Abstract
Eukaryotic transcription activators stimulate the expression of specific sets of target genes through recruitment of co-activators such as the RNA polymerase II-interacting Mediator complex1,2. Aberrant function of transcription activators has been implicated in several diseases. However, therapeutic targeting efforts have been hampered by a lack of detailed molecular knowledge of the mechanisms of gene activation by disease-associated transcription activators. We previously identified an activator-targeted three-helix bundle KIX domain in the human MED15 Mediator subunit that is structurally conserved in Gal11/Med15 Mediator subunits in fungi3,4. The Gal11/Med15 KIX domain engages pleiotropic drug resistance transcription factor (Pdr1) orthologues, which are key regulators of the multidrug resistance pathway in Saccharomyces cerevisiae and in the clinically important human pathogen Candida glabrata5,6. The prevalence of C. glabrata is rising, partly owing to its low intrinsic susceptibility to azoles, the most widely used antifungal agent7,8. Drug-resistant clinical isolates of C. glabrata most commonly contain point mutations in Pdr1 that render it constitutively active9,10,11,12,13,14, suggesting that this transcriptional activation pathway represents a linchpin in C. glabrata multidrug resistance. Here we perform sequential biochemical and in vivo high-throughput screens to identify small-molecule inhibitors of the interaction of the C. glabrata Pdr1 activation domain with the C. glabrata Gal11A KIX domain. The lead compound (iKIX1) inhibits Pdr1-dependent gene activation and re-sensitizes drug-resistant C. glabrata to azole antifungals in vitro and in animal models for disseminated and urinary tract C. glabrata infection. Determining the NMR structure of the C. glabrata Gal11A KIX domain provides a detailed understanding of the molecular mechanism of Pdr1 gene activation and multidrug resistance inhibition by iKIX1. We have demonstrated the feasibility of small-molecule targeting of a transcription factor-binding site in Mediator as a novel therapeutic strategy in fungal infectious disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Protein Data Bank
Data deposits
Coordinates and NMR resonance assignments have been deposited in the Protein Data Bank (PDB code 4D7X) and Biological Magnetic Resonance Data Bank (BMRB code 25372). RNA-seq data have been deposited in the Gene Expression Omnibus (GEO) under accession GSE74361.
References
Conaway, R. C. & Conaway, J. W. Function and regulation of the Mediator complex. Curr. Opin. Genet. Dev. 21, 225–230 (2011)
Poss, Z. C., Ebmeier, C. C. & Taatjes, D. J. The Mediator complex and transcription regulation. Crit. Rev. Biochem. Mol. Biol. 48, 575–608 (2013)
Thakur, J. K. et al. A nuclear receptor-like pathway regulating multidrug resistance in fungi. Nature 452, 604–609 (2008)
Yang, F. et al. An ARC/Mediator subunit required for SREBP control of cholesterol and lipid homeostasis. Nature 442, 700–704 (2006)
Paul, S. & Moye-Rowley, W. S. Multidrug resistance in fungi: regulation of transporter-encoding gene expression. Front. Physiol. 5, 143 (2014)
Prasad, R. & Goffeau, A. Yeast ATP-binding cassette transporters conferring multidrug resistance. Annu. Rev. Microbiol. 66, 39–63 (2012)
Pfaller, M. A. et al. Variation in susceptibility of bloodstream isolates of Candida glabrata to fluconazole according to patient age and geographic location in the United States in 2001 to 2007. J. Clin. Microbiol. 47, 3185–3190 (2009)
Pfaller, M. A., Messer, S. A., Moet, G. J., Jones, R. N. & Castanheira, M. Candida bloodstream infections: comparison of species distribution and resistance to echinocandin and azole antifungal agents in Intensive Care Unit (ICU) and non-ICU settings in the SENTRY Antimicrobial Surveillance Program (2008–2009). Int. J. Antimicrob. Agents 38, 65–69 (2011)
Ferrari, S. et al. Gain of function mutations in CgPDR1 of Candida glabrata not only mediate antifungal resistance but also enhance virulence. PLoS Pathog. 5, e1000268 (2009)
Ferrari, S., Sanguinetti, M., Torelli, R., Posteraro, B. & Sanglard, D. Contribution of CgPDR1-regulated genes in enhanced virulence of azole-resistant Candida glabrata. PLoS One 6, e17589 (2011)
Silva, L. V. et al. Milbemycins: more than efflux inhibitors for fungal pathogens. Antimicrob. Agents Chemother. 57, 873–886 (2013)
