Abstract
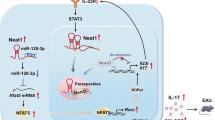
T helper 17 (TH17) lymphocytes protect mucosal barriers from infections, but also contribute to multiple chronic inflammatory diseases. Their differentiation is controlled by RORγt, a ligand-regulated nuclear receptor. Here we identify the RNA helicase DEAD-box protein 5 (DDX5) as a RORγt partner that coordinates transcription of selective TH17 genes, and is required for TH17-mediated inflammatory pathologies. Surprisingly, the ability of DDX5 to interact with RORγt and coactivate its targets depends on intrinsic RNA helicase activity and binding of a conserved nuclear long noncoding RNA (lncRNA), Rmrp, which is mutated in patients with cartilage-hair hypoplasia. A targeted Rmrp gene mutation in mice, corresponding to a gene mutation in cartilage-hair hypoplasia patients, altered lncRNA chromatin occupancy, and reduced the DDX5–RORγt interaction and RORγt target gene transcription. Elucidation of the link between Rmrp and the DDX5–RORγt complex reveals a role for RNA helicases and lncRNAs in tissue-specific transcriptional regulation, and provides new opportunities for therapeutic intervention in TH17-dependent diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
04 July 2018
This Article has been retracted; see accompanying Retraction. Corrected online 20 January 2016. In this Article, author Frank Rigo was incorrectly listed with a middle initial; this has been corrected in the online versions of the paper.
20 January 2016
A Correction to this paper has been published: https://doi.org/10.1038/nature16968
References
Weaver, C. T., Hatton, R. D., Mangan, P. R. & Harrington, L. E. IL-17 family cytokines and the expanding diversity of effector T cell lineages. Annu. Rev. Immunol. 25, 821–852 (2007)
Ivanov, I. I. et al. The orphan nuclear receptor RORγt directs the differentiation program of proinflammatory IL-17+ T helper cells. Cell 126, 1121–1133 (2006)
Leppkes, M. et al. RORγ-expressing Th17 cells induce murine chronic intestinal inflammation via redundant effects of IL-17A and IL-17F. Gastroenterology 136, 257–267 (2009)
Genovese, M. C. et al. LY2439821, a humanized anti-interleukin-17 monoclonal antibody, in the treatment of patients with rheumatoid arthritis: a phase I randomized, double-blind, placebo-controlled, proof-of-concept study. Arthritis Rheum. 62, 929–939 (2010)
Huh, J. R. et al. Digoxin and its derivatives suppress TH17 cell differentiation by antagonizing RORγt activity. Nature 472, 486–490 (2011)
Kondo, Y. et al. Involvement of RORγt-overexpressing T cells in the development of autoimmune arthritis in mice. Arthritis Res. Ther. 17, 105 (2015)
Gaffen, S. L., Jain, R., Garg, A. V. & Cua, D. J. The IL-23–IL-17 immune axis: from mechanisms to therapeutic testing. Nature Rev. Immunol. 14, 585–600 (2014)
Ciofani, M. et al. A validated regulatory network for Th17 cell specification. Cell 151, 289–303 (2012)
O’Malley, B. W. & Kumar, R. Nuclear receptor coregulators in cancer biology. Cancer Res. 69, 8217–8222 (2009)
Huang, Y. & Liu, Z. R. The ATPase, RNA unwinding, and RNA binding activities of recombinant p68 RNA helicase. J. Biol. Chem. 277, 12810–12815 (2002)
Fuller-Pace, F. V. & Moore, H. C. RNA helicases p68 and p72: multifunctional proteins with important implications for cancer development. Future Oncol. 7, 239–251 (2011)
