Abstract
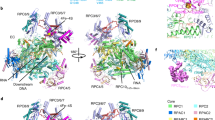
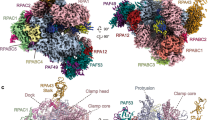
Transcription of genes encoding small structured RNAs such as transfer RNAs, spliceosomal U6 small nuclear RNA and ribosomal 5S RNA is carried out by RNA polymerase III (Pol III), the largest yet structurally least characterized eukaryotic RNA polymerase. Here we present the cryo-electron microscopy structures of the Saccharomyces cerevisiae Pol III elongating complex at 3.9 Å resolution and the apo Pol III enzyme in two different conformations at 4.6 and 4.7 Å resolution, respectively, which allow the building of a 17-subunit atomic model of Pol III. The reconstructions reveal the precise orientation of the C82–C34–C31 heterotrimer in close proximity to the stalk. The C53–C37 heterodimer positions residues involved in transcription termination close to the non-template DNA strand. In the apo Pol III structures, the stalk adopts different orientations coupled with closed and open conformations of the clamp. Our results provide novel insights into Pol III-specific transcription and the adaptation of Pol III towards its small transcriptional targets.
This is a preview of subscription content, access via your institution
Access options





Similar content being viewed by others
Accession codes
Primary accessions
Electron Microscopy Data Bank
Protein Data Bank
Data deposits
The 3.9 Å cryo-EM map of elongating Pol III and the two maps of apo Pol III have been deposited in the Electron Microscopy Data Bank with accession codes EMD-3178 (elongating Pol III) and EMD-3179 (‘closed clamp’ apo Pol III) EMD-3180 (‘open clamp’ apo Pol III). The coordinates of the corresponding atomic models have been deposited in the Protein Data Bank under accession code 5fj8 (elongating Pol III) and 5fj9 (‘closed clamp’ apo Pol III), 5fja (‘open clamp’ apo Pol III).
References
Dieci, G., Fiorino, G., Castelnuovo, M., Teichmann, M. & Pagano, A. The expanding RNA polymerase III transcriptome. Trends Genet. 23, 614–622 (2007)
White, R. J. RNA polymerases I and III, non-coding RNAs and cancer. Trends Genet. 24, 622–629 (2008)
Vannini, A. & Cramer, P. Conservation between the RNA polymerase I, II, and III transcription initiation machineries. Mol. Cell 45, 439–446 (2012)
Arimbasseri, A. G. & Maraia, R. J. Mechanism of transcription termination by RNA polymerase III utilizes a non-template strand sequence-specific signal element. Mol. Cell 58, 1124–1132 (2015)
Chédin, S., Riva, M., Schultz, P., Sentenac, A. & Carles, C. The RNA cleavage activity of RNA polymerase III is mediated by an essential TFIIS-like subunit and is important for transcription termination. Genes Dev. 12, 3857–3871 (1998)
Werner, M., Thuriaux, P. & Soutourina, J. Structure-function analysis of RNA polymerases I and III. Curr. Opin. Struct. Biol. 19, 740–745 (2009)
Jasiak, A. J., Armache, K. J., Martens, B., Jansen, R. P. & Cramer, P. Structural biology of RNA polymerase III: subcomplex C17/25 X-ray structure and 11 subunit enzyme model. Mol. Cell 23, 71–81 (2006)
Kassavetis, G. A., Prakash, P. & Shim, E. The C53/C37 subcomplex of RNA polymerase III lies near the active site and participates in promoter opening. J. Biol. Chem. 285, 2695–2706 (2010)
Landrieux, E. et al. A subcomplex of RNA polymerase III subunits involved in transcription termination and reinitiation. EMBO J. 25, 118–128 (2006)
Carter, R. & Drouin, G. The increase in the number of subunits in eukaryotic RNA polymerase III relative to RNA polymerase II is due to the permanent recruitment of general transcription factors. Mol. Biol. Evol. 27, 1035–1043 (2010)
Geiger, S. R. et al. RNA polymerase I contains a TFIIF-related DNA-binding subcomplex. Mol. Cell 39, 583–594 (2010)
Fernández-Tornero, C. et al. Insights into transcription initiation and termination from the electron microscopy structure of yeast RNA polymerase III. Mol. Cell 25, 813–823 (2007)
