Abstract
U4/U6.U5 tri-snRNP represents a substantial part of the spliceosome before activation. A cryo-electron microscopy structure of Saccharomyces cerevisiae U4/U6.U5 tri-snRNP at 3.7 Å resolution led to an essentially complete atomic model comprising 30 proteins plus U4/U6 and U5 small nuclear RNAs (snRNAs). The structure reveals striking interweaving interactions of the protein and RNA components, including extended polypeptides penetrating into subunit interfaces. The invariant ACAGAGA sequence of U6 snRNA, which base-pairs with the 5′-splice site during catalytic activation, forms a hairpin stabilized by Dib1 and Prp8 while the adjacent nucleotides interact with the exon binding loop 1 of U5 snRNA. Snu114 harbours GTP, but its putative catalytic histidine is held away from the γ-phosphate by hydrogen bonding to a tyrosine in the amino-terminal domain of Prp8. Mutation of this histidine to alanine has no detectable effect on yeast growth. The structure provides important new insights into the spliceosome activation process leading to the formation of the catalytic centre.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Electron Microscopy Data Bank
Protein Data Bank
Data deposits
The cryo-EM maps have been deposited in the Electron Microscopy Data Bank with accession codes EMD-8006, EMD-8007, EMD-8008, EMD-8009, EMD-8010, EMD-8011, EMD-8012, EMD-8013 and EMD-8014. The coordinates of the atomic models have been deposited in the Protein Data Bank under accession codes 5GAN (overall), 5GAP (body domain), 5GAO (head domain) and 5GAM (foot domain).
Change history
17 February 2016
The Electron Microscopy Data Bank accession codes were listed in full, rather than as a range.
References
Will, C. L. & Lührmann, R. Spliceosome structure and function. Cold Spring Harb. Perspect. Biol. 3, a003707 (2011)
Chen, C. H. et al. Functional and physical interactions between components of the Prp19p-associated complex. Nucleic Acids Res. 30, 1029–1037 (2002)
Fabrizio, P. et al. The evolutionarily conserved core design of the catalytic activation step of the yeast spliceosome. Mol. Cell 36, 593–608 (2009)
Ohrt, T. et al. Molecular dissection of step 2 catalysis of yeast pre-m RNA splicing investigated in a purified system. RNA 19, 902–915 (2013)
Staley, J. P. & Guthrie, C. An RNA switch at the 5′ splice site requires ATP and the DEAD box protein Prp28p. Mol. Cell 3, 55–64 (1999)
Raghunathan, P. L. & Guthrie, C. RNA unwinding in U4/U6 snRNPs requires ATP hydrolysis and the DEIH-box splicing factor Brr2. Curr. Biol. 8, 847–855 (1998)
Laggerbauer, B., Achsel, T. & Lührmann, R. The human U5-200kD DEXH-box protein unwinds U4/U6 RNA duplices in vitro. Proc. Natl Acad. Sci. USA 95, 4188–4192 (1998)
Fica, S. M. et al. RNA catalyses nuclear pre-mRNA splicing. Nature 503, 229–234 (2013)
Madhani, H. D. & Guthrie, C. Dynamic RNA-RNA interactions in the spliceosome. Annu. Rev. Genet. 28, 1–26 (1994)
Toor, N., Keating, K. S., Taylor, S. D. & Pyle, A. M. Crystal structure of a self-spliced group II intron. Science 320, 77–82 (2008)
Fica, S. M., Mefford, M. A., Piccirilli, J. A. & Staley, J. P. Evidence for a group II intron-like catalytic triplex in the spliceosome. Nature Struct. Mol. Biol. 21, 464–471 (2014)
Newman, A. J. & Norman, C. U5 snRNA interacts with exon sequences at 5′ and 3′ splice sites. Cell 68, 743–754 (1992)
Sontheimer, E. J. & Steitz, J. A. The U5 and U6 small nuclear RNAs as active site components of the spliceosome. Science 262, 1989–1996 (1993)
Stark, H. & Lührmann, R. Cryo-electron microscopy of spliceosomal components. Annu. Rev. Biophys. Biomol. Struct. 35, 435–457 (2006)
Bai, X. C., McMullan, G. & Scheres, S. H. How cryo-EM is revolutionizing structural biology. Trends Biochem. Sci. 40, 49–57 (2015)
Nguyen, T. H. et al. The architecture of the spliceosomal U4/U6.U5 tri-snRNP. Nature 523, 47–52 (2015)
Ohi, M. D., Ren, L., Wall, J. S., Gould, K. L. & Walz, T. Structural characterization of the fission yeast U5.U2/U6 spliceosome complex. Proc. Natl Acad. Sci. USA 104, 3195–3200 (2007)