Clark, E. L. et al. The RNA helicase p68 is a novel androgen receptor coactivator involved in splicing and is overexpressed in prostate cancer. Cancer Res. 68, 7938–7946 (2008)
Linder, P. & Jankowsky, E. From unwinding to clamping — the DEAD box RNA helicase family. Nature Rev. Mol. Cell Biol. 12, 505–516 (2011)
Arun, G., Akhade, V. S., Donakonda, S. & Rao, M. R. mrhl RNA, a long noncoding RNA, negatively regulates Wnt signaling through its protein partner Ddx5/p68 in mouse spermatogonial cells. Mol. Cell. Biol. 32, 3140–3152 (2012)
Caretti, G. et al. The RNA helicases p68/p72 and the noncoding RNA SRA are coregulators of MyoD and skeletal muscle differentiation. Dev. Cell 11, 547–560 (2006)
Lin, C., Yang, L., Yang, J. J., Huang, Y. & Liu, Z. R. ATPase/helicase activities of p68 RNA helicase are required for pre-mRNA splicing but not for assembly of the spliceosome. Mol. Cell. Biol. 25, 7484–7493 (2005)
Jalal, C., Uhlmann-Schiffler, H. & Stahl, H. Redundant role of DEAD box proteins p68 (Ddx5) and p72/p82 (Ddx17) in ribosome biogenesis and cell proliferation. Nucleic Acids Res. 35, 3590–3601 (2007)
Rosenbluh, J. et al. RMRP is a non-coding RNA essential for early murine development. PLoS ONE 6, e26270 (2011)
Hsieh, C. L. et al. The gene for the RNA component of the mitochondrial RNA-processing endoribonuclease is located on human chromosome 9p and on mouse chromosome 4. Genomics 6, 540–544 (1990)
Esakova, O. & Krasilnikov, A. S. Of proteins and RNA: the RNase P/MRP family. RNA 16, 1725–1747 (2010)
Mäkitie, O., Kaitila, I. & Savilahti, E. Susceptibility to infections and in vitro immune functions in cartilage-hair hypoplasia. Eur. J. Pediatr. 157, 816–820 (1998)
Bonafé, L. et al. Evolutionary comparison provides evidence for pathogenicity of RMRP mutations. PLoS Genet. 1, e47 (2005)
Bacchetta, J. et al. Autoimmune hypoparathyroidism in a 12-year-old girl with McKusick cartilage hair hypoplasia. Pediatr. Nephrol. 24, 2449–2453 (2009)
Jensen, E. D. et al. p68 (Ddx5) interacts with Runx2 and regulates osteoblast differentiation. J. Cell. Biochem. 103, 1438–1451 (2008)
Dardenne, E. et al. RNA helicases DDX5 and DDX17 dynamically orchestrate transcription, miRNA, and splicing programs in cell differentiation. Cell Rep. 7, 1900–1913 (2014)
Wortham, N. C. et al. The DEAD-box protein p72 regulates ERα-/oestrogen-dependent transcription and cell growth, and is associated with improved survival in ERα-positive breast cancer. Oncogene 28, 4053–4064 (2009)
Ivanov, I. I. et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 139, 485–498 (2009)
Powrie, F. et al. Inhibition of Th1 responses prevents inflammatory bowel disease in scid mice reconstituted with CD45RBhi CD4+ T cells. Immunity 1, 553–562 (1994)
Hirota, K. et al. Fate mapping of IL-17-producing T cells in inflammatory responses. Nature Immunol. 12, 255–263 (2011)
Wang, Y. et al. The transcription factors T-bet and Runx are required for the ontogeny of pathogenic interferon-γ-producing T helper 17 cells. Immunity 40, 355–366 (2014)
Sun, Z. et al. Requirement for RORγ in thymocyte survival and lymphoid organ development. Science 288, 2369–2373 (2000)
Chu, C., Quinn, J. & Chang, H. Y. Chromatin isolation by RNA purification (ChIRP). J. Vis. Exp. (2012)
Bonasio, R. & Shiekhattar, R. Regulation of transcription by long noncoding RNAs. Annu. Rev. Genet. 48, 433–455 (2014)
Gomez, J. A. et al. The NeST long ncRNA controls microbial susceptibility and epigenetic activation of the interferon-γ locus. Cell 152, 743–754 (2013)