Vermitsky, J. P. et al. Pdr1 regulates multidrug resistance in Candida glabrata: gene disruption and genome-wide expression studies. Mol. Microbiol. 61, 704–722 (2006)
Sanguinetti, M. et al. Mechanisms of azole resistance in clinical isolates of Candida glabrata collected during a hospital survey of antifungal resistance. Antimicrob. Agents Chemother. 49, 668–679 (2005)
Caudle, K. E. et al. Genomewide expression profile analysis of the Candida glabrata Pdr1 regulon. Eukaryot. Cell 10, 373–383 (2010)
Roehrl, M. H., Wang, J. Y. & Wagner, G. A general framework for development and data analysis of competitive high-throughput screens for small-molecule inhibitors of protein-protein interactions by fluorescence polarization. Biochemistry 43, 16056–16066 (2004)
DeRisi, J. et al. Genome microarray analysis of transcriptional activation in multidrug resistance yeast mutants. FEBS Lett. 470, 156–160 (2000)
Sanglard, D., Ischer, F., Calabrese, D., Majcherczyk, P. A. & Bille, J. The ATP binding cassette transporter gene CgCDR1 from Candida glabrata is involved in the resistance of clinical isolates to azole antifungal agents. Antimicrob. Agents Chemother. 43, 2753–2765 (1999)
Silva, L. V. et al. Milbemycins: more than efflux inhibitors for fungal pathogens. Antimicrob. Agents Chemother. 57, 873–886 (2012)
Arvanitis, M., Glavis-Bloom, J. & Mylonakis, E. Invertebrate models of fungal infection. Biochim. Biophys. Acta 1832, 1378–1383 (2013)
Vale-Silva, L., Ischer, F., Leibundgut-Landmann, S. & Sanglard, D. Gain-of-function mutations in PDR1, a regulator of antifungal drug resistance in Candida glabrata, control adherence to host cells. Infect. Immun. 81, 1709–1720 (2013)
Chen, Y. L. et al. Convergent evolution of calcineurin pathway roles in thermotolerance and virulence in Candida glabrata . G3 2, 675–691 (2012)
Farmakiotis, D., Tarrand, J. J. & Kontoyiannis, D. P. Drug-resistant Candida glabrata infection in cancer patients. Emerg. Infect. Dis. 20, 1833–1840 (2014)
Pfaller, M. A. et al. Frequency of decreased susceptibility and resistance to echinocandins among fluconazole-resistant bloodstream isolates of Candida glabrata. J. Clin. Microbiol. 50, 1199–1203 (2011)
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014)
EUCAST definitive document EDef 7.1: method for the determination of broth dilution MICs of antifungal agents for fermentative yeasts. Clin. Microbiol. Infect. 14, 398–405 (2008)
Krissinel, E. & Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D 60, 2256–2268 (2004)
Acknowledgements
We are grateful to P. Coote, E. Papadopoulos, R. Oh and R. E. Luna for helpful discussions and advice with data analysis and manuscript preparation. We acknowledge the ICCB-Longwood Screening Facility at Harvard Medical School for assistance with the high-throughput screens and access to the compound libraries, and the MGH Next Gen sequencing core for RNA-seq library construction. Mouse plasma and microsomal stability experiments were carried out at the Scripps Research Institute and iKIX1 pharmacokinetic parameters were assessed by Sai Life Sciences Limited. We acknowledge support from the National Institute of Health (grants GM047467 to G.W. and A.M.N. and EB002026 to G.W.). J.L.N. was supported by an NSERC fellowship.
Author information
Authors and Affiliations
Contributions
J.L.N., A.B., G.W., A.M.N. and H.A. conceived and designed the studies. A.B. and H.A. performed experiments relating to protein structure, small molecule screening and small molecule-protein interaction and data analysis. J.B. and G.M. performed the docking and free energy calculations. V.G., S.J.B. and N.S.G. designed the synthesis for iKIX1 and its analogues. J.L.N. performed the in vivo small molecule screen, luciferase, ChIP, transcription, efflux, spot plating, combination index and mammalian cell culture (HepG2) experiments. Y.-J.S. performed transcription and efflux experiments. J.L.N. prepared samples for RNA-seq analysis; bioinformatic analysis was carried out by F.J. and R.I.S.; L.A.V.-S. and D.S. designed and performed moth survival and adhesion assays. R.T., B.P. and M.S. designed and executed mouse fungal burden and UTI model studies. J.L.N., A.B., G.W., A.M.N. and H.A. wrote the manuscript with input from the team.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 List of compound libraries screened and top hits from the secondary in vivo viability screen.