Chen, W. et al. Endogenous U2·U5·U6 snRNA complexes in S. pombe are intron lariat spliceosomes. RNA 20, 308–320 (2014)
Yan, C. et al. Structure of a yeast spliceosome at 3.6-angstrom resolution. Science 349, 1182–1191 (2015)
Hang, J., Wan, R., Yan, C. & Shi, Y. Structural basis of pre-mRNA splicing. Science 349, 1191–1198 (2015)
Stevens, S. W. et al. Biochemical and genetic analyses of the U5, U6, and U4/U6 x U5 small nuclear ribonucleoproteins from Saccharomyces cerevisiae. RNA 7, 1543–1553 (2001)
Gottschalk, A. et al. Identification by mass spectrometry and functional analysis of novel proteins of the yeast [U4/U6.U5] tri-snRNP. EMBO J. 18, 4535–4548 (1999)
Bai, X.-C., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. W. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, e11182 (2015)
Galej, W. P., Oubridge, C., Newman, A. J. & Nagai, K. Crystal structure of Prp8 reveals active site cavity of the spliceosome. Nature 493, 638–643 (2013)
Reuter, K., Nottrott, S., Fabrizio, P., Lührmann, R. & Ficner, R. Identification, characterization and crystal structure analysis of the human spliceosomal U5 snRNP-specific 15 kD protein. J. Mol. Biol. 294, 515–525 (1999)
Fabrizio, P., Laggerbauer, B., Lauber, J., Lane, W. S. & Lührmann, R. An evolutionarily conserved U5 snRNP-specific protein is a GTP-binding factor closely related to the ribosomal translocase EF-2. EMBO J. 16, 4092–4106 (1997)
Jørgensen, R. et al. Two crystal structures demonstrate large conformational changes in the eukaryotic ribosomal translocase. Nature Struct. Biol. 10, 379–385 (2003)
Wittinghofer, A. & Vetter, I. R. Structure-function relationships of the G domain, a canonical switch motif. Annu. Rev. Biochem. 80, 943–971 (2011)
Tourigny, D. S., Fernández, I. S., Kelley, A. C. & Ramakrishnan, V. Elongation factor G bound to the ribosome in an intermediate state of translocation. Science 340, 1235490 (2013)
Lin, J., Gagnon, M. G., Bulkley, D. & Steitz, T. A. Conformational changes of elongation factor G on the ribosome during tRNA translocation. Cell 160, 219–227 (2015)
Brenner, T. J. & Guthrie, C. Genetic analysis reveals a role for the C terminus of the Saccharomyces cerevisiae GTPase Snu114 during spliceosome activation. Genetics 170, 1063–1080 (2005)
Maracci, C., Peske, F., Dannies, E., Pohl, C. & Rodnina, M. V. Ribosome-induced tuning of GTP hydrolysis by a translational GTPase. Proc. Natl Acad. Sci. USA 111, 14418–14423 (2014)
Small, E. C., Leggett, S. R., Winans, A. A. & Staley, J. P. The EF-G-like GTPase Snu114p regulates spliceosome dynamics mediated by Brr2p, a DExD/H box ATPase. Mol. Cell 23, 389–399 (2006)
Bartels, C., Urlaub, H., Lührmann, R. & Fabrizio, P. Mutagenesis suggests several roles of Snu114p in pre-mRNA splicing. J. Biol. Chem. 278, 28324–28334 (2003)
Brenner, T. J. & Guthrie, C. Assembly of Snu114 into U5 snRNP requires Prp8 and a functional GTPase domain. RNA 12, 862–871 (2006)
Liu, S. et al. Binding of the human Prp31 Nop domain to a composite RNA-protein platform in U4 snRNP. Science 316, 115–120 (2007)
Ayadi, L. et al. Functional and structural characterization of the Prp3 binding domain of the yeast Prp4 splicing factor. J. Mol. Biol. 284, 673–687 (1998)
Liu, S. et al. A composite double-/single-stranded RNA-binding region in protein Prp3 supports tri-snRNP stability and splicing. eLife 4, e07320 (2015)
Nottrott, S., Urlaub, H. & Lührmann, R. Hierarchical, clustered protein interactions with U4/U6 snRNA: a biochemical role for U4/U6 proteins. EMBO J. 21, 5527–5538 (2002)
Makarov, E. M., Makarova, O. V., Achsel, T. & Lührmann, R. The human homologue of the yeast splicing factor prp6p contains multiple TPR elements and is stably associated with the U5 snRNP via protein-protein interactions. J. Mol. Biol. 298, 567–575 (2000)
Galisson, F. & Legrain, P. The biochemical defects of prp4-1 and prp6-1 yeast splicing mutants reveal that the PRP6 protein is required for the accumulation of the [U4/U6.U5] tri-snRNP. Nucleic Acids Res. 21, 1555–1562 (1993)