Carpenter, S. et al. A long noncoding RNA mediates both activation and repression of immune response genes. Science 341, 789–792 (2013)
Maenner, S., Muller, M., Frohlich, J., Langer, D. & Becker, P. B. ATP-dependent roX RNA remodeling by the helicase maleless enables specific association of MSL proteins. Mol. Cell 51, 174–184 (2013)
Ilik, I. A. et al. Tandem stem-loops in roX RNAs act together to mediate X chromosome dosage compensation in Drosophila . Mol. Cell 51, 156–173 (2013)
Calo, E. et al. RNA helicase DDX21 coordinates transcription and ribosomal RNA processing. Nature 518, 249–253 (2015)
Yang, J., Sundrud, M. S., Skepner, J. & Yamagata, T. Targeting Th17 cells in autoimmune diseases. Trends Pharmacol. Sci. 35, 493–500 (2014)
Lee, Y. J., Holzapfel, K. L., Zhu, J., Jameson, S. C. & Hogquist, K. A. Steady-state production of IL-4 modulates immunity in mouse strains and is determined by lineage diversity of iNKT cells. Nature Immunol. 14, 1146–1154 (2013)
Takatori, H. et al. Lymphoid tissue inducer-like cells are an innate source of IL-17 and IL-22. J. Exp. Med. 206, 35–41 (2009)
Luci, C. et al. Influence of the transcription factor RORγt on the development of NKp46+ cell populations in gut and skin. Nature Immunol. 10, 75–82 (2009)
Chien, Y. H., Zeng, X. & Prinz, I. The natural and the inducible: interleukin (IL)-17-producing γδ T cells. Trends Immunol. 34, 151–154 (2013)
Kang, H. S. et al. Gene expression profiling reveals a regulatory role for RORα and RORγ in phase I and phase II metabolism. Physiol. Genomics 31, 281–294 (2007)
Yang, Y. et al. Focused specificity of intestinal TH17 cells towards commensal bacterial antigens. Nature 510, 152–156 (2014)
Sano, T. et al. An IL-23R/IL-22 circuit regulates epithelial serum amyloid A to promote local effector Th17 responses. Cell (2015)
Stanley, S. et al. Profiling of glucose-sensing neurons reveals that GHRH neurons are activated by hypoglycemia. Cell Metab. 18, 596–607 (2013)
Nicol, S. M. et al. The RNA helicase p68 (DDX5) is selectively required for the induction of p53-dependent p21 expression and cell-cycle arrest after DNA damage. Oncogene 32, 3461–3469 (2013)
Croswell, A., Amir, E., Teggatz, P., Barman, M. & Salzman, N. H. Prolonged impact of antibiotics on intestinal microbial ecology and susceptibility to enteric Salmonella infection. Infect. Immun. 77, 2741–2753 (2009)
Ostanin, D. V. et al. T cell transfer model of chronic colitis: concepts, considerations, and tricks of the trade. Am. J. Physiol. Gastrointest. Liver Physiol. 296, G135–G146 (2009)
Kim, S. V. et al. GPR15-mediated homing controls immune homeostasis in the large intestine mucosa. Science 340, 1456–1459 (2013)
Manel, N., Unutmaz, D. & Littman, D. R. The differentiation of human TH-17 cells requires transforming growth factor-β and induction of the nuclear receptor RORgammat. Nature Immunol. 9, 641–649 (2008)
Meng, L. et al. Towards a therapy for Angelman syndrome by targeting a long non-coding RNA. Nature 518, 409–412 (2015)
Bates, G. J. et al. The DEAD box protein p68: a novel transcriptional coactivator of the p53 tumour suppressor. EMBO J. 24, 543–553 (2005)
Morita, S., Kojima, T. & Kitamura, T. Plat-E: an efficient and stable system for transient packaging of retroviruses. Gene Ther. 7, 1063–1066 (2000)
Santori, F. R. et al. Identification of natural RORγ ligands that regulate the development of lymphoid cells. Cell Metab. 21, 286–297 (2015)
