorded). f, g, Bar graphs and cumulative distribution plots showing mIPSC frequency/inter-event interval (f) and amplitude (g) of mIPSCs recorded from eGFP-positive neurons in P20 Emx1-cre mice that were intracortically injected with AAVs driving the expression of control (black, n = 20) or IGF1-OE AAVs (red, n = 20). Amplitude, P = 0.99; frequency, P = 0.01, Mann–Whitney U-test. h–j, Scatter plots of IGF1-interacting proteins showing the log10 normalized mean expression values in Vip-cre neurons versus each of the other Cre lines (Emx1 (h), Pv (i), Sst (j)). k, Quantification of EPSC paired-pulse recordings from VIP neurons infected with control shRNA- (black n = 9) or Igf1 shRNA- (red n = 11) expressing AAVs. The ratio of the second EPSC amplitude divided by the first EPSC amplitude is plotted against inter-stimulus interval (P = 0.1, two-way ANOVA).
Extended Data Figure 8 VIP neuron-derived IGF1 does not disrupt ocular dominance plasticity.
a, Widespread infection of VIP neurons by AAV-shRNA-hUbc-Flex-eGFP. High-titre injection of AAVs into the visual cortex of P18–20 Vip-cre/+ mice leads to infection of the majority VIP neurons (green, eGFP; red, anti-VIP; blue, DAPI; arrowheads, infected VIP neurons; scale bars, 150 μm, 20 μm in the inlet). b, c, Average of VEP traces recorded in the visual cortices of mice that were injected with AAVs expressing control shRNA (black/grey) or shRNA against Igf1 (red/pink) shRNA and that were (grey, pink) or were not (black, red) subjected to monocular deprivation in the eye contralateral to the recording site (MD versus NoMD, respectively). d, Monocular deprivation induces a significant reduction in the VEP amplitude in response to low spatial frequency stimulation in mice that had AAVs expressing control shRNA and Igf1 shRNA injected into their visual cortices (control shRNA NoMD, n = 5 mice; control shRNA MD, n = 7 mice; Igf1 shRNA NoMD, n = 5; Igf1 shRNA MD, n = 8. *P < 0.05, Mann–Whitney U-test). e, Mice that had AAVs expressing control shRNA (black) and Igf1 shRNA (red) injected into their visual cortices display normal ocular dominance plasticity as monocular deprivation (MD) induces a shift to the ispilateral eye in both groups (control shRNA NoMD, n = 5 mice; control shRNA MD, n = 7; Igf1 shRNA NoMD, n = 5 ; Igf1 shRNA MD, n = 8 mice. **P < 0.0001, one-way ANOVA with Tukey’s post hoc test).
Supplementary information
Supplementary Table 1
Table of expression values (normalized reads) of all expressed genes. (XLSX 11852 kb)
Supplementary Table 2
Tables of expression values (normalized reads) of all genes that are experience-regulated genes. (XLSX 1017 kb)
Supplementary Table 3
Tables of expression values (normalized reads) of all genes that are enriched in one cell-type. (XLSX 288 kb)
Supplementary Table 4
Tables of expression values (normalized reads) of all genes that are enriched in one cell-type and experience-regulated. (XLSX 41 kb)
Supplementary Table 5
Tables of top GO terms of all genes that are enriched in one cell-type and experience-regulated. (XLSX 8 kb)
Supplementary Tables 6 -8
This file comprises: (6) Table of primers used for quantitative PCR (qPCR); (7) Table of primers used for cloning of ISH riboprobes; (8) Table of AAV stocks used. (PDF 231 kb)
Rights and permissions
About this article
Cite this article
Mardinly, A., Spiegel, I., Patrizi, A. et al. Sensory experience regulates cortical inhibition by inducing IGF1 in VIP neurons. Nature 531, 371–375 (2016). https://doi.org/10.1038/nature17187
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17187
This article is cited by
-
Unique transcriptomes of sensory and non-sensory neurons: insights from Splicing Regulatory States
Molecular Systems Biology (2024)
-
Control of neuronal excitation–inhibition balance by BMP–SMAD1 signalling
Nature (2024)
-
Excitation–transcription coupling, neuronal gene expression and synaptic plasticity
Nature Reviews Neuroscience (2023)
-
The neurobiology of insulin-like growth factor I: From neuroprotection to modulation of brain states
Molecular Psychiatry (2023)
-
Aberrant maturation and connectivity of prefrontal cortex in schizophrenia—contribution of NMDA receptor development and hypofunction
Molecular Psychiatry (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.