Nguyen, T. H. D. et al. Structural basis of Brr2-Prp8 interactions and implications for U5 snRNP biogenesis and the spliceosome active site. Structure 21, 910–919 (2013)
Hahn, D., Kudla, G., Tollervey, D. & Beggs, J. D. Brr2p-mediated conformational rearrangements in the spliceosome during activation and substrate repositioning. Genes Dev. 26, 2408–2421 (2012)
van Nues, R. W. & Beggs, J. D. Functional contacts with a range of splicing proteins suggest a central role for Brr2p in the dynamic control of the order of events in spliceosomes of Saccharomyces cerevisiae. Genetics 157, 1451–1467 (2001)
Wolf, E. et al. Exon, intron and splice site locations in the spliceosomal B complex. EMBO J. 28, 2283–2292 (2009)
Boehringer, D. et al. Three-dimensional structure of a pre-catalytic human spliceosomal complex B. Nature Struct. Mol. Biol. 11, 463–468 (2004)
De, I. et al. The RNA helicase Aquarius exhibits structural adaptations mediating its recruitment to spliceosomes. Nature Struct. Mol. Biol. 22, 138–144 (2015)
Li, Z. & Brow, D. A. A spontaneous duplication in U6 spliceosomal RNA uncouples the early and late functions of the ACAGA element in vivo. RNA 2, 879–894 (1996)
Kuhn, A. N. & Brow, D. A. Suppressors of a cold-sensitive mutation in yeast U4 RNA define five domains in the splicing factor Prp8 that influence spliceosome activation. Genetics 155, 1667–1682 (2000)
Chan, S. P. & Cheng, S. C. The Prp19-associated complex is required for specifying interactions of U5 and U6 with pre-mRNA during spliceosome activation. J. Biol. Chem. 280, 31190–31199 (2005)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nature Methods 10, 584–590 (2013)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Scheres, S. H. A Bayesian view on cryo-EM structure determination. J. Mol. Biol. 415, 406–418 (2012)
Scheres, S. H. Beam-induced motion correction for sub-megadalton cryo-EM particles. eLife 3, e03665 (2014)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nature Methods 11, 63–65 (2014)
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nature Methods 9, 853–854 (2012)
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011)
Nicholls, R. A., Fischer, M., McNicholas, S. & Murshudov, G. N. Conformation-independent structural comparison of macromolecules with ProSMART. Acta Crystallogr. D 70, 2487–2499 (2014)
Brown, A. et al. Tools for macromolecular model building and refinement into electron cryo-microscopy reconstructions. Acta Crystallogr. D 71, 136–153 (2015)
Scheres, S. H. W., Núñez-Ramírez, R., Sorzano, C. O. S., Carazo, J. M. & Marabini, R. Image processing for electron microscopy single-particle analysis using XMIPP. Nature Protocols 3, 977–990 (2008)
Goddard, T. D., Huang, C. C. & Ferrin, T. E. Visualizing density maps with UCSF Chimera. J. Struct. Biol. 157, 281–287 (2007)
Kunkel, T. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Proc. Natl Acad. Sci. USA 82, 488–492 (1985)
Turner, I. A., Norman, C. M., Churcher, M. J. & Newman, A. J. Dissection of Prp8 protein defines multiple interactions with crucial RNA sequences in the catalytic core of the spliceosome. RNA 12, 375–386 (2006)
Frazer, L. N., Lovell, S. C. & O’Keefe, R. T. Analysis of synthetic lethality reveals genetic interactions between the GTPase Snu114p and snRNAs in the catalytic core of the Saccharomyces cerevisiae spliceosome. Genetics 183, 497–515 (2009)
Acknowledgements
We thank C. Savva, S. Chen, G. McMullan, J. Grimmett and T. Darling for running the electron microscopy and computing facilities, A. Brown, P. Emsley, G. Murshudov for advice and help with model building and refinement, R. O’Keefe for the ∆Snu114 yeast strain, the members of the spliceosome group for help and advice throughout the project and R. Leiro for help with data processing. We thank S. Fica for critical reading of the manuscript and J. Löwe, V. Ramakrishnan, and R. Henderson for their continuing support and encouragements. The project was supported by the Medical Research Council (MC_U105184330 to K.N. and MC_UP_A025_1013 to S.H.W.S.).