Left, table of compound libraries that were screened using a fluorescence polarization assay at the Institute of Chemistry & Cell Biology (ICCB) facility at Harvard Medical School. Right, an S. cerevisiae viability screen identifies small molecules that preferentially inhibit growth of S. cerevisiae in a concentration-dependent manner in the presence of 5 μM ketoconazole. Top hits from the screen are shown; A600 nm values are the average of values from duplicate plates. Error bars denote mean ± s.d.
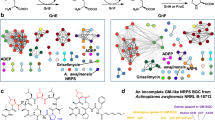
Extended Data Figure 2 Mapping the interaction interface of iKIX1 and CgPdr1 AD on the CgGal11A KIX domain and molecular details of iKIX1 interaction with the CgGal11A KIX domain based on targeted docking.
a, 2-dimensional representation of the H-bonding network between the CgGal11A KIX domain and iKIX1 based on docking studies. b, Chemical shift perturbations (CSPs) of ILV methyl resonances. Left, 1H-13C heteronuclear single quantum coherence (HSQC) spectrum showing ILV methyl resonances of CgGal11A KIX domain in presence (brown) and absence (teal) of CgPdr1 AD (twofold excess). Right, 1H-13C HSQC spectrum showing ILV methyl resonances of CgGal11A KIX domain in presence (purple) and in absence (teal) of iKIX1 (fourfold excess). Three leucines (L19, L23, L51) show significant CSPs in both spectra. c, Sequence alignment of the C. glabrata Gal11A and S. cerevisiae Gal11/Med15 KIX domains24.
Extended Data Figure 3 iKIX1 blocks Gal11/Med15 recruitment and upregulation of Pdr1 target genes.
a, iKIX1 prevents the ketoconazole-induced recruitment of ScGal11/Med15/Mediator to the upstream activating sequences (UAS) of the PDRE-regulated promoter ScSNQ2 and transcriptional upregulation of ScSNQ2. b, Haemagglutinin (HA)–tagged Pdr1 occupies PDRE-regulated promoters of ScPDR5 and ScSNQ2 in the presence of 20 μM iKIX1 or vehicle (DMSO) control before and following ketoconazole addition. c, 20 μM iKIX1 inhibits ketoconazole-induced upregulation of ScPdr1 target genes ScPDR5 and ScSNQ2 in the HA–Pdr1 strain. RNA was harvested concurrently with representative chromatin immunoprecipitation experiment shown in b at t = 0 min (DMSO, 20 μM iKIX1) and t = 15 min after ketoconazole induction (DMSO + KET, 20 μM iKIX1 + KET). Transcripts are normalized to ScSCR1 and un-induced DMSO control. a–c, Representative experiment from two biological replicates is shown. Error bars represent mean ±s.d. of technical replicates; *P < 0.05, **P < 0.01 and ***P < 0.001 as calculated by two-tailed Student’s t-test. d, RNA-seq analysis of a wild-type S. cerevisiae strain (BY4741) pre-treated with iKIX1 or vehicle alone then induced with ketoconazole (iKIX1 + KET and KET, respectively) demonstrate a blunted induction of Pdr1 target genes following iKIX1 pre-treatment. Data represent means of three biological replicates. e, iKIX1 pre-treatment does not significantly alter the transcript levels of PDR1 or GAL11/GAL11A in S. cerevisiae or C. glabrata after azole induction. Cells were pre-incubated with vehicle (DMSO) or iKIX1 and then induced with 40 μM ketoconazole (+KET) for 15 min before harvest. Average value of three biological replicates is shown and error bars represent mean ±s.d.; *P < 0.05, **P < 0.001 as compared to DMSO or DMSO+KET control, calculated by two-tailed Student’s t-test.
