Abstract
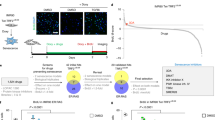
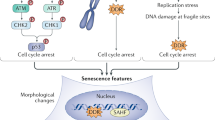
Cellular senescence, a stress-induced irreversible growth arrest often characterized by expression of p16Ink4a (encoded by the Ink4a/Arf locus, also known as Cdkn2a) and a distinctive secretory phenotype, prevents the proliferation of preneoplastic cells and has beneficial roles in tissue remodelling during embryogenesis and wound healing. Senescent cells accumulate in various tissues and organs over time, and have been speculated to have a role in ageing. To explore the physiological relevance and consequences of naturally occurring senescent cells, here we use a previously established transgene, INK-ATTAC, to induce apoptosis in p16Ink4a-expressing cells of wild-type mice by injection of AP20187 twice a week starting at one year of age. We show that compared to vehicle alone, AP20187 treatment extended median lifespan in both male and female mice of two distinct genetic backgrounds. The clearance of p16Ink4a-positive cells delayed tumorigenesis and attenuated age-related deterioration of several organs without apparent side effects, including kidney, heart and fat, where clearance preserved the functionality of glomeruli, cardio-protective KATP channels and adipocytes, respectively. Thus, p16Ink4a-positive cells that accumulate during adulthood negatively influence lifespan and promote age-dependent changes in several organs, and their therapeutic removal may be an attractive approach to extend healthy lifespan.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Campisi, J. Aging, cellular senescence, and cancer. Annu. Rev. Physiol. 75, 685–705 (2013)
Sharpless, N. E. & Sherr, C. J. Forging a signature of in vivo senescence. Nature Rev. Cancer 15, 397–408 (2015)
van Deursen, J. M. The role of senescent cells in ageing. Nature 509, 439–446 (2014)
Baker, D. J. et al. Clearance of p16Ink4a-positive senescent cells delays ageing-associated disorders. Nature 479, 232–236 (2011)
Baker, D. J. et al. Opposing roles for p16Ink4a and p19Arf in senescence and ageing caused by BubR1 insufficiency. Nature Cell Biol. 10, 825–836 (2008)
Jun, J. I. & Lau, L. F. The matricellular protein CCN1 induces fibroblast senescence and restricts fibrosis in cutaneous wound healing. Nature Cell Biol. 12, 676–685 (2010)
Krizhanovsky, V. et al. Senescence of activated stellate cells limits liver fibrosis. Cell 134, 657–667 (2008)
Demaria, M. et al. An essential role for senescent cells in optimal wound healing through secretion of PDGF-AA. Dev. Cell 31, 722–733 (2014)
Muñoz-Espin, D. et al. Programmed cell senescence during mammalian embryonic development. Cell 155, 1104–1118 (2013)
Storer, M. et al. Senescence is a developmental mechanism that contributes to embryonic growth and patterning. Cell 155, 1119–1130 (2013)
Wang, W., Wu, J., Zhang, Z. & Tong, T. Characterization of regulatory elements on the promoter region of p16INK4a that contribute to overexpression of p16 in senescent fibroblasts. J. Biol. Chem. 276, 48655–48661 (2001)
Tchkonia, T. et al. Fat tissue, aging, and cellular senescence. Aging Cell 9, 667–684 (2010)
Liu, Y. et al. Expression of p16INK4a in peripheral blood T-cells is a biomarker of human aging. Aging Cell 8, 439–448 (2009)
Wang, C., Li, Q., Redden, D. T., Weindruch, R. & Allison, D. B. Statistical methods for testing effects on “maximum lifespan”. Mech. Ageing Dev. 125, 629–632 (2004)
Goodrick, C. L., Ingram, D. K., Reynolds, M. A., Freeman, J. R. & Cider, N. Effects of intermittent feeding upon body weight and lifespan in inbred mice: interaction of genotype and age. Mech. Ageing Dev. 55, 69–87 (1990)
Blackwell, B. N., Bucci, T. J., Hart, R. W. & Turturro, A. Longevity, body weight, and neoplasia in ad libitum-fed and diet-restricted C57BL6 mice fed NIH-31 open formula diet. Toxicol. Pathol. 23, 570–582 (1995)