Author information
Authors and Affiliations
Contributions
T.H.D.N. purified yeast tri-snRNP and prepared EM grids, T.H.D.N., W.P.G. and X.-C.B collected all EM images. T.H.D.N. processed data and calculated the maps with the help of X.-C.B and S.H.W.S.; T.H.D.N., W.P.G. and C.O. built a model into the map and refined the structure. T.H.D.N. performed mutagenesis experiments with the help of A.J.N.; T.H.D.N. and W.P.G. prepared all illustrations. K.N. initiated and orchestrated the project. T.H.D.N., W.P.G. and K.N. analysed the structure and wrote the paper with invaluable contributions from all other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Image processing procedures.
a, Representative micrograph. b, Representative 2D class averages obtained from reference-free 2D classification. c, Classification and refinement procedures used in this study.
Extended Data Figure 2 Local and overall resolutions of tri-snRNP maps.
Local resolution estimation by Resmap55 of a, the overall 3.7 Å map and b, maps of the head, body and foot domains obtained from masked refinements with signal subtraction23. c, Gold-standard FSC curves for the overall map and the maps of the head, body and foot domains obtained from masked refinements. Their resolutions are estimated at FSC = 0.143. d–g, FSC curves of model versus map and cross-validation of model refinement by half-maps for the body, foot, head and overall maps, respectively. The red curves show FSC between the atomic model and the half-map it was refined against (half1) and the blue curves show FSC between the atomic model and the other half-map (half2) it was not refined against. The black curves show FSC between the atomic model and the sum map which the model was refined against.
Extended Data Figure 3 Representative EM density for different components of the map.
a, Snu114 in the Foot domain with a bound GTP (magenta). The inset shows the GTP-binding pocket. b, Brr2 in the head domain with a bound single-stranded region of U4 snRNA. The inset shows the density in the RNA binding tunnel. c, Density for Prp8 large and RNase-like domains. The inset shows the density in the core of Prp8. d–f, Prp3, Prp31 and Prp6 densities, respectively, with extended polypeptides.
Extended Data Figure 4 Secondary structure of the snRNAs in tri-snRNP.
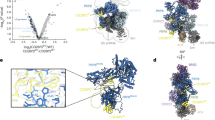
a, U4/U6 snRNA; c, U5 snRNA. The coloured nucleotides with red, green and blue background were built de novo into our EM density. The region near the ACAGAGA sequence of U6 snRNA forms a stem-loop that was not predicted previously. b, d, Representative EM density for U4/U6 snRNA duplex and U5 snRNA, respectively.
Extended Data Figure 5 Interactions of Snu114 with guanine nucleotides and the N-terminal domain of Prp8 in the S. cerevisiae U4/U6.U5 tri-snRNP and S. pombe ILS complexes.