Wu, C. C., Lin, Y. C. & Chen, H. T. The TFIIF-like Rpc37/53 dimer lies at the center of a protein network to connect TFIIIC, Bdp1, and the RNA polymerase III active center. Mol. Cell. Biol. 31, 2715–2728 (2011)
Lane, L. A. et al. Mass spectrometry reveals stable modules in holo and apo RNA polymerases I and III. Structure 19, 90–100 (2011)
Wang, Z. & Roeder, R. G. Three human RNA polymerase III-specific subunits form a subcomplex with a selective function in specific transcription initiation. Genes Dev. 11, 1315–1326 (1997)
Wu, C. C. et al. RNA polymerase III subunit architecture and implications for open promoter complex formation. Proc. Natl Acad. Sci. USA 109, 19232–19237 (2012)
Thuillier, V., Stettler, S., Sentenac, A., Thuriaux, P. & Werner, M. A mutation in the C31 subunit of Saccharomyces cerevisiae RNA polymerase III affects transcription initiation. EMBO J. 14, 351–359 (1995)
Brun, I., Sentenac, A. & Werner, M. Dual role of the C34 subunit of RNA polymerase III in transcription initiation. EMBO J. 16, 5730–5741 (1997)
Khoo, S. K., Wu, C. C., Lin, Y. C., Lee, J. C. & Chen, H. T. Mapping the protein interaction network for TFIIB-related factor Brf1 in the RNA polymerase III preinitiation complex. Mol. Cell. Biol. 34, 551–559 (2014)
Boguta, M. Maf1, a general negative regulator of RNA polymerase III in yeast. Biochim. Biophys. Acta 1829, 376–384 (2013)
Vannini, A. et al. Molecular basis of RNA polymerase III transcription repression by Maf1. Cell 143, 59–70 (2010)
Sainsbury, S., Bernecky, C. & Cramer, P. Structural basis of transcription initiation by RNA polymerase II. Nature Rev. Mol. Cell Biol. 16, 129–143 (2015)
Svetlov, V. & Nudler, E. Basic mechanism of transcription by RNA polymerase II. Biochim. Biophys. Acta 1829, 20–28 (2013)
Fernández-Tornero, C. et al. Crystal structure of the 14-subunit RNA polymerase I. Nature 502, 644–649 (2013)
Engel, C., Sainsbury, S., Cheung, A. C., Kostrewa, D. & Cramer, P. RNA polymerase I structure and transcription regulation. Nature 502, 650–655 (2013)
Fernández-Tornero, C. et al. Conformational flexibility of RNA polymerase III during transcriptional elongation. EMBO J. 29, 3762–3772 (2010)
Moreno-Morcillo, M. et al. Solving the RNA polymerase I structural puzzle. Acta Crystallogr. D 70, 2570–2582 (2014)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Barnes, C. O. et al. Crystal structure of a transcribing RNA polymerase II complex reveals a complete transcription bubble. Mol. Cell 59, 258–269 (2015)
Gnatt, A. L., Cramer, P., Fu, J., Bushnell, D. A. & Kornberg, R. D. Structural basis of transcription: an RNA polymerase II elongation complex at 3.3Å resolution. Science 292, 1876–1882 (2001)
Kettenberger, H., Armache, K. J. & Cramer, P. Complete RNA polymerase II elongation complex structure and its interactions with NTP and TFIIS. Mol. Cell 16, 955–965 (2004)
Lefèvre, S. et al. Structure-function analysis of hRPC62 provides insights into RNA polymerase III transcription initiation. Nature Struct. Mol. Biol. 18, 352–358 (2011)
Aravind, L., Anantharaman, V., Balaji, S., Babu, M. M. & Iyer, L. M. The many faces of the helix-turn-helix domain: transcription regulation and beyond. FEMS Microbiol. Rev. 29, 231–262 (2005)
He, Y., Fang, J., Taatjes, D. J. & Nogales, E. Structural visualization of key steps in human transcription initiation. Nature 495, 481–486 (2013)
Schramm, L. & Hernandez, N. Recruitment of RNA polymerase III to its target promoters. Genes Dev. 16, 2593–2620 (2002)
Ferri, M. L. et al. A novel subunit of yeast RNA polymerase III interacts with the TFIIB-related domain of TFIIIB70. Mol. Cell. Biol. 20, 488–495 (2000)
Plaschka, C. et al. Architecture of the RNA polymerase II-Mediator core initiation complex. Nature 518, 376–380 (2015)
Arimbasseri, A. G., Rijal, K. & Maraia, R. J. Comparative overview of RNA polymerase II and III transcription cycles, with focus on RNA polymerase III termination and reinitiation. Transcription 5, e27369 (2014)