Huh, J. R. et al. Identification of potent and selective diphenylpropanamide RORgamma Inhibitors. ACS Med. Chem. Lett. 4, 79–84, (2013)
Chu, C., Qu, K., Zhong, F. L., Artandi, S. E. & Chang, H. Y. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Mol. Cell 44, 667–678 (2011)
Flynn, R. A. et al. Dissecting noncoding and pathogen RNA–protein interactomes. RNA 21, 135–143 (2015)
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9. (2015)
Acknowledgements
We thank M. V. Pokrovskii for unpublished ATAC-seq data and L. X. Garmire for suggestions on our manuscript. This work was supported by a Cancer Research Institute Irvington Postdoctoral Fellowship (W.H.), Institutional NRSA T32 CA009161_Levy (W.H.), National Multiple Sclerosis Society postdoctoral fellowship FG 2089-A-1 (L.W.), Career Development Award (329388) from the Crohn’s and Colitis Foundation of America (S.V.K.), Dale and Betty Frey Fellowship of the Damon Runyon Cancer Research Foundation 2105-12 (J.A.H.), HHMI Exceptional Research Opportunities Program (N.R.M. and N.H.), NIH F30 1F30CA189514-01 (R.A.F.), NIH P50-HG007735 and R01HG004361 (H.Y.C.), NIH R01AI080885 (D.R.L.), NIH R01DK103358 (R.B. and D.R.L.), and the Howard Hughes Medical Institute (H.Y.C. and D.R.L.).
Author information
Authors and Affiliations
Contributions
W.H. and D.R.L. designed experiments, analysed data and wrote the manuscript with input from the co-authors; B.T. and O.A. performed mass spectrometry studies; F.Ri. designed and synthesized control and Rmrp ASOs; S.J.G. and L.W. performed MOG-EAE immunization and blinded scoring; S.V.K. performed blinded histology scoring on colitis sections; W.H. and A.I.D. designed and performed ribosome TRAP-seq studies. S.M. and R.M.M. performed library preparation for RNA sequencing studies; N.R.M. and N.G.H. performed microscopy studies; F.Ra. provided recombinant full length His-tagged hRORγt, and F.V.F.-P. generated DDX5 conditional mutant animals. J.A.H. performed RORγt ChIP studies. C.P.N performed DDX5 studies in the thymus. R.A.F., W.H. and H.Y.C. performed ChIRP-seq experiments. E.R.M and R.B. performed statistical analyses on ChIRP-seq experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Identification of DDX5 as a RORγt-interacting partner.
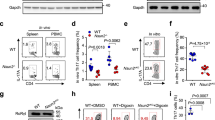
a, Mass spectrometry experimental workflow. Sorted naive CD4+ T cells from wild-type mice were cultured in vitro in TH17-polarizing conditions for 48 h. Immunoprecipitation of endogenous RORγt was performed using RORγ/γt-specific antibodies on whole-cell lysates. RORγt enrichment in pull-down was confirmed by immunoblot. Immunoprecipitated proteins were digested and analysed by mass spectrometry. The listed DDX5 peptides were identified in the TH17 RORγt immunoprecipitate. b, Co-immunoprecipitaton of DDX5 with anti-RORγt in lysates of in vitro polarized TH17 cells. c, Cell surface phenotype of splenic and lymph node DAPI−CD4+CD8α−CD19− T cells from wild-type and DDX5-T mice, examined by flow cytometry. d, Immunoblot of RORγt protein expression whole-cell lysate of cultured TH17 cells from wild-type or DDX5-T animals. For uncropped gels (b, d), see Supplementary Fig. 1. e, Immunofluorescence staining of RORγt in cultured TH17 cells from wild-type or DDX5-T mice. f, Immunofluorescence staining revealed nuclear localization of DDX5 in TH17 cells.