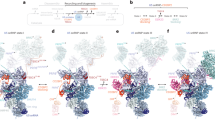
a, Conformation of the Snu114(Cwf10)-bound GDP refined in the S. pombe ILS spliceosomal complex19,20 (red, PDB 3JB9), was overlaid on GDPs found in other guanine-nucleotide binding proteins (grey, PDB coordinates: 1DAR, 2E1R, 2WRI, 1Z0I, 5CA8, 1XTQ, 4YLG, 1SF8, 5BXQ). b, Guanine nucleotide refined as GDP in Snu114 of the S. cerevisiae U4/U6.U5 tri-snRNP (blue) is overlaid on GDPs found in the PDB coordinates as in a. c, Conformation of guanine nucleotide refined as GTP in Snu114 of the S. cerevisiae U4/U6.U5 tri-snRNP (blue) agrees well with GTP or GTP analogues in other guanine-nucleotide binding proteins (PDB code: 2BV3, 2DY1, 2J7K, 4YW9, 1ASO, 1LF0 (grey)). d, Superposition of the active site of Snu114-GTP and Cwf10-GDP. e, Superposition of the GDP-bound EF-G (2WRI), GMP-PCP bound EF-G (4JUW) and Snu114 (S. cerevisiae tri-snRNP) active sites. His218 (His87 in EF-G) positions water molecule crucial for GTP hydrolysis. f, Comparison of Prp8N-term domain, Snu114 and U5 snRNA in the S. cerevisiae U4/U6.U5 complex and S. pombe ILS complex. g, Growth of serial dilutions of yeast strains carrying wild-type Snu114, His218Arg or His218Ala Snu114 mutants at different temperatures. Cells were spotted on YPD plates and grown at 14 °C for 10 days, 30 °C and 37 °C for 2 days. h, Growth of serial dilutions of yeast strains carrying wild-type Prp8, Tyr403Phe and Tyr403Ala mutants. Cells were spotted on YPD plates and grown at 14 °C for 9 days, 30 °C for 3 days. This yeast strain does not survive at 37 °C and thus is not shown.
Extended Data Figure 6 Conformational flexibility of tri-snRNP observed by classification.
a, Different conformations of the arm domain demonstrated by the unsharpened maps of the three major classes (purple, magenta and red) obtained from masked classification of the arm domain alone followed by masked refinement with the body and arm domains. The body domain was included in the refinement because the arm domain is too small for accurate alignments. b, The sharpened map of one of the three classes with Prp3 and LSm models shown. In the improved domain maps for the arm domain, extra density for the N-terminal helix of Prp3 could be observed to extend to the LSm proteins. c, The sharpened map of the tri-snRNP and the locations of Snu66 and Prp8. d, The open and closed conformations of the head and foot domains of the tri-snRNP observed by global classification. The unsharpened maps for the two major classes obtained from global classification with finer angular sampling (1.8°) followed by 3D auto-refinement are shown. The open and closed states are indicated. e, Superposition of the unsharpened maps of the open (grey) and closed (yellow) states shown in d. The arrows indicate the rotations of the head and foot domains.
Extended Data Figure 7 Brr2 helicase and its U4/U6 snRNA substrate.
a, Domain structure of Brr2 helicase comprising the N-terminal domain and two helicase cassettes. Individual domains of N-terminal helicase cassette (NHC) are colour-coded. b, Extensive interactions of Brr2 with U4/U6 snRNA and Prp3. The single-stranded region of U4 snRNA extending from stem I enters the active site near the β-finger (red). c, 3′ stem of U4 snRNA interacts with the HLH domain of NHC. d, The N-terminal domain (NTD) of Brr2 interacts with a long helix of Prp3 and inserts a loop into U4/U6 Stem II. e, Snu66 has a long extended region that wraps around both helicase cassettes of Brr2.
Supplementary information
Supplementary Data
This zipped file contains the pymol session file of the PDB coordinate. (ZIP 4054 kb)
Rights and permissions
About this article
Cite this article
Nguyen, T., Galej, W., Bai, Xc. et al. Cryo-EM structure of the yeast U4/U6.U5 tri-snRNP at 3.7 Å resolution. Nature 530, 298–302 (2016). https://doi.org/10.1038/nature16940
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16940
This article is cited by
-
3DFlex: determining structure and motion of flexible proteins from cryo-EM
Nature Methods (2023)
-
A multi-factor trafficking site on the spliceosome remodeling enzyme BRR2 recruits C9ORF78 to regulate alternative splicing
Nature Communications (2022)
-
Conserved protein Pir2ARS2 mediates gene repression through cryptic introns in lncRNAs
Nature Communications (2020)
-
Bottom-up structural proteomics: cryoEM of protein complexes enriched from the cellular milieu
Nature Methods (2020)
-
Structures of the human pre-catalytic spliceosome and its precursor spliceosome
Cell Research (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.