Hu, H. L., Wu, C. C., Lee, J. C. & Chen, H. T. A region of Bdp1 necessary for transcription initiation that is located within the RNA polymerase III active site cleft. Mol. Cell. Biol. 35, 2831–2840 (2015)
Rijal, K. & Maraia, R. J. RNA polymerase III mutants in TFIIFα-like C37 that cause terminator readthrough with no decrease in transcription output. Nucleic Acids Res. 41, 139–155 (2013)
Shaaban, S. A., Krupp, B. M. & Hall, B. D. Termination-altering mutations in the second-largest subunit of yeast RNA polymerase III. Mol. Cell. Biol. 15, 1467–1478 (1995)
Richard, P. & Manley, J. L. Transcription termination by nuclear RNA polymerases. Genes Dev. 23, 1247–1269 (2009)
Whitehall, S. K., Bardeleben, C. & Kassavetis, G. A. Hydrolytic cleavage of nascent RNA in RNA polymerase III ternary transcription complexes. J. Biol. Chem. 269, 2299–2306 (1994)
Iben, J. R. et al. Point mutations in the Rpb9-homologous domain of Rpc11 that impair transcription termination by RNA polymerase III. Nucleic Acids Res. 39, 6100–6113 (2011)
Ehara, H., Sekine, S. & Yokoyama, S. Crystal structure of the C17/25 subcomplex from Schizosaccharomyces pombe RNA polymerase III. Protein Sci. 20, 1558–1565 (2011)
Chakraborty, A. et al. Opening and closing of the bacterial RNA polymerase clamp. Science 337, 591–595 (2012)
Jun, S. H. et al. The X-ray crystal structure of the euryarchaeal RNA polymerase in an open-clamp configuration. Nature Commun. 5, 5132 (2014)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nature Methods 10, 584–590 (2013)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999)
Scheres, S. H. Beam-induced motion correction for sub-megadalton cryo-EM particles. eLife 3, e03665 (2014)
van Heel, M. & Schatz, M. Fourier shell correlation threshold criteria. J. Struct. Biol. 151, 250–262 (2005)
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013)
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nature Methods 11, 63–65 (2014)
Kosinski, J., Barbato, A. & Tramontano, A. MODexplorer: an integrated tool for exploring protein sequence, structure and function relationships. Bioinformatics 29, 953–954 (2013)
Söding, J. Protein homology detection by HMM-HMM comparison. Bioinformatics 21, 951–960 (2005)
Šali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Kosinski, J. et al. Xlink Analyzer: software for analysis and visualization of cross-linking data in the context of three-dimensional structures. J. Struct. Biol. 189, 177–183 (2015)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Bui, K. H. et al. Integrated structural analysis of the human nuclear pore complex scaffold. Cell 155, 1233–1243 (2013)
Ashkenazy, H., Erez, E., Martz, E., Pupko, T. & Ben-Tal, N. ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res. 38, W529–W533 (2010)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Harding, M. M. Geometry of metal-ligand interactions in proteins. Acta Crystallogr. D 57, 401–411 (2001)
Harding, M. M. Small revisions to predicted distances around metal sites in proteins. Acta Crystallogr. D 62, 678–682 (2006)
Colliex, C. et al. in International Tables of Crystallography Vol. C (ed Prince, E. ), 259–429 (International Union of Crystallography, 2006)
Sachse, C. et al. High-resolution electron microscopy of helical specimens: a fresh look at tobacco mosaic virus. J. Mol. Biol. 371, 812–835 (2007)
DiMaio, F., Zhang, J., Chiu, W. & Baker, D. Cryo-EM model validation using independent map reconstructions. Protein Sci. 22, 865–868 (2013)
Jiang, J. S. & Brünger, A. T. Protein hydration observed by X-ray diffraction. Solvation properties of penicillopepsin and neuraminidase crystal structures. J. Mol. Biol. 243, 100–115 (1994)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Schrodinger, L. L. C. The PyMOL Molecular Graphics System, Version 1.3r1 (2010)
Acknowledgements