Extended Data Figure 2 DDX5 coregulates a subset of RORγt transcriptional targets in polarized TH17 cells.
a, Venn diagram of distinct and overlapping genes regulated by RORγt and/or DDX5, as determined from RNA-seq studies. b, Ingenuity Pathway Analysis (Qiagen) of DDX5- and RORγt-coregulated genes. c, IGV browser view showing biological replicate RNA-seq coverage tracks of control and DDX5-T from in vitro polarized TH17 cell samples at the Il17a, Il22, Ddx5 and Rorc loci. d, Independent RT–qPCR validation of RNA-seq results confirming effects of DDX5 deletion on RORγt target gene expression. Graphs show mean ± s.d.
Extended Data Figure 3 DDX5 chromatin localization in TH17 cells.
a, ChIP-seq-generated heatmap of DDX5 occupancy in regions centred on 16,003 RORγt-occupied sites (±2 kilobases (kb)). K-means linear normalization was used for clustering analysis by SeqMiner. Metagene analysis on cluster 1 depicts RORγt-occupied regions with DDX5 enrichment in wild-type but not DDX5-T cells; cluster 2 represents RORγt-occupied regions without DDX5 enrichment. b, IGV browser view of Il17a, Il17f and Rorc loci with ChIP-seq enrichment for RNA Pol II, RORγt and DDX5. c, Independent ChIP-qPCR of DDX5 in polarized TH17 cells. DDX5 occupancy at the Il17a and Il17f loci (as identified by RORγt ChIP-seq MACS peak called 32 and 39, respectively, from b) in control, DDX5-T or RORγt-deficient (RORγKO) cells. Results are representative of two independent experiments. Each experiment was performed with two technical replicates. Graph shows mean ± s.d. **P < 0.01 (unpaired, t-test).
Extended Data Figure 4 Influence of DDX5 on T-cell phenotypes in autoimmune disease models.
a, At 8 weeks after T-cell transfer, large intestine lamina propria mononuclear cells were evaluated for amounts of Il17a and Ifng mRNA by RT–qPCR. Results are representative of two independent experiments. Each experiment was performed using large intestines from three mice in each condition. RT–qPCR was performed with two technical replicates. Graph shows mean ± s.d. *P < 0.03 (unpaired, t-test). b, Gating strategy for analysis of TH17 and TH1 cells from large intestine of Rag2-deficient recipients of wild-type or DDX5-T naive T cells analysed at 8 weeks after T-cell transfer. c, Representative IL-17A and IFNγ intracellular staining of Aqua−CD4+RORγt+ TH17 cells in spinal cord of MOG-immunized animals on day 21.
Extended Data Figure 5 Noncoding RNAs enriched in DDX5 and RORγt RIP-seq studies.
a, DDX5-T cells were transduced with wild-type or helicase-mutant DDX5 and evaluated for DDX5 expression by immunofluorescence (left) and immunoblot (right) with anti-DDX5 antibody. For uncropped gels, see Supplementary Fig. 1. b, Venn diagram of noncoding RNAs detected by RIP-seq of ribosome-depleted TH17 cell lysates with anti-DDX5 and anti-RORγt antibodies. c, Abundance of top noncoding RNAs enriched in DDX5 and RORγt immunoprecipitates from polarized TH17 cell lysates depleted of ribosomes. Top, abundance of the noncoding RNAs in total lysate. d, RIP-qPCR experiments to compare Rmrp association with DDX5 in cultured TH17 and total thymocytes ex vivo. Results are representative of three independent experiments. Each experiment was performed with two technical replicates. Graph shows mean ± s.d. **P < 0.001 (unpaired, t-test).