Bronson, R. T. & Lipman, R. D. Reduction in rate of occurrence of age related lesions in dietary restricted laboratory mice. Growth Dev. Aging 55, 169–184 (1991)
Conti, B. et al. Transgenic mice with a reduced core body temperature have an increased life span. Science 314, 825–828 (2006)
Coschigano, K. T. et al. Deletion, but not antagonism, of the mouse growth hormone receptor results in severely decreased body weights, insulin, and insulin-like growth factor I levels and increased life span. Endocrinology 144, 3799–3810 (2003)
Forster, M. J., Morris, P. & Sohal, R. S. Genotype and age influence the effect of caloric intake on mortality in mice. FASEB J. 17, 690–692 (2003)
Ikeno, Y. et al. Housing density does not influence the longevity effect of calorie restriction. J. Gerontol. A Biol. Sci. Med. Sci. 60, 1510–1517 (2005)
Kunstyr, I. & Leuenberger, H. G. Gerontological data of C57BL/6J mice. I. Sex differences in survival curves. J. Gerontol. 30, 157–162 (1975)
Selman, C. et al. Evidence for lifespan extension and delayed age-related biomarkers in insulin receptor substrate 1 null mice. FASEB J. 22, 807–818 (2008)
Sohal, R. S., Ku, H. H., Agarwal, S., Forster, M. J. & Lal, H. Oxidative damage, mitochondrial oxidant generation and antioxidant defenses during aging and in response to food restriction in the mouse. Mech. Ageing Dev. 74, 121–133 (1994)
Srivastava, V. K., Tilley, R. D., Hart, R. W. & Busbee, D. L. Effect of dietary restriction on the fidelity of DNA polymerases in aging mice. Exp. Gerontol. 26, 453–466 (1991)
Turturro, A. et al. Growth curves and survival characteristics of the animals used in the biomarkers of aging program. J. Gerontol. A Biol. Sci. Med. Sci. 54, B492–B501 (1999)
Yuan, R. et al. Genetic coregulation of age of female sexual maturation and lifespan through circulating IGF1 among inbred mouse strains. Proc. Natl Acad. Sci. USA 109, 8224–8229 (2012)
Zhang, Y. et al. The starvation hormone, fibroblast growth factor-21, extends lifespan in mice. Elife 1, e00065 (2012)
Richardson, A. et al. Measures of healthspan as indices of aging in mice-a recommendation. J. Gerontol. A Biol. Sci. Med. Sci. http://dx.doi.org/10.1093/gerona/glv080 (2015)
Razzaque, M. S. Does renal ageing affect survival? Ageing Res. Rev. 6, 211–222 (2007)
Ferder, L. F., Inserra, F. & Basso, N. Effects of renin-angiotensin system blockade in the aging kidney. Exp. Gerontol. 38, 237–244 (2003)
Paul, M., Poyan Mehr, A. & Kreutz, R. Physiology of local renin-angiotensin systems. Physiol. Rev. 86, 747–803 (2006)
Bernhard, D. & Laufer, G. The aging cardiomyocyte: a mini-review. Gerontology 54, 24–31 (2008)
Sudhir, R., Sukhodub, A., Du, Q., Jovanovic, S. & Jovanovic, A. Ageing-induced decline in physical endurance in mice is associated with decrease in cardiac SUR2A and increase in cardiac susceptibility to metabolic stress: therapeutic prospects for up-regulation of SUR2A. Biogerontology 12, 147–155 (2011)
Baker, D. J. et al. Increased expression of BubR1 protects against aneuploidy and cancer and extends healthy lifespan. Nature Cell Biol. 15, 96–102 (2012)
Childs, B. G., Durik, M., Baker, D. J. & van Deursen, J. M. Cellular senescence in aging and age-related disease: from mechanisms to therapy. Nature Med. 21, 1424–1435 (2015)
Baker, D. J., Weaver, R. L. & van Deursen, J. M. p21 both attenuates and drives senescence and aging in BubR1 progeroid mice. Cell Rep. 3, 1164–1174 (2013)
Baker, D. J. et al. Increased expression of BubR1 protects against aneuploidy and cancer and extends healthy lifespan. Nature Cell Biol. 15, 96–102 (2013)
Zingman, L. V. et al. Kir6.2 is required for adaptation to stress. Proc. Natl Acad. Sci. USA 99, 13278–13283 (2002)