We thank Y. Sadian, H. Grötsch and R. Wetzel for help in Pol III purification. We are grateful to F. Thommen and M. Wahlers for set-up and maintenance of the high-performance computational environment of RELION. We acknowledge F. Schur, S. Fromm, C. Bertipaglia, M. Beck and G. Hofhaus for helpful advice regarding sample preparation and image processing. We also thank the ‘Fermentation et culture de microorganisms’ (IFR88, CNRS, Marseille). M.M.-M. and A.J.J. were supported by Marie-Sklodowska-Curie fellowships (FP7-PEOPLE-2011IEF301002, PIEF-GA-2012-331285). N.A.H. acknowledges support by the EMBL International PhD program, A.J.J. and J.K. by postdoctoral fellowships from the EMBL Interdisciplinary Postdoc Program (EIPOD) under Marie Curie COFUND actions (PCOFUND-GA-2008-229597) and C.W.M. by an ERC Advanced Grant (ERC-2013-AdG340964-POL1PIC).
Author information
Authors and Affiliations
Contributions
C.W.M. and C.S. initiated and supervised the project. N.A.H. established and performed Pol III experiments with input from C.W.M., S.G. and M.M.-M.; N.A.H., C.S. and W.J.H.H. established the grid freezing and acquisition parameters, N.A.H. and W.J.H.H. collected all cryo-EM data. N.A.H., M.M.-M. and A.J.J. built the initial model. N.A.H., A.J.J. and C.S. analysed cryo-EM data. J.K. performed homology modelling and alignments, A.J.J. established the refinement protocol. A.J.J. and N.A.H. optimized the refinement protocol and revised the models. N.A.H., A.J.J., S.G., M.M.-M., C.S. and C.W.M. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Pol III processing pipeline, Fourier-shell correlation curves and local resolution assessment.
a, Exemplary micrograph of elongating Pol III. All micrographs were low-pass filtered for particle picking. b, General processing pipeline. The orange boxes display micrograph number and particles for elongating Pol III (left) and apo Pol III (right). The middle panel shows the general workflow that was followed for both data sets. For elongating Pol III (bottom left), a local classification step yielded one class with 49,543 particles (purple) that was subsequently refined and post-processed. For apo Pol III (bottom right), local classification diverged into two classes (purple) with 68,818 particles and 52,423 particles that were subsequently refined and post-processed. c, Fourier-shell correlation (FSC) and local resolution assessment with RESMAP55. All FSC calculations were performed with two independent half maps using RELION’s masking procedure53. The resolution for the elongating Pol III cryo-EM map (top panel) is 3.9 Å according to the FSC 0.143 criterion, indicated by the black dashed line. The two apo Pol III cryo-EM reconstructions have a resolution of 4.6 Å (closed clamp Pol III, middle panel) and 4.7 Å (open clamp Pol III, bottom panel) according to the FSC 0.143 criterion. Local resolution is displayed on the post-processed full maps (first image column on the right) and a cross-section representation (second image column on the right). In both apo Pol III reconstructions, the peripheral subcomplexes show a strong decay in resolution compared to the core. In the elongating Pol III reconstruction, the resolution is more uniformly distributed, indicating stabilization of peripheral subunits.
Extended Data Figure 2 Representative sections of the cryo-EM density for elongating Pol III.
a, Cross-section of elongating Pol III in ribbon and stick representation, embedded in the experimental density at 3.9 Å, displayed in dark blue. b, Section displaying the core subunits C160 (grey) and C128 (wheat) shown in stick and ribbon representation. The experimental density of the core (dark blue) is well defined and has been filtered at 3.5 Å resolution for display. c, Section of stalk subunits C25 (blue) and C17 (pink). The estimated local resolution in this part is lower compared to the core (Extended Data Table 2). In c, d and e, the cryo-EM density is shown at 3.9 Å resolution. d, Section showing subunits C53 (blue) and C37 (purple). e, Close-up view of C82-WH1 (brown), C82-WH2 (green) and C31 (yellow) interface.