Extended Data Figure 6 Rmrp and DDX5 knockdown in mouse and human TH17 cells.
a, RNA FISH analysis, using probes specific for Rmrp (green) and Malat1 (red) lncRNAs, in TH17 cells at 72 h after nucleofection with control (CTL) or Rmrp ASOs. b, Effect of Rmrp ASOs targeting different regions of Rmrp transcript on levels of Rmrp, Il17f and Ccr6 RNAs in polarized TH17 cells. c, Knockdown of DDX5 reduced IL-17A production in in vitro polarized human RORγt+ TH17 cells. **P < 0.01 (paired, t-test). Representative result shown in left panel. Each dot represents a different healthy donor (n = 4). Graphs show mean ± s.d.
Extended Data Figure 7 Effects of wild-type and mutant Rmrp in T-cell differentiation.
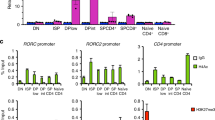
a, Il17a mRNA in cell lysates of in vitro polarized mouse TH17 cells at 96 h after transduction of control vector or wild-type Rmrp. Results are representative of two independent experiments. b, IFNγ production in polarized mouse TH1 cells at 96 h after transduction of control or Rmrp-encoding vector. Representative of two independent experiments. Each experiment was performed with two technical replicates. c, Comparison of human and mouse Rmrp sequences. Several mutations identified in CHH patients are highlighted. d, IL-17A production in polarized mouse TH17 cells at 96 h after transduction of wild-type or mutant Rmrp vectors. Representative of two independent experiments. e, Venn diagram depicting the number of distinct and overlapping genes regulated by RORγt, DDX5 and Rmrp in in vitro polarized TH17 cells. f, Expression of cytokine and Foxp3 mRNAs in T cells from wild-type or RmrpG270T/G270T mice cultured in vitro in TH17-, iTreg-, TH1- and TH2-polarizing conditions. Results are representative of two independent experiments. Each experiment was performed with two technical replicates. ***P < 0.001 (unpaired, t-test). g, ChIP-qPCR experiment using anti-RORγ/γt antibodies on chromatin of TH17 cells from wild-type or mutant mice cultured for 48 h in vitro. Each dot represents a different biological sample. Wild type, n = 2; RmrpG270T, n = 2. Results are representative of three separate independent experiments. Graphs show mean ± s.d. (unpaired, t-test).
Extended Data Figure 8 Effect of Ddx5 and Rmrp mutations in inflammation and thymocyte development.
a, Left, percentage weight change in Rag2−/− recipients of wild-type (black circles) or RmrpG270T/G270T (grey squares) naive CD4+ T cells in the transfer model of colitis. Animal weight was measured on day 56 (wild type, n = 8; RmrpG270T/G270T, n = 8, combined from three independent experiments). Graphs show mean ± s.d. ***P < 0.001 (unpaired, t-test). Middle, histology score (scale of 0–24) (wild type, n = 8; RmrpG270T/G270T, n = 5), combined from two independent experiments. **P < 0.01 (unpaired, t-test). Right, representative H&E staining of large intestine from Rag2−/− mice on day 56 after naive T-cell transfer. b, Mice with deletion of Ddx5 in early common lymphoid progenitors (DDX5-clpKO) have normal thymic development. Left, immunoblot of thymocyte lysates with anti-DDX5 antibody confirmed depletion of DDX5. Right, percentage of CD4 single-positive (SP), CD8α SP, double-positive (DP) and double-negative (DN) cells among total thymocytes. Each bar represents the result from one mouse (WT/het, n = 9; DDX5-clpKO, n = 6). For uncropped gels, see Supplementary Fig. 1. c, Thymocyte and peripheral T-cell surface phenotypes of wild-type and RmrpG270T/G270T knock-in mice at steady state. Peripheral T-cell gate, DAPI− CD19−CD8α−CD4+.
