Abstract
Arrestins are cytosolic proteins that regulate G-protein-coupled receptor (GPCR) desensitization, internalization, trafficking and signalling1,2. Arrestin recruitment uncouples GPCRs from heterotrimeric G proteins, and targets the proteins for internalization via clathrin-coated pits3,4. Arrestins also function as ligand-regulated scaffolds that recruit multiple non-G-protein effectors into GPCR-based ‘signalsomes’5,6. Although the dominant function(s) of arrestins vary between receptors, the mechanism whereby different GPCRs specify these divergent functions is unclear. Using a panel of intramolecular fluorescein arsenical hairpin (FlAsH) bioluminescence resonance energy transfer (BRET) reporters7 to monitor conformational changes in β-arrestin2, here we show that GPCRs impose distinctive arrestin ‘conformational signatures’ that reflect the stability of the receptor–arrestin complex and role of β-arrestin2 in activating or dampening downstream signalling events. The predictive value of these signatures extends to structurally distinct ligands activating the same GPCR, such that the innate properties of the ligand are reflected as changes in β-arrestin2 conformation. Our findings demonstrate that information about ligand–receptor conformation is encoded within the population average β-arrestin2 conformation, and provide insight into how different GPCRs can use a common effector for different purposes. This approach may have application in the characterization and development of functionally selective GPCR ligands8,9 and in identifying factors that dictate arrestin conformation and function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Ferguson, S. S. Evolving concepts in G protein-coupled receptor endocytosis: the role in receptor desensitization and signaling. Pharmacol. Rev. 53, 1–24 (2001)
Gurevich, V. V. & Gurevich, E. V. Structural determinants of arrestin functions. Prog. Mol. Biol. Transl. Sci. 118, 57–92 (2013)
Goodman, O. B., Jr et al. β-Arrestin acts as a clathrin adaptor in endocytosis of the β2-adrenergic receptor. Nature 383, 447–450 (1996)
Laporte, S. A. et al. The β2-adrenergic receptor/βarrestin complex recruits the clathrin adaptor AP-2 during endocytosis. Proc. Natl Acad. Sci. USA 96, 3712–3717 (1999)
Shenoy, S. K. & Lefkowitz, R. J. Angiotensin II-stimulated signaling through G proteins and β-arrestin. Sci. STKE 2005, cm14 (2005)
Luttrell, L. M. & Gesty-Palmer, D. Beyond desensitization: physiological relevance of arrestin-dependent signaling. Pharmacol. Rev. 62, 305–330 (2010)
Hoffmann, C. et al. A FlAsH-based FRET approach to determine G protein-coupled receptor activation in living cells. Nature Methods 2, 171–176 (2005)
Kenakin, T. Functional selectivity through protean and biased agonism: who steers the ship? Mol. Pharmacol. 72, 1393–1401 (2007)
Luttrell, L. M. Minireview : More than just a hammer: ligand “bias” and pharmaceutical discovery. Mol. Endocrinol. 28, 281–294 (2014)
Han, M. et al. Crystal structure of β-arrestin at 1.9 Å: possible mechanism of receptor binding and membrane translocation. Structure 9, 869–880 (2001)
Zhan, X. et al. Crystal structure of arrestin3 reveals the basis of the difference in receptor binding between two non-visual subtypes. J. Mol. Biol. 406, 467–478 (2011)
Shukla, A. K. et al. Structure of active β-arrestin-1 bound to a G-protein-coupled receptor phosphopeptide. Nature 497, 137–141 (2013)
Kim, Y. J. et al. Crystal structure of pre-activated arrestin p44. Nature 497, 142–146 (2013)
Kang, Y. et al. Crystal structure of rhodopsin bound to arrestin by femtosecond X-ray laser. Nature 523, 561–567 (2015)
Charest, P. G., Terrillon, S. & Bouvier, M. Monitoring agonist-promoted conformational changes of β-arrestin in living cells by intramolecular BRET. EMBO Rep. 6, 334–340 (2005)
Shukla, A. K. et al. Distinct conformational changes in β-arrestin report biased agonism at seven-transmembrane receptors. Proc. Natl Acad. Sci. USA 105, 9988–9993 (2008)
Oakley, R. H., Laporte, S. A., Holt, J. A., Caron, M. G. & Barak, L. S. Differential affinities of visual arrestin, βarrestin1, and βarrestin2 for G protein-coupled receptors delineate two major classes of receptors. J. Biol. Chem. 275, 17201–17210 (2000)
Appleton, K. M. et al. Biasing the parathyroid hormone receptor: relating in vitro ligand efficacy to in vivo biological activity. Methods Enzymol. 522, 229–262 (2013)
Barak, L. S., Ferguson, S. S., Zhang, J. & Caron, M. G. A β-arrestin/green fluorescent protein biosensor for detecting G protein-coupled receptor activation. J. Biol. Chem. 272, 27497–27500 (1997)
Wu, G. Krupnick, J. G., Benovic, J. L. & Lanier, S. M. Interaction of arrestins with intracellular domains of muscarinic and α2-adrenergic receptors. J. Biol. Chem. 272, 17836–17842 (1997)