Wijshake, T. et al. Reduced life- and healthspan in mice carrying a mono-allelic BubR1 MVA mutation. PLoS Genet. 8, e1003138 (2012)
Buenz, E. J. et al. Apoptosis of hippocampal pyramidal neurons is virus independent in a mouse model of acute neurovirulent picornavirus infection. Am. J. Pathol. 175, 668–684 (2009)
Maskey, R. S. et al. Spartan deficiency causes genomic instability and progeroid phenotypes. Nature Commun. 5, 5744 (2014)
Blatner, N. R. et al. Expression of RORγt marks a pathogenic regulatory T cell subset in human colon cancer. Sci. Transl. Med. 4, 164ra159 (2012)
Baker, D. J. et al. BubR1 insufficiency causes early onset of aging-associated phenotypes and infertility in mice. Nature Genet. 36, 744–749 (2004)
Stollewerk, A., Klambt, C. & Cantera, R. Electron microscopic analysis of Drosophila midline glia during embryogenesis and larval development using β-galactosidase expression as endogenous cell marker. Microsc. Res. Tech. 35, 294–306 (1996)
Garelick, M. G. et al. Chronic rapamycin treatment or lack of S6K1 does not reduce ribosome activity in vivo. Cell Cycle 12, 2493–2504 (2013)
Schreiber, K. H. et al. Rapamycin-mediated mTORC2 inhibition is determined by the relative expression of FK506-binding proteins. Aging Cell 14, 265–273 (2015)
Baker, D. J. et al. BubR1 insufficiency causes early onset of aging-associated phenotypes and infertility in mice. Nature Genet. 36, 744–749 (2004)
Burd, C. E. et al. Monitoring tumorigenesis and senescence in vivo with a p16INK4a-luciferase model. Cell 152, 340–351 (2013)
Ricke, R. M., Jeganathan, K. B., Malureanu, L., Harrison, A. M. & van Deursen, J. M. Bub1 kinase activity drives error correction and mitotic checkpoint control but not tumor suppression. J. Cell Biol. 199, 931–949 (2012)
Acknowledgements
We thank M. Hamada, J. Rainey, Q. Guo, S. Bornschlegl, M. Li, C. M. Roos, N. Hamada and B. Zhang for assistance, and C. Burd, S. Reyes Ramirez, P. Galardy and D. Katzmann and the members of Program Project Grant AG041122 for discussions. We are grateful to R. Miller and S. Austad for help with the design and interpretation of our lifespan studies. We thank C. Burd and J. Campisi for sharing p16-FLUC and 3MR MEFs, respectively. This work was supported by the National Institutes of Health (J.M.v.D. R01CA96985 and AG041122 project 1) and (J.D.M., HL111121), the Paul F. Glenn Foundation (J.M.v.D. and D.J.B.), the Ellison Medical Foundation (D.J.B.), the Noaber Foundation (J.M.v.D.), the Children’s Research Center (D.J.B.) and the Robert and Arlene Kogod Center on Aging (D.J.B.).
Author information
Authors and Affiliations
Contributions
D.J.B. performed all lifespan and most healthspan analyses on ATTAC mice. B.G.C. designed and conducted experiments to identify and quantify X-gal-positive cells by TEM, and analysed mice for cardiomyocyte hypertrophy and local RAAS activity in kidney. M.E.W., J.Z. and R.A.S. assisted with various aspects of healthspan analyses: the extent of their contributions is reflected in the authorship order. C.J.S. conducted the lifespan analysis of C57BL/6-129Sv/E hybrid mice on diets containing 5% or 9% fat. K.B.J. investigated somatotrophic axis signaling. G.C.C.V., J.D.M. and M.D. analysed resting heart functions by echocardiography. M.D. designed and conducted cardiac stress tests. A.P. and K.K. analysed leukocyte subpopulations. J.M.v.D., D.J.B. and B.G.C. wrote the paper with input from all co-authors. J.M.v.D. directed and supervised all aspects of the study in collaboration with D.J.B.
Corresponding authors
Ethics declarations
Competing interests
J.M.v.D. and D.J.B. are inventors on patents licensed to Unity Biotechnology by Mayo Clinic and J.M.v.D. is a co-founder of Unity Biotechnology.