Extended Data Figure 3 Comparison of electron microscopy densities with X-ray electron densities for shared subunits ABC23 (Rpb6) and ABC14.5 (Rpb8).
Top left shows Pol III in front view, a stretch in ABC23 (cyan) and ABC14.5 (green) is coloured. The red boxes indicate the regions that are enlarged in the neighbouring panels. Corresponding density is displayed in tungsten. Models of Pol II and Pol I at nominally higher resolution are available, but for better comparison models in a similar resolution range are shown. For the 2Fo − Fc electron density maps obtained by X-ray crystallography a threshold of 1σ was used for display. The top right shows three close-up views of the shared subunit ABC23 from elongating Pol III, Pol II (PDB 1wcm) and Pol I (PDB 4c3j). The bottom panels show 6 strands of shared subunit ABC14.5. Front view of the β-sheet and orthogonal views of individual strands in elongating Pol III, Pol II (PDB 1wcm) and Pol I (PDB 4c3j).
Extended Data Figure 4 Model validation and temperature factor distribution of atomic models.
a–c, FSC curves calculated between the refined atomic model and the half map used in refinement (FSCwork) are shown in blue, those calculated between the refined atomic model and the second half map not used for refinement (FSCtest) in red. Vertical lines mark the regular FSC 0.143 cutoff and the resolution target used in refinement as shown. Close agreement between FSCwork and FSCtest and the absence of a sharp drop beyond the refinement target resolution indicate that no overfitting took place. The respective FSC between the refined atomic model and the map obtained from 3D reconstruction using the entire data set (FSCref) is also shown (black). d, e, Atomic B-factor distributions mapped on ribbon representations of elongating and apo Pol III. The overall distribution and relative differences between core and peripheral subunits for the different models correlate well with the distribution of local resolution (Extended Data Fig. 1).
Extended Data Figure 5 Pol III-specific features of subunits C160 and C128 and comparison to the homologous Pol II and Pol I subunits.
a, Top (left) and front (right) view of Pol III, with subunits C160 and C128 displayed in ribbon representation and the other subunits in surface representation (grey). Coloured stretches highlight characteristic features denoted in b. b, Bar diagram shows the domain organization of Pol III C160. Arrows and corresponding numbers below the bar diagram indicate insertions and deletions of five or more residues in Pol III relative to Pol II subunit Rpb1 as indicated by structure-based alignment. Coloured regions are also shown in Pol III subunit C160 (lower panel, left) and in a. Lower panel middle and right show Pol II Rpb1 and Pol I A190 subunits, respectively. c, Same as in b for Pol III subunit C128. Insertions and deletions compared to Pol II subunit Rpb2 are displayed in the box diagram.
Extended Data Figure 6 Open and closed clamp conformation in Pol III compared to other RNA polymerases.
a, Top view of aligned elongating Pol III and apo Pol III (left panel, closed clamp; middle panel, open clamp) and both closed clamp and open clamp apo Pol III conformations (right panel). r.m.s.d. values (core-heterodimer:all) for elongating Pol III – apo Pol III (closed clamp) (0.43 Å3,490 Cα atoms aligned:0.43 Å4,813 Cα atoms aligned), elongating Pol III – apo Pol III (open clamp) (0.71 Å3,496 Cα atoms aligned:2.73 Å4,795 Cα atoms aligned) and both apo Pol III open and closed clamp (0.71 Å3,540 Cα atoms aligned:2.71 Å4,829 Cα atoms aligned) demonstrate the similarity between closed clamp apo Pol III and elongating Pol III conformations. b, Schematic representation of Pol III in top view showing the conformational changes of clamp head, heterotrimer and stalk. The closed clamp conformation (elongating Pol III and closed clamp apo Pol III) is displayed in red, the open clamp conformation (open clamp apo Pol III) in green. The DNA–RNA duplex is shown in blue, the core and heterodimer in grey. c, Front view on open and closed clamp conformations in other RNA polymerases. The closed clamp state (green) and open clamp state (red) is indicated for archaeal polymerase (left panel, PDB 4ayb and 4qiw), for Pol II (middle panel, PDB 1wcm and 1twf) and for Pol III (right panel). Green and red angles describe the cleft opening in the closed and open clamp conformations. Black arrows and corresponding values indicate the relative distance of the subunits between the two conformations. d, Front view of apo Pol III closed clamp (left panel), apo Pol III closed clamp vs apo Pol II (middle panel, Pol II (PDB 1wcm) in red) and apo Pol III closed clamp vs apo Pol I (right panel, Pol I (PDB 4c3i) in blue). The cleft opening is indicated by a dashed line and the Cα-Cα distance across the cleft (black for Pol III, red for Pol II and blue for Pol I). However, some of the observed differences in cleft width between Pol I, Pol II and Pol III might also reflect differences between conditions of cryo-EM and crystal structures as well as different packing contacts in the crystals.