Extended Data Figure 4 iKIX1 pre-treatment durably represses azole-induced CgPdr1-dependent transcription but not basal levels of CgPdr1-dependent transcripts.
a, b, With iKIX1 pre-treatment, CgPdr1-dependent transcription of (a) CgCDR1 and (b) CgYOR1 remains repressed 120 min after ketoconazole induction. SFY114 (PDR1 wild-type) cells were pre-incubated with vehicle (DMSO) or iKIX1 and then induced with 40 μM ketoconazole (+KET). Transcript levels were assessed by quantitative RT–PCR before and for 120 min following ketoconazole induction. Transcript levels are normalized to CgRDN25-1 and un-induced vehicle control (DMSO) at t = 0. c, iKIX1 treatment alone does not have significant effects on CgPdr1 target gene induction either in the presence of wild-type (SFY114) or gain-of-function mutant CgPDR1 (amino acid alterations indicated). d, Table of average CgCDR1 delta Cp values (CpCgCDR1 – CpCgRDN25-1) and corresponding standard deviation for quantitative real-time PCR experiments shown in Fig. 3 f and Extended Data Fig. 4c. a–d, For all panels of Extended Data Fig. 4, average value of three biological replicates is shown and error bars represent ±s.d.; *P < 0.05, **P < 0.01, and ***P < 0.005 as compared to vehicle + ketoconazole control. P values calculated using two-tailed Student’s t-test.
Extended Data Figure 5 iKIX1 inhibits efflux of rhodamine 6 G in PDR1 wild-type, PDR1L280F and PDR1Y584C strains.
Data points indicate mean of three biological replicates and error bars represent mean ±s.d.
Extended Data Figure 6 iKIX1 increases the sensitivity of Cg strains bearing wild-type CgPDR1 to azole treatment and synergistically enhances azole sensitivity in a CgPDR1L280F mutant strain.
a, b, Two strains bearing wild-type CgPDR1 alleles (SFY114, DSY759) were plated at concentrations differing by tenfold (10×, 1×) on plates containing increasing concentrations of iKIX1 to 300 μM in the presence or absence of 1 μM ketoconazole (a) or iKIX1 to 250 μM in the presence or absence of 50 μM fluconazole (b). c, iKIX1 and ketoconazole have additive effects on the growth of a CgPDR1 wild-type strain. d, iKIX1 and ketoconazole synergistically inhibit the growth of the CgPDR1L280F mutant. c, d, The EUCAST broth microdilution method25 was used to assess the effects of iKIX1 and ketoconazole combination treatment. Growth, as assessed by A540 nm, was normalized to no drug control. All combination indices (CI) for the CgPDR1L280F mutant were less than 1, indicating synergy. A representative of three biological replicates is shown and the red line indicates a combination index of 1.
Extended Data Figure 7 iKIX1 exhibits favourable drug-like properties and an inactive analogue highlights the importance of the electron withdrawing chloride groups on the aromatic ring of iKIX1.
a–d, A structurally similar iKIX1 analogue (A2) lacking electron-withdrawing groups increases the IC50 in the fluorescence polarization assay (a) and abolishes activity in the S. cerevisiae luciferase reporter assay (b), repression of CgCDR1 expression (c), and synergistic C. glabrata cell growth inhibition with azoles (d). Error bars in b, c indicate mean ±s.d. of technical replicates (reads/real-time PCR reactions, respectively). **P < 0.005; statistical differences calculated using two-tailed Student’s t-test. e, iKIX1 inhibits viability of HepG2 cells at concentrations >50 μM. The mean of 3 biological replicates is shown; error bars represent means ±s.d. f, iKIX1 exhibits no effect on transcription of SREBP-target genes in HepG2 cells at concentrations up to 100 μM. Biological duplicates were assessed; representative experiment is shown and error bars represent means ±s.d. of technical (real-time PCR) replicates. g, Mouse plasma stability of iKIX1 and mouse and human microsomal stability of iKIX1, n = 1. h, In vivo pharmacokinetic parameters of iKIX1, n = 3 mice per time point.
Extended Data Figure 8 iKIX1-fluconazole combination treatment reduces fungal tissue burden by CgPDR1+ or CgPDR1 gain-of-function mutant strains in disseminated disease models; iKIX1 alone reduces adherence and fungal burden in a UTI model.