Extended data figures and tables
Extended Data Figure 1 ATTAC transgene expression tracks with expression of senescence markers in iWAT and induces apoptosis of senescent cells after AP administration.
a, Comparative analysis of SA-β-Gal activity in intact iWAT. Scale bar, 0.5 cm. b, Analysis of endogenous Ink4a and ATTAC transcript SA-β-Gal activity in iWAT by qRT–PCR. H/H denotes BubR1H/H mice (n = 4 mice per group). c, FACS-based quantification of iWAT progenitor cell numbers in 18-month-old ATTAC mice treated with vehicle or AP. ASC, adipocyte stem cells; PAC, preadipocytes. d, Expression of the ATTAC transgene and senescence markers in iWAT as determined by qRT–PCR (n = 4 mice per group). Asterisks above individual bars denote significant changes to 2-month-old mice; asterisks above brackets denote significant differences between 18-month-old vehicle and AP-treated mice. e, Perirenal, mesenteric, subscapular and brown adipose tissue depot weights. SSAT, subscapular adipose tissue. f, SA-β-Gal activity in iWAT from 2-month-old ATTAC mice treated with vehicle or AP beginning at weaning age. g, p16Ink4a levels in iWAT from the mice described in f. Actin was used a loading control. h, Expression of ATTAC and senescence marker mRNA in the mice described in f (n = 3 mice per group). i–k, Early passage non-senescent ATTAC MEFs express p16Ink4a but are not susceptible to FKBP–Casp8-mediated elimination when cultured in the presence of AP. i, Levels of p16Ink4a in passage 3 ATTAC MEFs, with and without AP treatment. j, Growth curves of passage 3 ATTAC MEFs (n = 4 independently generated MEF lines per group), with or without AP treatment. k, Expression of ATTAC and senescence marker mRNA in passage 3 ATTAC MEFs (n = 3 independently generated MEF lines per group), with or without AP treatment. Error bars indicate s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001 (unpaired two-tailed t-tests). For gel source data, see Supplementary Fig. 1.
Extended Data Figure 2 ATTAC lacks promoter elements required for expression in replication-competent cells or aged lymphocytes expressing high levels of endogenous p16Ink4a.
a–c, SV40 large-T-antigen-immortalized ATTAC MEFs robustly express endogenous p16Ink4a (owing to SV40 large-T-antigen-mediated inactivation of Rb) but fail to engage the ATTAC transgene and are not subject to FKBP–Casp8-mediated elimination. a, p16Ink4a protein levels in passage 4 (P4) primary MEFs and MEFs immortalized with SV40 large T antigen. Actin was used as a loading control. b, p16Ink4a protein levels in immortalized MEFs treated with vehicle or two concentrations of AP. Actin was used as a loading control. c, Expression of ATTAC and senescence marker transcripts in passage 4 primary MEFs, vehicle-treated immortalized MEFs, and AP-treated immortalized MEFs (n = 3 independently generated MEF lines per group). d, Schematic representation of the endogenous Ink4a locus and the various Ink4a promoter regions driving ATTAC, 3MR and firefly luciferase (FLUC). ATTAC and p16-3MR mice have 2.6 kb and ~50 kb Ink4a promoter fragments driving transgene activity, respectively. p16-FLUC has firefly luciferase knocked into the endogenous Ink4a locus, which keeps the entire promoter region intact but ablates p16Ink4a protein expression. e, p16Ink4a protein levels in early passage primary and SV40 large-T-antigen-immortalized p16-3MR MEFs. f, Expression of senescence marker mRNA in early and late passage primary MEFs and SV40 large-T-antigen-immortalized p16-3MR MEFs (n = 1 independently generated MEF line per group performed in triplicate). g, Expression of senescence marker mRNA in early and late passage primary MEFs and SV40 large-T-antigen-immortalized p16-FLUC MEFs (n = 1 independently generated MEF line per group performed in triplicate). h, Expression of ATTAC and senescence markers in CD3+ T cells from 12- and 18-month-old ATTAC mice (n = 5 mice per group). Error bars indicate s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001 (unpaired two-tailed t-tests). We note that all values in f and g have P < 0.05 compared to passage 4 MEFs, with the exception of the one marked NS for not significant. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 3 ATTAC-mediated clearance of senescent cells is partial and tissue selective and attenuates expression of inflammation markers.
a, Expression of the ATTAC transgene and a senescence marker panel, as determined by RT–PCR, in gastrocnemius, eye, kidney, heart (atria), spleen, lung, liver and colon (n = 4 females per group). b, Expression of Il6, Il1a and Tnfa as determined by qRT–PCR in mouse iWAT, kidney and skeletal muscle at different ages (n = 4 females per group). Il6 values are as indicated in Extended Data Fig. 1d (iWAT) and in a (kidney and gastrocnemius). Expression levels of inflammation markers in unmanipulated 18-month-old C57BL/6 females suggests that repeated vehicle injections were not a source of tissue inflammation. Error bars indicate s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001 (unpaired two-tailed t-tests). Asterisks above individual bars in a denote significant changes to 2-month-old mice; asterisks above brackets denote significant differences between 18-month-old vehicle and AP-treated mice.