Extended Data Figure 7 Pol III-specific subunits C82, C31 and C11.
a, Left panel: overall surface representation of Pol III with the C82–C34–C31 heterotrimer in ribbon representation. Right panels: two enlarged and orthogonal views of the region marked with a dotted black square. In subunit C82 WH4 inserts in the DNA binding cleft passing through a canyon in the clamp head. WH2 and WH3 extensions reach over the clamp head and are positioned in close proximity to downstream DNA. b, Ribbon model of Pol III fitted into the electron microscopy density of the open clamp apo Pol III filtered at 6 Å resolution. For C31, additional density is visible in the cavity between the stalk and the heterotrimer, as shown in the top right panel. The described densities are also present in the closed clamp apo Pol III and the elongating Pol III reconstructions. No attempts were made to fit atomic models into these densities. c, Electron microscopy density of the C11 TFIIS-like domain at 6 Å resolution as observed in the open clamp apo Pol III reconstruction. The left panel shows a side view of Pol III, the middle and right panels show close-ups at two different density thresholds.
Extended Data Figure 8 C53–C37 heterodimer and stalk subunits C25–C17.
a, Visualization of the photo-crosslinks between C53–C37 heterodimer and subunit C128. Pol III is shown in surface, C53–C37 and C11 in ribbon representation. In addition, the C128 lobe is shown in cartoon representation (small inset). Purple spheres on the lobe indicate residues that photo-crosslink to C37 (ref. 13), beige spheres on C37 indicate residues that photo-crosslink to C128 (ref. 13). The dashed line marks the tentative path of the non-template DNA strand. The experimental photo-crosslinks fit well to the cryo-EM structure. The C37 loop is disordered between Glu196 and Asn225, although photo-crosslinks indicate that this region is in close proximity to the lobe and the non-template DNA strand. b, Bottom view of Pol III in surface representation, with C53–C37 and C11 shown in ribbon representation. The black dotted square indicates the enlarged area in the centre of the image (small inset). The red density (shown at 4.5 Å) was not of sufficient quality to build an atomic model. However, photo-crosslinks from C37 and C128 to C53 (blue spheres on C37 ribbon and C128 surface mark crosslink positions)13 indicate that C53 N-terminal residues are located in this region. c, Stalk anchoring with C160 extensions. Top view (left panel) and bottom view (right panel) of the stalk subunits C17 (magenta), C25 (blue) and the C160 extensions (grey). Electron microscopy density corresponding to the C160 N- and C-terminal extensions is shown in tungsten blue. Individual entities and subunits are labelled.
Rights and permissions
About this article
Cite this article
Hoffmann, N., Jakobi, A., Moreno-Morcillo, M. et al. Molecular structures of unbound and transcribing RNA polymerase III. Nature 528, 231–236 (2015). https://doi.org/10.1038/nature16143
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16143
This article is cited by
-
Truncated PARP1 mediates ADP-ribosylation of RNA polymerase III for apoptosis
Cell Discovery (2022)
-
Structural insights into nuclear transcription by eukaryotic DNA-dependent RNA polymerases
Nature Reviews Molecular Cell Biology (2022)
-
A cancer-associated RNA polymerase III identity drives robust transcription and expression of snaR-A noncoding RNA
Nature Communications (2022)
-
Mechanism of RNA polymerase III termination-associated reinitiation-recycling conferred by the essential function of the N terminal-and-linker domain of the C11 subunit
Nature Communications (2021)
-
Activation and repression at the heart of human RNA polymerase III
Nature Structural & Molecular Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.