DeFea, K. A. et al. β-Arrestin-dependent endocytosis of proteinase-activated receptor 2 is required for intracellular targeting of activated ERK1/2. J. Cell Biol. 148, 1267–1281 (2000)
Luttrell, L. M. et al. Activation and targeting of extracellular signal-regulated kinases by β-arrestin scaffolds. Proc. Natl Acad. Sci. USA 98, 2449–2454 (2001)
Zimmerman, B., Simaan, M., Lee, M.-H., Luttrell, L. M. & Laporte, S. A. c-Src-mediated phosphorylation of AP-2 reveals a general mechanism for receptors internalizing through the clathrin pathway. Cell. Signal. 21, 103–110 (2009)
Wei, H. et al. Independent β-arrestin 2 and G protein-mediated pathways for angiotensin II activation of extracellular signal-regulated kinases 1 and 2. Proc. Natl Acad. Sci. USA 100, 10782–10787 (2003)
Shenoy, S. K. et al. β-Arrestin-dependent, G protein-independent ERK1/2 activation by the β2 adrenergic receptor. J. Biol. Chem. 281, 1261–1273 (2006)
Oakley, R. H., Laporte, S. A., Holt, J. A., Barak, L. S. & Caron, M. G. Association of β-arrestin with G protein-coupled receptors during clathrin-mediated endocytosis dictates the profile of receptor resensitization. J. Biol. Chem. 274, 32248–32257 (1999)
Zimmerman, B. et al. Differential β-arrestin-dependent conformational signaling and cellular responses revealed by angiotensin analogs. Sci. Signal. 5, ra33 (2012)
Tobin, A. B., Butcher, A. J. & Kong, K. C. Location, location, location...site-specific GPCR phosphorylation offers a mechanism for cell-type-specific signalling. Trends Pharmacol. Sci. 29, 413–420 (2008)
Kommaddi, R. P. & Shenoy, S. K. Arrestins and protein ubiquitination. Prog. Mol. Biol. Transl. Sci. 118, 175–204 (2013)
Nuber, S. et al. β-Arrestin biosensors reveal a rapid, receptor-dependent activation/deactivation cycle. Nature http://dx.doi.org/10.1038/nature17198 (this issue)
Yon, J. & Fried, M. Precise gene fusion by PCR. Nucleic Acids Res. 17, 4895 (1989)
Binkowski, B. F., Fan, F. & Wood, K. V. Luminescent biosensors for real-time monitoring of intracellular cAMP. Methods Mol. Biol. 756, 263–271 (2011)
Leonard, A. P., Appleton, K. M., Luttrell, L. M. & Peterson, Y. K. A high-content, live-cell, and real-time approach to the quantitation of ligand-induced β-arrestin2 and class A/class B GPCR mobilization. Microsc. Microanal. 19, 150–170 (2013)
Wilson, P. C. et al. The arrestin-selective angiotensin AT1 receptor agonist [Sar1,Ile4,Ile8]-AngII negatively regulates bradykinin B2 receptor signaling via AT1-B2 receptor heterodimers. J. Biol. Chem. 288, 18872–18884 (2013)
Acknowledgements
This work was supported by National Institutes of Health grants DK055524 (L.M.L.) and GM095497 (L.M.L.), funds provided by Dialysis Clinics, Inc. (T.A.M), and the Research Service of the Charleston, SC Veterans Affairs Medical Center (L.M.L.). Supported by Canadian Institutes of Health Research operating grant MOP-74603 (S.A.L.). National Institutes of Health grant RR027777 (L.M.L.) supported the FLIPRTETRA facility. The contents of this article do not represent the views of the Department of Veterans Affairs or the United States Government.
Author information
Authors and Affiliations
Contributions
M.-H.L., K.A.M., E.G.S., J.Y.K, and S.A.L. performed experimental measurements and data analysis. T.A.M., Y.K.P., and S.A.L. provided technical expertise. M.-H.L. and L.M.L. conceived the project. All authors contributed to preparation of the manuscript and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
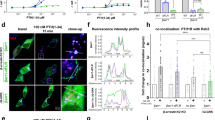
Extended Data Figure 1 Time-course and relationship of the β-arrestin2 intramolecular FlAsH BRET signal to receptor occupancy.
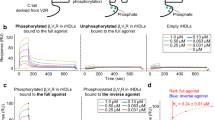
a, Time-course of AT1AR-induced changes in intramolecular FlAsH BRET. HEK293 cells were co-transfected with plasmid cDNA encoding AT1AR and the indicated rLuc–β-arrestin2–FlAsH reporter. Stimulations were carried out at a saturating concentration of AngII for the indicated times. The graph depicts the mean ± s.e.m. of independent biological replicates of ligand-induced Δnet BRET for each rLuc–β-arrestin2–FlAsH construct (n = 6). b, Ligand concentration dependence of PTH1R- and α1BAR-induced changes in intramolecular FlAsH BRET. HEK293 cells were co-transfected with plasmid cDNA encoding the PTH1R or α1BAR and the rLuc–β-arrestin2–FlAsH5 reporter. Stimulations were for 2 min using the indicated agonist concentration. The graph depicts the mean ± s.e.m. of independent biological replicates of ligand-induced Δnet BRET (n = 5). The EC50 for PTH(1–34) (PTH1R) and phenylephrine (α1BAR) were 30 nM and 80 nM, respectively. In all panels: *P < 0.05, #P < 0.005, greater or less than vehicle stimulated control.