Extended Data Figure 4 Comparison of lifespans under different diets and housing facilities.
a, Survival curves of unmanipulated wild-type C57BL/6-129Sv mice fed a 5% versus 9% fat diet. Median lifespan (in days) are indicated. b, Survival curves of unmanipulated wild-type C57BL/6-129Sv mice fed a 9% fat diet plotted against those of vehicle-treated (−AP) and AP-treated (+AP) C57BL/6-129Sv-FVB ATTAC mice from Fig. 2b. These data suggest that the lifespans of vehicle-injected C57BL/6-129Sv-FVB control mice were quite normal for the diet that they were on, and unlikely to be negatively affected by repeated intraperitoneal injections. *P < 0.01; **P < 0.001; ***P < 0.001 (log-rank tests). c, d, Median survival data of unmanipulated C57BL/6 male (c) and female (d) mice from various laboratories for comparsion to the results obtained from our facility.
Extended Data Figure 5 Senescent cell clearance delays tumour and cataract formation.
a, b, Survival curves of mixed (a) and C57BL/6 (b) ATTAC mice dying of cancer (mice that had an overt tumour at time of death; only mice with lymphomas, sarcomas and carcinomas were included). Median survival (in days) and percentage increase are indicated. c, d, Incidence of macroscopically detectable neoplasms (lymphomas, sarcomas and carcinomas) at time of death in mixed (c) and C57BL/6 (d) ATTAC mice from survival cohorts. e, Survival curves of C57BL/6-129Sv-FVB mice dying without cancer (mice that had an overt tumour at time of death, including lymphoma, sarcoma and/or carcinoma, were excluded). Median survival (in days) and percentage increase are indicated. f, g, Cataract incidence for mixed (f) and C57BL/6 (g) ATTAC mouse cohorts. *P < 0.05; **P < 0.01; ***P < 0.001 (log-rank tests).
Extended Data Figure 6 Senescent cell clearance does not affect coordination, memory or exercise ability of 18-month-old ATTAC mice.
a, Time spent balanced during a fixed speed rotarod test for 18-month-old ATTAC mice (n = 6 male and 8 female mice per group). b, Novel object investigation test. The percentage of investigations of a novel object divided by the total investigations is graphed. Key and animal numbers are as indicated in a. c–e, Time-to-exhaustion (c), distance (d) and work (e) during a treadmill exercise test. Animal numbers are as indicated in c. f, Gastrocnemius muscle weight of ATTAC mice (n = 6 12-month-old males and females; n = 4 18-month-old −AP males and females; n = 4 18-month-old +AP males and females). g–i, Myofibre diameter measurements on isolated gastrocnemius (g), abdominal (h) and paraspinal muscle (i). Animal numbers are as indicated in f. j, Analysis of forelimb grip strength of ATTAC mice. Error bars indicate s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001 (unpaired two-tailed t-tests).
Extended Data Figure 7 Senescent cell clearance has no effect on haematological parameters and age-related changes in leukocyte populations.
a–l, Haematology results of 3- and 10-month-old untreated ATTAC C57BL/6 mice and 18-month-old vehicle- and AP-treated ATTAC C57BL/6 mice. White blood cell count (a), platelet count (b), red blood cell count (c), haemaglobin concentration (d), haematocrit (e), mean corpuscular volume (f), mean corpuscular haemoglobin (g), neutrophils (h), lymphocytes (i), basophils (j), monocytes (k) and eosinophils (l). m–q, Assessment for leukocyte subpopulations in 3- and 10-month-old untreated ATTAC C57BL/6 mice and 18-month-old vehicle- and AP-treated ATTAC C57BL/6 mice. CD4+ T cells (percentage of peripheral blood mononuclear cells, PBMC) (m), CD8+ T cells (percentage of PBMC) (n), CD44hi CD4+ T cells (percentage of CD4+) (o), CD44hi CD8+ T cells (percentage of CD8+) (p), and NK1.1+ cells (percentage of PBMC) (q). Error bars indicate s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001 (unpaired two-tailed t-tests).