a, Clinical isolates DSY562/DSY565 (azole sensitive and PDR1L280F azole-resistant strains, respectively) behave similarly to SFY114/SFY115 (isogenic PDR1+ and PDR1L280F strains, shown in Fig. 4d) in the mouse infection model. n = 10 mice for each treatment condition; *P < 0.01, **P < 0.005 and ***P < 0.0001. b, iKIX1 combination treatment with fluconazole reduces fungal tissue burdens in the spleen or kidney of mice injected with C. glabrata PDR1P822L (SFY116). n = 5 mice for each treatment condition; **P < 0.01 and *P < 0.05. c, 100 mg kg−1 day−1 iKIX1 (high iKIX1) treatment of mice infected with SFY93 (pdr1Δ) significantly reduces fungal burden in a mouse infection model (colony forming units (CFU) per g of kidney) alone as compared to SFY114 (PDR1+) or SFY115 (PDR1L280F). n = 10 mice for each treatment condition; *P < 0.01, **P < 0.005, ***P < 0.0001. d, Mice infected with SFY114 (PDR1+), SFY115 (PDR1L280F) or SFY93 (pdr1Δ) were treated with low (10 mg kg−1 day−1) iKIX1, low fluconazole (low FLU; 25 mg kg−1 day−1), fluconazole at 100 mg kg−1 day−1 or combination with the two. iKIX1 did not confer additional reductions in CFU per g of kidney with SFY93 infection. n = 10 mice for each treatment condition. ***P < 0.0005. e, iKIX1 and ketoconazole reduce adherence of CgPDR1L280F (SFY115) to CHO-Lec2 cells. Adherence is normalized to SFY114 DMSO control; each column represents the average of 4 biological replicates. *P < 0.05 as compared to SFY114 DMSO control. f, iKIX1 (100 mg kg−1 day−1) or fluconazole significantly reduces fungal burden in the bladder and kidney in a urinary tract infection model in mice. n = 15 mice were infected in each group and points at 0 log10 CFU per g of organ fell below the detection limit of the method (50 CFU per g of organ). *P < 0.05, **P < 0.005. a–f, Statistical differences were measured using a Mann–Whitney/Wilcoxon rank-sum test as compared to no treatment control; error bars represent means ±s.d.
Extended Data Figure 9 Model of iKIX1 function.
iKIX1 functions as a co-therapeutic in combination with an azole, blocking the azole-induced recruitment of Gal11/Med15-Mediator to Pdr1 target genes upon azole-treatment and preventing the upregulation of Pdr1 target genes, including those which encode drug efflux pumps. This figure has been adapted from ref. 3.
Supplementary information
Supplementary Information
This file contains Supplementary Methods and Supplementary References. (PDF 423 kb)
Supplementary Table 1
This file contains RNA-Seq data for C. glabrata as RPKM (reads per kilobase per million mapped reads); key CgPdr1 targets are in bold. Genes are ranked by log2 fold change of expression between KET and iKIX1/KET treated cells (column logFC). (PDF 671 kb)
Supplementary Table 2
This file contains RNA-Seq data for S. cerevisiae as RPKM (reads per kilobase per million mapped reads); key ScPdr1 targets are in bold. Genes are ranked by log2 fold change of expression between KET and iKIX1/KET treated cells (column logFC). (PDF 836 kb)
Supplementary Table 3
This file contains gene ontology (GO) analysis of differentially expressed genes in S. cerevisiae upon iKIX1 treatment. Oxidative stress response gene expression is prominently elevated. (PDF 42 kb)
Supplementary Tables 4-6
This file contains Supplementary Tables 4-6. Supplementary Table 4 contains a list of strains used in the study, Supplementary Table 5 contains a list of plasmids used in the study and Supplementary Table 6 contains a list of primers used in the study. (XLSX 47 kb)
Rights and permissions
About this article
Cite this article
Nishikawa, J., Boeszoermenyi, A., Vale-Silva, L. et al. Inhibiting fungal multidrug resistance by disrupting an activator–Mediator interaction. Nature 530, 485–489 (2016). https://doi.org/10.1038/nature16963
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16963
This article is cited by
-
Molecular mechanisms governing antifungal drug resistance
npj Antimicrobials and Resistance (2023)
-
Sterol homeostasis in yeast
Nature Chemical Biology (2022)
-
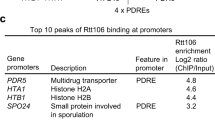
SWI/SNF and the histone chaperone Rtt106 drive expression of the Pleiotropic Drug Resistance network genes
Nature Communications (2022)
-
The Mediator complex as a master regulator of transcription by RNA polymerase II
Nature Reviews Molecular Cell Biology (2022)
-
Transcriptional regulation of the caspofungin-induced cell wall damage response in Candida albicans
Current Genetics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.