Extended Data Figure 9 Association of Rmrp lncRNA with DDX5 and RORγt in vitro.
a, In vitro translated (TNT) HA-tagged wild-type or helicase-dead DDX5 and Flag-tagged RORγt were incubated with in vitro transcribed Rmrp. After capture on anti-HA or anti-Flag beads, the amount of lncRNA was determined by RT–qPCR. Data are representative of two independent experiments, and each experiment was performed with two technical replicates. b, Helicase requirement for in vitro interaction of DDX5 and RORγt. Recombinant GST–DDX5 (wild type or helicase-dead mutant) and His–RORγt full-length protein were synthesized in Escherichia coli, purified, and assayed for binding with or without in vitro transcribed Rmrp RNA in the presence exogenous ATP. c, Association of in vitro transcribed wild-type and mutant Rmrp with recombinant GST–DDX5 captured on glutathione beads (left) or with recombinant GST–DDX5 and His–RORγt captured with anti-His antibody. Amounts of associated Rmrp were quantified using RT–qPCR. Data are representative of two independent experiments. Each experiment was performed with two technical replicates. d, Comparison of ability of in vitro transcribed wild-type and RmrpG270T lncRNA to promote interaction between recombinant RORγt and DDX5 in vitro. All graphs show mean ± s.d. ***P < 0.001 (unpaired, t-test). For uncropped gels, see Supplementary Fig. 1.
Extended Data Figure 10 Rmrp chromatin localization in TH17 cells.
a, ChIRP-seq sample validation of Rmrp RNA pull-down over other nuclear noncoding RNAs using pools of ‘even’ or ‘odd’ capture probes. Graphs show mean ± s.d. b, ChIRP-qPCR of Rmrp RNA pull-down from wild-type TH17 cell lysate treated with or without RNase (n = 2). qPCR for each sample was performed with two technical replicates. Graph shows mean ± s.d. **P < 0.001 (unpaired, t-test). c, HOMER motif analysis reveals top three DNA motifs within Rmrp-enriched peaks. d, Significance of peak overlaps between Rmrp ChIRP-seq and ChIP-seq for BATF (n = 2), IRF4 (n = 7), STAT3 (n = 2), c-Maf (n = 2), RORγt (n = 2), CTCF (n = 2), RNA Pol II (n = 2), H3K27me3 (n = 4) and H3K4me3 (n = 3) in TH17 cells (hypergeometric distribution). Each dot represents a separate biological replicate of ChIP-seq experiments. e, Venn diagram depicting changes in peaks called from Rmrp (ChIRP-seq) experiments in wild-type and DDX5-T TH17 cells. f, Comparison of Rmrp chromatin occupancy (ChIRP-seq) at known RORγt occupied loci in in vitro polarized TH17 cells from wild-type and RmrpG270T/G270T mice.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1 and gel images. (PDF 477 kb)
Source data
Rights and permissions
About this article
Cite this article
Huang, W., Thomas, B., Flynn, R. et al. DDX5 and its associated lncRNA Rmrp modulate TH17 cell effector functions. Nature 528, 517–522 (2015). https://doi.org/10.1038/nature16193
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16193
This article is cited by
-
Construction of immune-related lncRNA signature to predict aggressiveness, immune landscape, and drug resistance of colon cancer
BMC Gastroenterology (2022)
-
The Airway Microbiome-IL-17 Axis: a Critical Regulator of Chronic Inflammatory Disease
Clinical Reviews in Allergy & Immunology (2022)
-
Nucleic Acids as Novel Therapeutic Modalities to Address Multiple Sclerosis Onset and Progression
Cellular and Molecular Neurobiology (2022)
-
Linc00473 potentiates cholangiocarcinoma progression by modulation of DDX5 expression via miR-506 regulation
Cancer Cell International (2020)
-
Functional polymorphisms in LncRNA HOTAIR contribute to susceptibility of pancreatic cancer
Cancer Cell International (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.