Extended Data Figure 8 Senescent cell removal does not affect somatotrophic axis signalling in vivo.
a, Glucose levels following intraperitoneal glucose administration after an overnight fast in 18-month-old vehicle- and AP-treated ATTAC C57BL/6 females. b, Normalized glucose levels after intraperitoneal insulin administration following a 4-h fast in 18-month-old vehicle- and AP-treated ATTAC C57BL/6 females. c, Serum Igf1 levels in ATTAC C57BL/6 mice (n = 4 mice of each group). d, Representative western blots for phospho-S6K, total S6K, phospho-AKTS473 and total AKT in iWAT, kidney and skeletal muscle tissue lysates from 18-month-old vehicle- and AP-treated ATTAC C57BL/6 females. e, Quantification of phospho-S6K to total S6K ratio in blots from d, n = 4 mice of each group. f, Quantification of phospho-AKTS473 to total AKT ratio in blots from d, n = 4 mice of each group. Error bars indicate s.e.m. No statistically significant differences were observed in a–c, e and f using unpaired two-tailed t-tests. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 9 Senescent cell clearance does not alter cardiac morphology and function in ‘resting’ mice and AP treatment has no effect on healthspan of mice lacking the ATTAC transgene.
a, Electron micrographs of X-Gal crystal containing cells in the aortic root. VSMC, vascular smooth muscle cell. Scale bars, 1 μm (main panel) and 200 nm (inset). b–g, Echocardiography measurements of heart rate (b), left ventricular mass (c), posterior wall thickness (d), left ventricular inner diameter (e), ejection fraction (f), and the fractional shortening of the heart (g) in 12-month-old untreated mice and 18-month-old ATTAC mice treated with vehicle or AP. h, Fat mass (n = 9 mice per group). 18-month-old ATTAC vehicle-treated mouse values are the same as indicated in Fig. 1. i, iWAT and eWAT depot weight (n = 4 mice per group). 18-month-old ATTAC vehicle-treated mouse values are the same as indicated in Fig. 1. j, k, Kidney sclerosis (j) and blood urea levels (k) (n = 4 mice per group). 18-month-old ATTAC vehicle-treated mouse values are the same as indicated in Fig. 4. l, Time to death after isoproterenol administration (n = 4 mice per group). 18-month-old ATTAC vehicle-treated mouse values are the same as indicated in Fig. 5. Error bars indicate s.e.m. No statistically significant differences were observed using unpaired two-tailed t-tests.
Extended Data Figure 10 Effect of senescent cell clearance on wound healing and tissue fibrosis.
a, Closure of 3-mm punch biopsy wounds in 18-month-old ATTAC females after treatment with vehicle or AP for 6 months and if drug treatment was stopped 2 days before skin puncture or continued during wound closure (n = 6 wounds for −AP;−AP and +AP; −AP and n = 10 wounds for −AP;+AP and +AP;+AP). AP administration during the wound healing process significantly attenuates the rate of wound closure independently of whether senescent cell removal had occurred before wounding. b, Closure of 3-mm punch biopsy wounds in 4-month-old ATTAC females after treatment with vehicle or AP following wounding (n = 10 wounds per group). Similar to 18-month-old mice, AP administration during the wound healing process dramatically attenuated the rate of wound closure. c, Quantification of total GFP+ cells isolated from 3-mm punch biopsy wounds of 4-month-old mice two days into the wound healing process treated with vehicle (black) or AP (red, n = 3 mice per group). d, PTAH-stained tissues sections from 18-month-old ATTAC mice for detection of fibrosis. Scale bars, 100 μm. Error bars indicate s.e.m. Mice receiving AP during the healing process in a and b are significantly different from those treated with vehicle from day 1.5 to day 9.5. *P < 0.05 (unpaired two-tailed t-tests).
Supplementary information
Supplementary Information
This file contains Supplementary Text, Supplementary References and Supplementary Figure 1. (PDF 14896 kb)
Supplementary Data
This file contains Supplementary Table 1. (XLSX 224 kb)
Supplementary data
This file contains Supplementary Table 2. (XLSX 65 kb)
Source data
Rights and permissions
About this article
Cite this article
Baker, D., Childs, B., Durik, M. et al. Naturally occurring p16Ink4a-positive cells shorten healthy lifespan. Nature 530, 184–189 (2016). https://doi.org/10.1038/nature16932
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16932
This article is cited by
-
Distinguishing between driver and passenger mechanisms of aging
Nature Genetics (2024)
-
A review of the pathophysiological mechanisms of doxorubicin-induced cardiotoxicity and aging
npj Aging (2024)
-
Cholesterol biosynthetic pathway induces cellular senescence through ERRα
npj Aging (2024)
-
Targeting aging and age-related diseases with vaccines
Nature Aging (2024)
-
Alzheimer’s disease, aging, and cannabidiol treatment: a promising path to promote brain health and delay aging
Molecular Biology Reports (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.