Extended Data Figure 2 G-protein-coupling profiles of selected GPCRs.
a, Representative time-courses of cAMP luminescence following stimulation of HEK293 GloSensor cAMP cells transfected with each of six GPCRs. For the Gi/o-coupled S1P1R and α2AAR, stimulations were carried out in the presence of 10 μM forskolin to detect inhibition of adenylyl cyclase. Each panel depicts the agonist effect (green) compared to the control response to 10 μM forskolin (grey) measured in adjacent wells. Data are presented in relative luminescence units (RLU). b, Representative time-courses of intracellular calcium fluorescence following stimulation of HEK293 cells transfected with the same panel of GPCRs. Each panel depicts the agonist effect (blue) compared to the control response to the calcium ionophore A23187 (lavender) measured in adjacent wells. Data are presented in relative fluorescence units (RFU).
Extended Data Figure 3 Pertussis toxin sensitivity of ERK1/2 activation by Gi/o-coupled GPCRs.
HEK293 cells transfected with the β2AR, S1PR1 or α2AAR were serum-deprived overnight in the presence or absence of 1 ng ml−1 Bordetella pertussis toxin (PTX) before 5 min stimulation with isoproterenol, S1P or UK14303, respectively. Representative phospho-ERK1/2 immunoblots are shown above bar graphs depicting the mean ± s.e.m. of independent biological replicates (n = 5, β2AR, S1P1R and α2AAR). Responses were normalized to the basal level of phospho-ERK1/2 in non-stimulated samples. *P < 0.05, #P < 0.005, less than stimulated response in the absence of pertussis toxin.
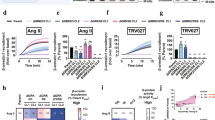
Extended Data Figure 4 Concentration-response relationship between FlAsH5 signal and arrestin-dependent ERK1/2 activation.
a, Relationship between α1BAR-induced change in FlAsH5 Δnet BRET and arrestin-dependent ERK1/2 activation at varying agonist concentration. The percent maximal phenylephrine-induced FlAsH5 Δnet BRET was determined in HEK293 cells transfected with α1BAR and rLuc–β-arrestin2–FlAsH5 expression plasmids (left). The concentration dependence of phenylephrine-stimulated ERK1/2 activation was determined in α1BAR-expressing HEK293 FRT/TO β-arrestin1/2 shRNA cells stimulated for 5 min (centre). β-arrestin1/2-dependent ERK1/2 activation was defined as the fold difference between agonist-stimulated ERK1/2 phosphorylation in the absence (total ERK1/2 signal) and presence (β-arrestin1/2-independent ERK1/2 signal) of doxycycline. A representative phospho-ERK1/2 immunoblot is shown above a graph depicting the mean ± s.e.m. of independent biological replicates (n = 4). EC50 for total ERK1/2, β-arrestin1/2-independent ERK1/2, and β-arrestin1/2-dependent ERK1/2 were 64 nM, 27 nM and 334 nM, respectively. Right, the relationship between percent maximal α1BAR-induced change in FlAsH5 Δnet BRET and β-arrestin1/2-dependent ERK1/2 activation over a range of agonist concentrations. In all panels, *P < 0.05, #P < 0.005, greater than nonstimulated. b, Relationship between GPCR-induced change in FlAsH5 Δnet BRET and arrestin-dependent ERK1/2 activation at saturating agonist concentration. The ligand-induced FlAsH5 Δnet BRET was determined in HEK293 cells transfected with the indicated GPCR and rLuc–β-arrestin2–FlAsH5 expression plasmids. The graph depicts the mean ± s.e.m. of independent biological replicates (n = 5).
Rights and permissions
About this article
Cite this article
Lee, MH., Appleton, K., Strungs, E. et al. The conformational signature of β-arrestin2 predicts its trafficking and signalling functions. Nature 531, 665–668 (2016). https://doi.org/10.1038/nature17154
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17154
This article is cited by
-
Molecular insights into intrinsic transducer-coupling bias in the CXCR4-CXCR7 system
Nature Communications (2023)
-
β-arrestin1 and 2 exhibit distinct phosphorylation-dependent conformations when coupling to the same GPCR in living cells
Nature Communications (2022)
-
Location bias contributes to functionally selective responses of biased CXCR3 agonists
Nature Communications (2022)
-
Structures of the ADGRG2–Gs complex in apo and ligand-bound forms
Nature Chemical Biology (2022)
-
How Carvedilol activates β2-adrenoceptors
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.